Search Count: 24,499
All
Selected
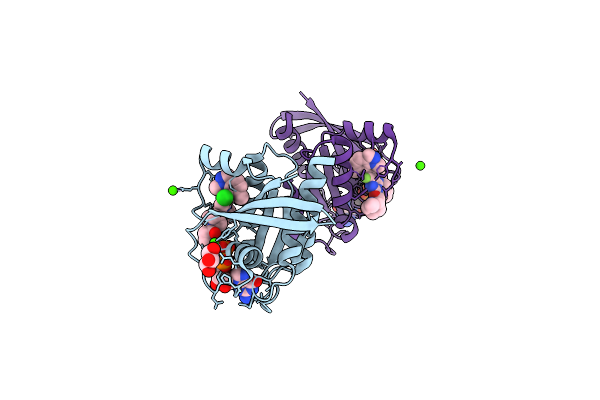 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-15 Classification: ONCOPROTEIN Ligands: CA, A1C6Y, GOL, GDP |
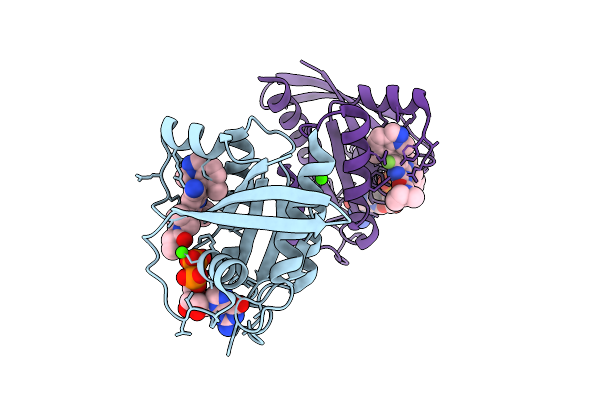 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-04-15 Classification: ONCOPROTEIN Ligands: GDP, A1C60, CA |
 |
Crystal Structure Of Putative L-Amino Acid N-Acyltransferase Mnat From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: EDO |
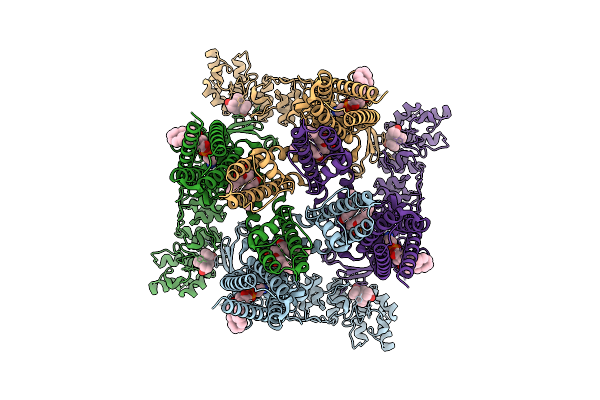 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: MEMBRANE PROTEIN Ligands: PEX, A1BMB |
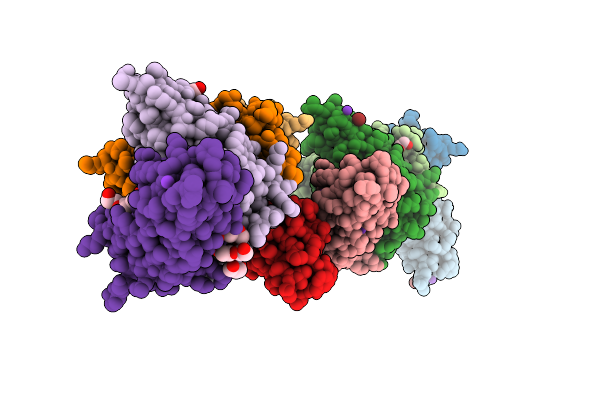 |
Crystal Structure Of The Third Immunoglobulin-Like Domain Of Human Muscle-Specific Kinase
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-15 Classification: SIGNALING PROTEIN Ligands: BR, K, PEG, PG4, PGE |
 |
Long-Wavelength Sad Crystal Structure Of The Third Immunoglobulin-Like Domain Of Human Muscle-Specific Kinase (Musk)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-15 Classification: SIGNALING PROTEIN Ligands: K, BR, PEG, PGE |
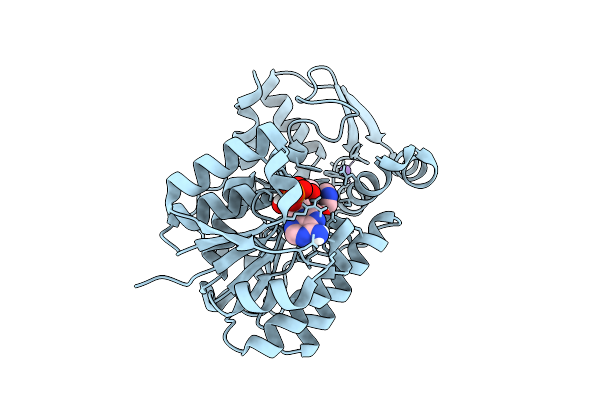 |
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-04-15 Classification: PLANT PROTEIN Ligands: NAP, NCA, NA |
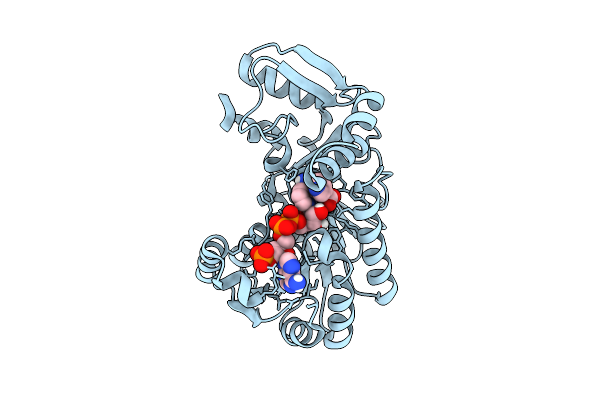 |
Crystal Structure Of An Nta622L Variant In Complex With Nadp+ And Nicotinic Acid
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-04-15 Classification: PLANT PROTEIN Ligands: NIO, NAP |
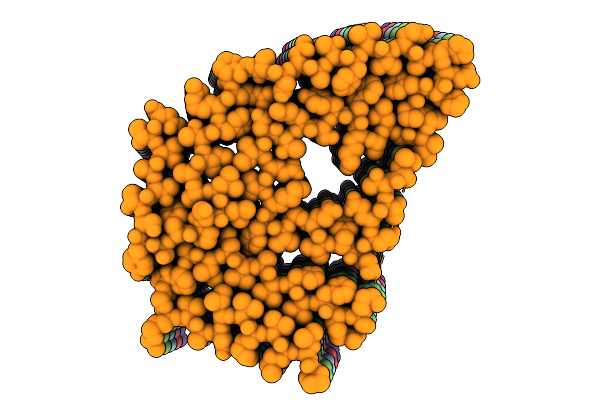 |
Cryo-Em Structure Of Hepatic Amyloid Fibril From A Variant Attrv122Delta, Single Filament Morphology
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
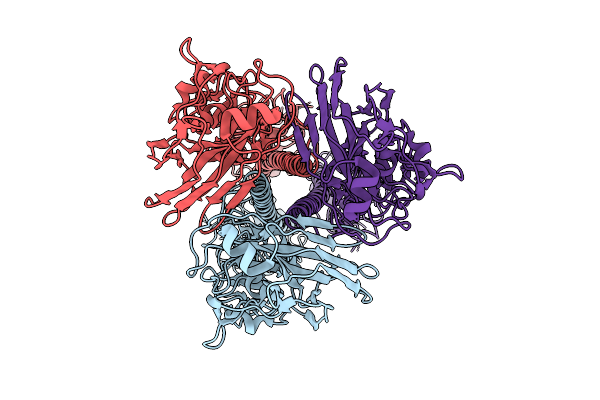 |
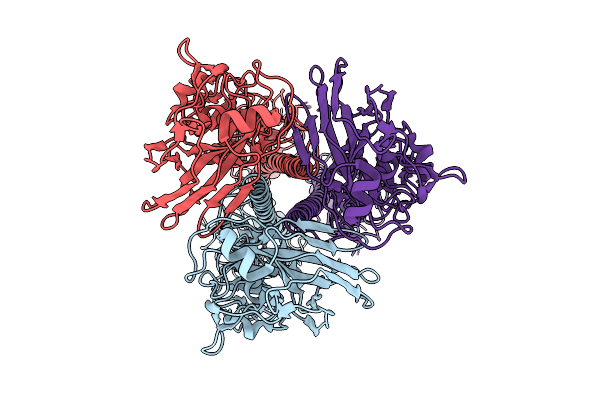
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.39 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.16 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
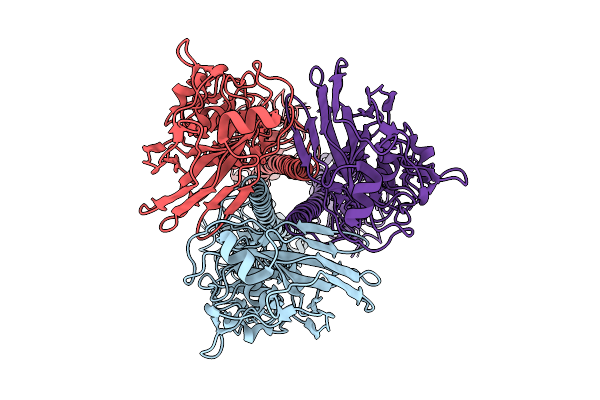
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.67 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
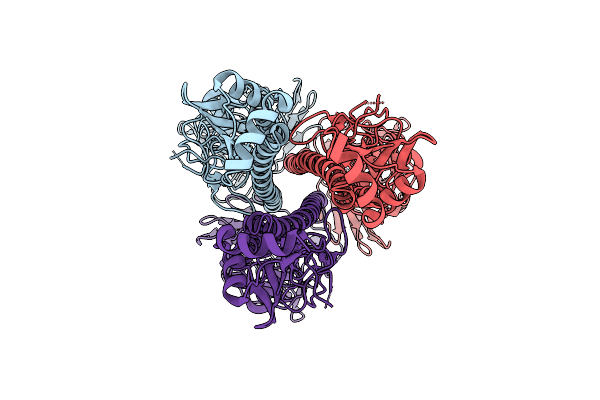
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.47 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
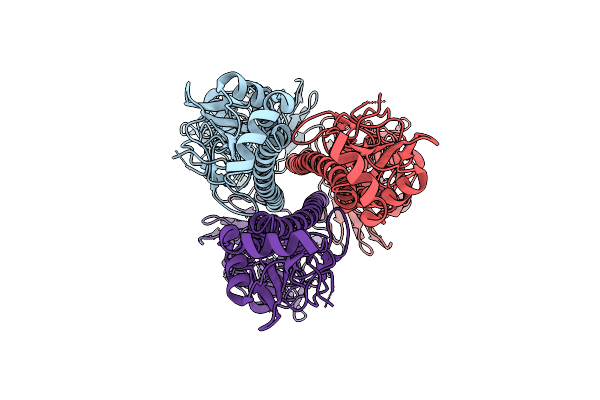
Influenza A Virus Hemagglutinin (A/California/04/2009 H1N1), E47K Ha2 Stabilizing Mutation (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
Influenza A Virus Hemagglutinin (A/California/04/2009 H1N1), E47K Ha2 Stabilizing Mutation (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.64 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
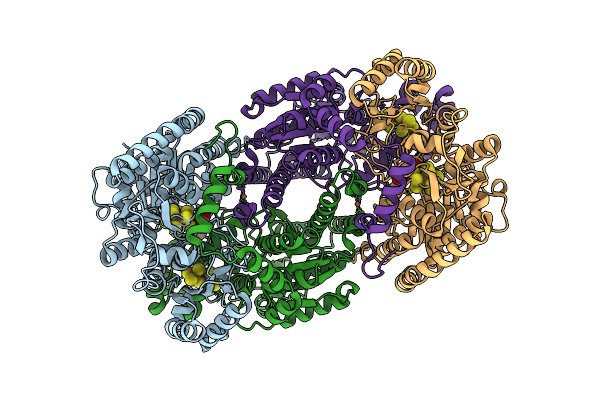 |
Azotobacter Vinelandii Mofep (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, CLF, FE |
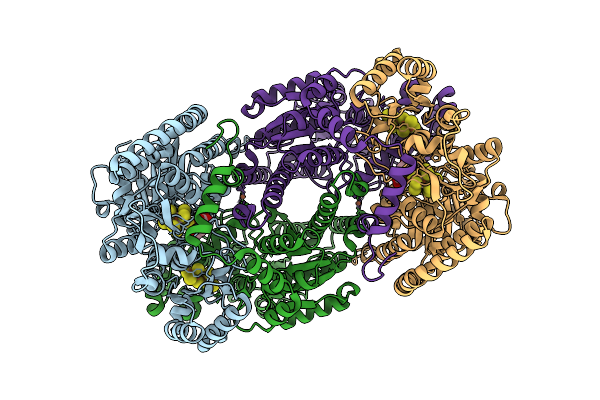 |
Azotobacter Vinelandii Mofep (C2 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.46 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
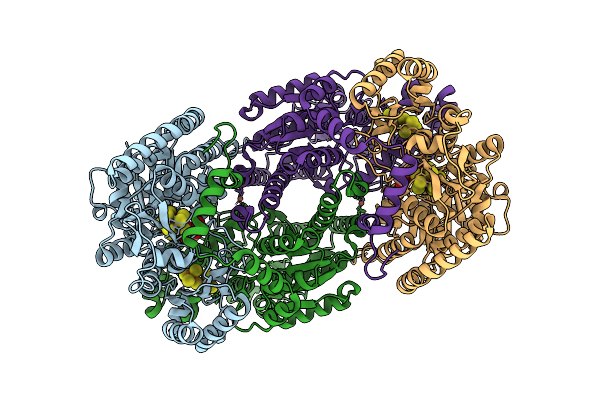 |
Azotobacter Vinelandii Mofep (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.16 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
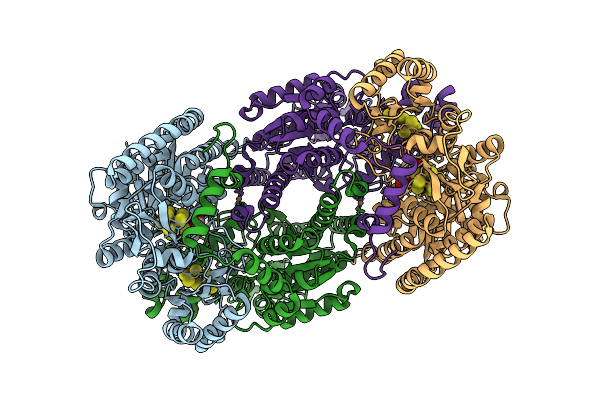 |
Azotobacter Vinelandii Mofep (C2 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
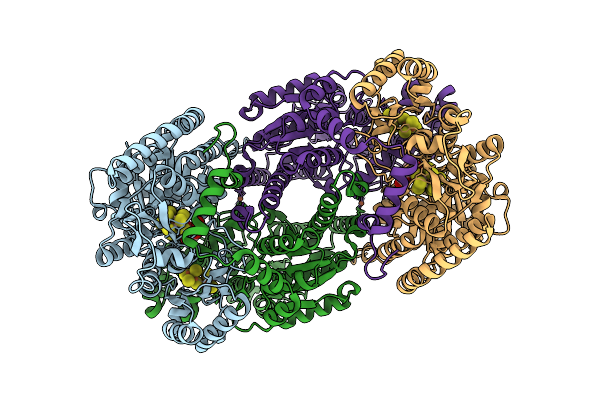 |
Azotobacter Vinelandii Mofep (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |

