Search Count: 940
All
Selected
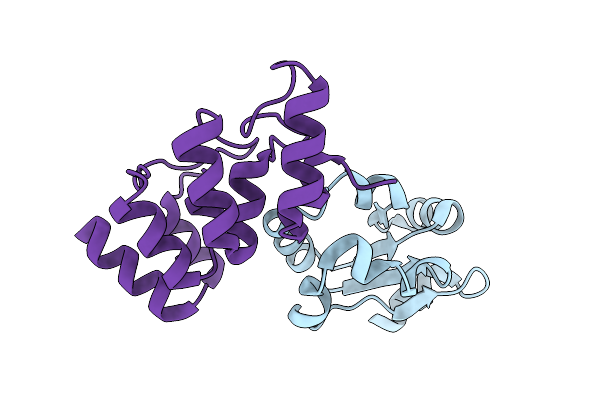 |
Crystal Structure Of The Acl1 Ankyrin Repeat Domain In Complex With The Second Rpl1 Domain
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-04-08 Classification: RIBOSOMAL PROTEIN |
 |
Organism: Oceanobacillus iheyensis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: RNA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: PROTEIN FIBRIL |
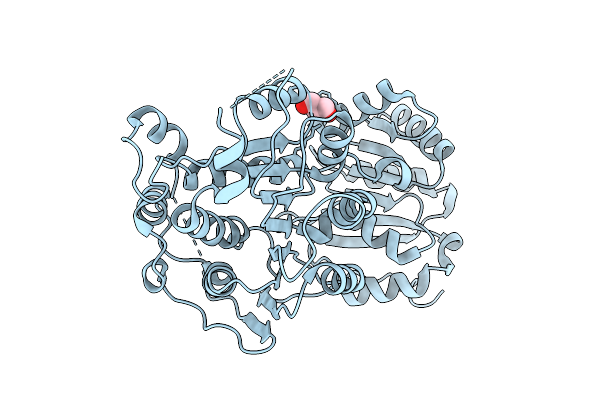 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-10-29 Classification: RNA BINDING PROTEIN Ligands: GOL |
 |
Organism: Streptococcus thermophilus
Method: SOLUTION NMR Release Date: 2025-10-22 Classification: NUCLEAR PROTEIN |
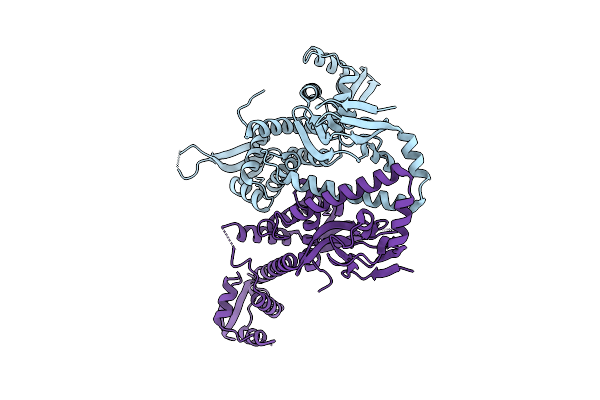 |
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2025-10-22 Classification: SIGNALING PROTEIN |
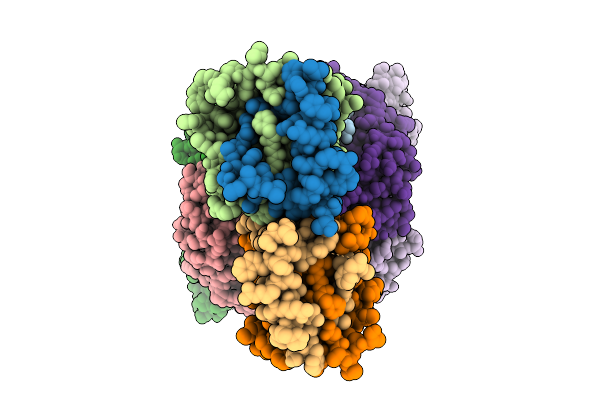 |
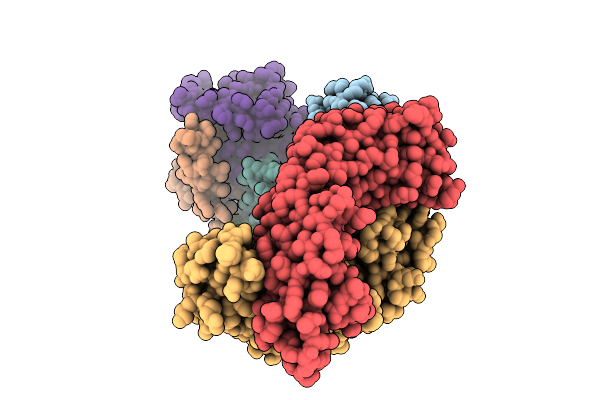
Thermus Thermophilus Mrec-Mred Complex With An Internal Mred Bril Fusion And An Anti-Bril Fab
Organism: Escherichia coli, Thermus thermophilus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN |
 |
Thermus Thermophilus Mrec-Mred Complex With A C-Terminal Mred Bril Fusion And An Anti-Bril Fab
Organism: Escherichia coli, Thermus thermophilus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN |
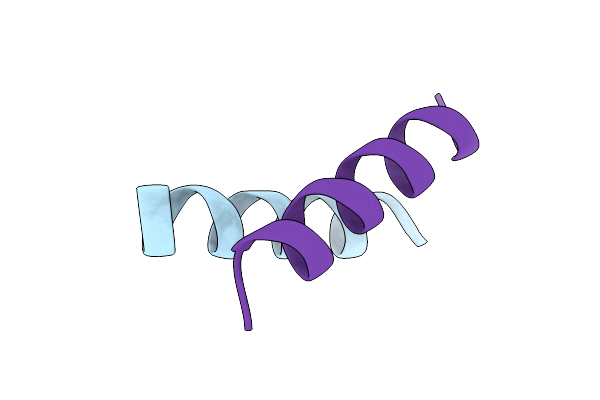 |
Organism: Ranoidea citropa
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2025-10-08 Classification: ANTIMICROBIAL PROTEIN |
 |
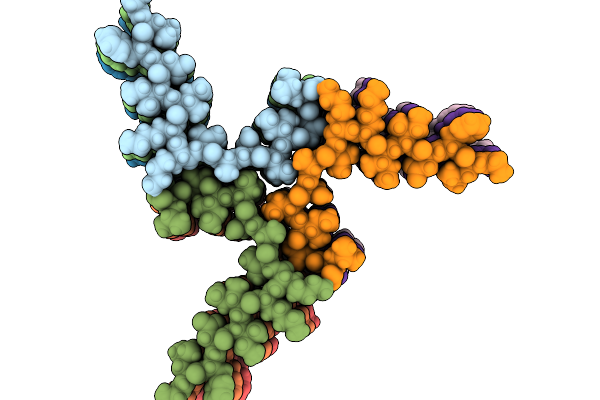
Amyloid Fibril From The Antimicrobial Peptide Citropin 1-3 - Polymorph Nr.3
Organism: Ranoidea citropa
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: ANTIMICROBIAL PROTEIN |
 |
Amyloid Fibril From The Antimicrobial Peptide Citropin 1-3 - Polymorph Nr.2
Organism: Ranoidea citropa
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: ANTIMICROBIAL PROTEIN |
 |
Amyloid Fibril From The Antimicrobial Peptide Citropin 1-3 - Polymorph Nr.1L
Organism: Ranoidea citropa
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: ANTIMICROBIAL PROTEIN |
 |
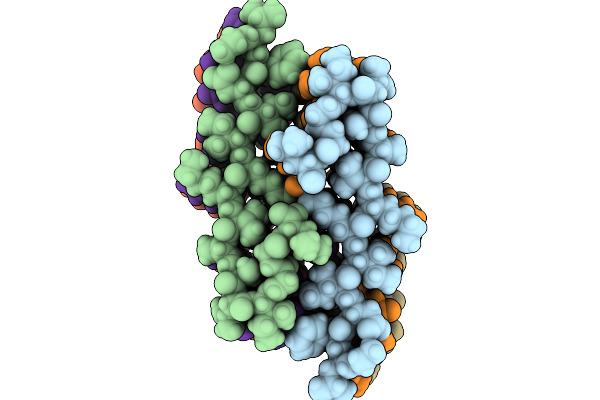
Amyloid Fibril From The Antimicrobial Peptide Citropin 1-3 - Polymorph Nr.3L
Organism: Ranoidea citropa
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: ANTIMICROBIAL PROTEIN |
 |
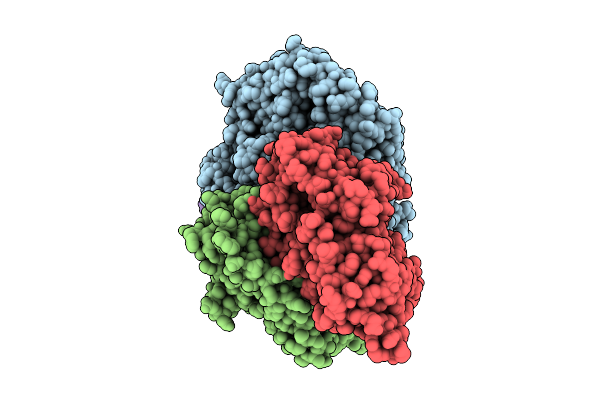
Amyloid Fibril From The Antimicrobial Peptide Citropin 1-3 - Polymorph Nr.2L
Organism: Ranoidea citropa
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: ANTIMICROBIAL PROTEIN |
 |
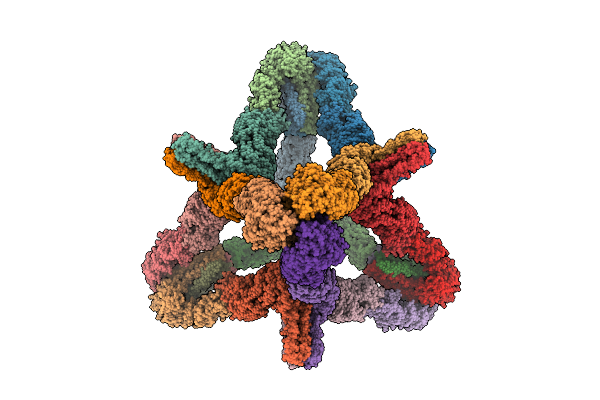
Organism: Rhodospirillum rubrum
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: OXIDOREDUCTASE Ligands: S5Q, CLF, ADP, MG, AF3, SF4 |
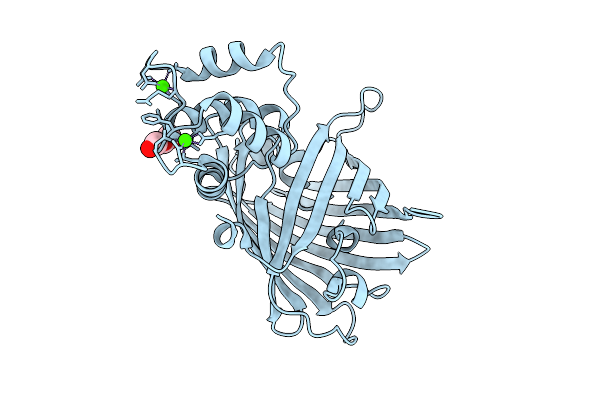 |
Crystal Structure Of The Genetically-Encoded Green Calcium Sensor Icbtnc2 In Its Calcium Bound-State
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-09-17 Classification: FLUORESCENT PROTEIN Ligands: PEG, CA, CL |
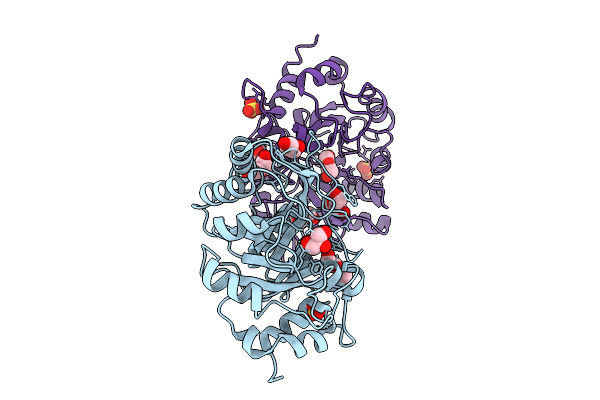 |
Organism: Metschnikowia pulcherrima
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: PEG, GOL, SO4, NAG |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: DE NOVO PROTEIN |
 |
Crystal Structure Of The Wuhan Sars-Cov-2 Spike Rbd (319-541) Complexed With 1P1B10 Nanobody
Organism: Severe acute respiratory syndrome coronavirus 2, Camelus bactrianus
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-07-16 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG, NA |

