Search Count: 46,761
All
Selected
 |
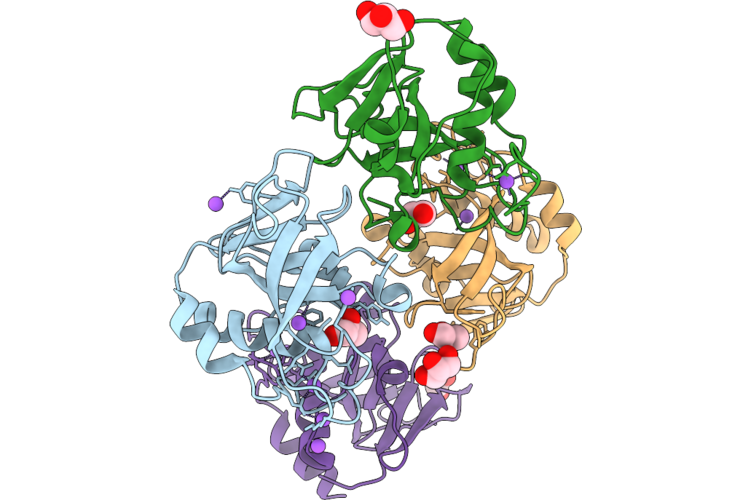
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: UDP, NOV |
 |
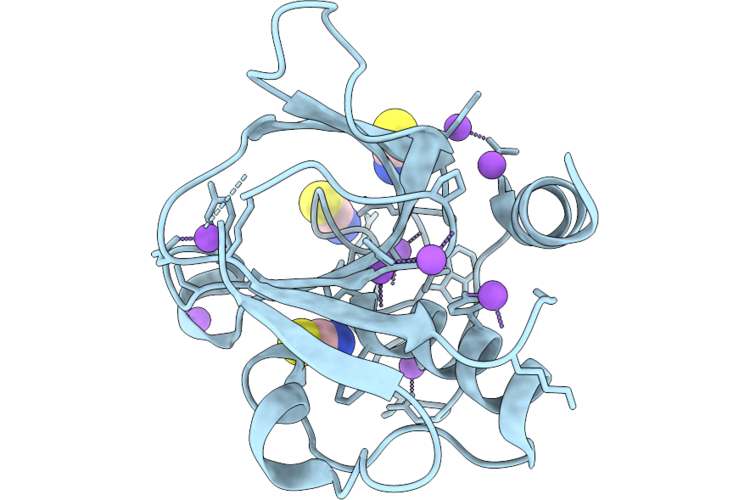
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2026-05-06 Classification: HYDROLASE |
 |
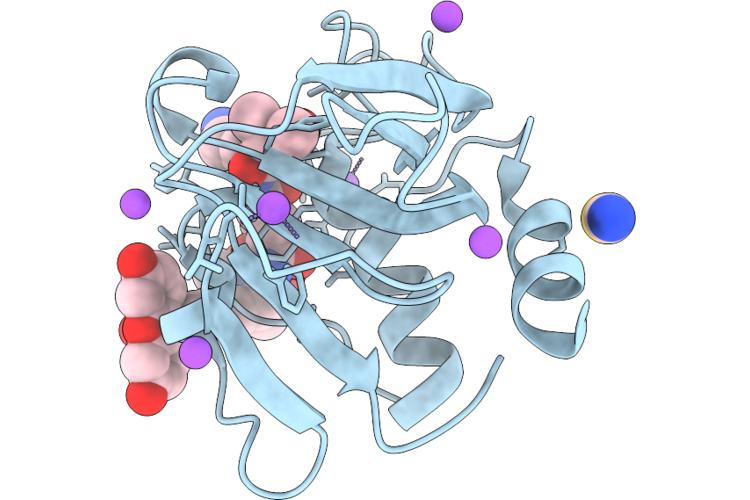
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-05-06 Classification: HYDROLASE |
 |
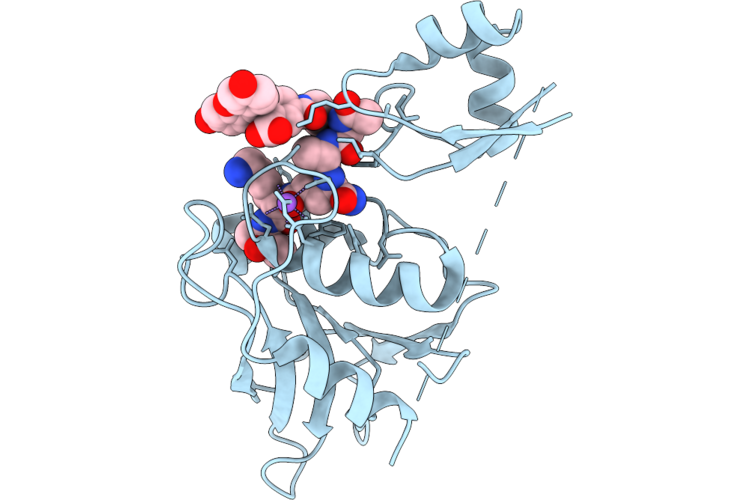
Chap Domain Of Staphylococcus Aureus-Specific Lysin L1 Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA, SCN |
 |
Staphylococcus Aureus-Specific Lysin L1-3 (Lysm-Chap) Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA |
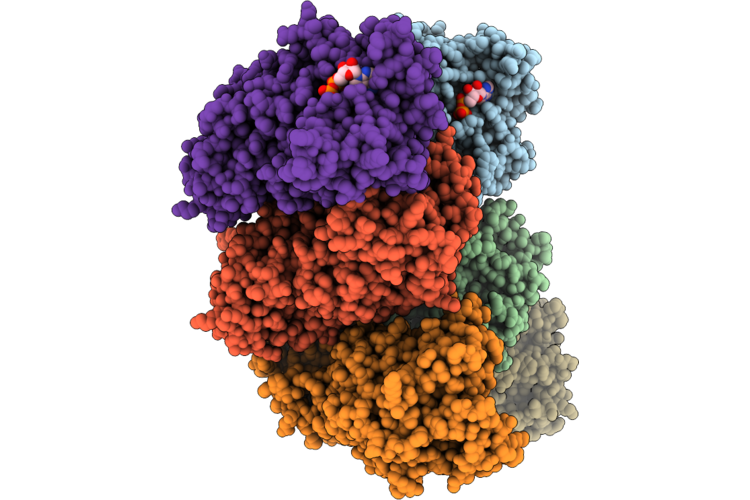 |
Crystal Structure Of Hera Like Helicae Yjgr With Ntpase Fold From Escherchia Coli
Organism: Escherichia coli str. k-12 substr. mg1655
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-05-06 Classification: GENE REGULATION Ligands: ATP, MG |
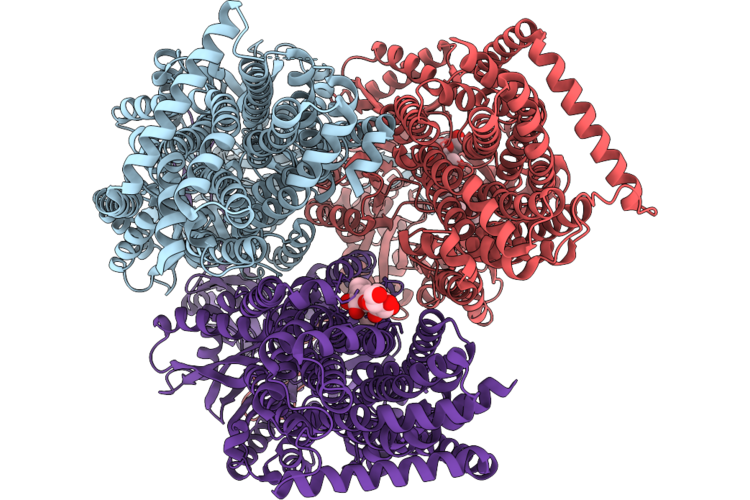 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: CA7 |
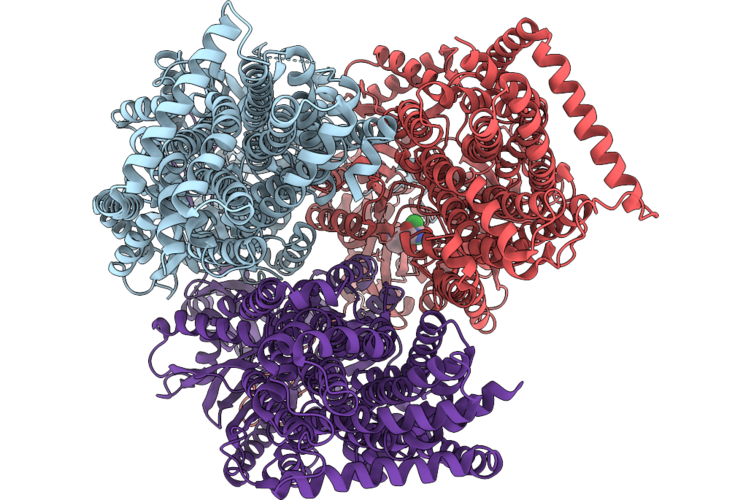 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: CLM |
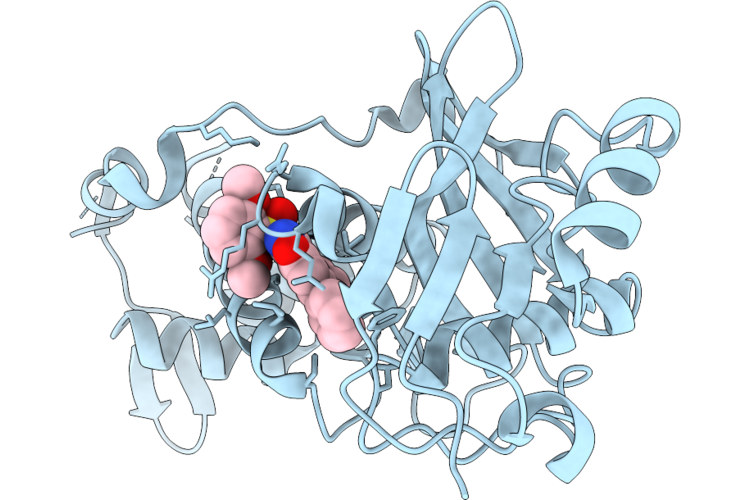 |
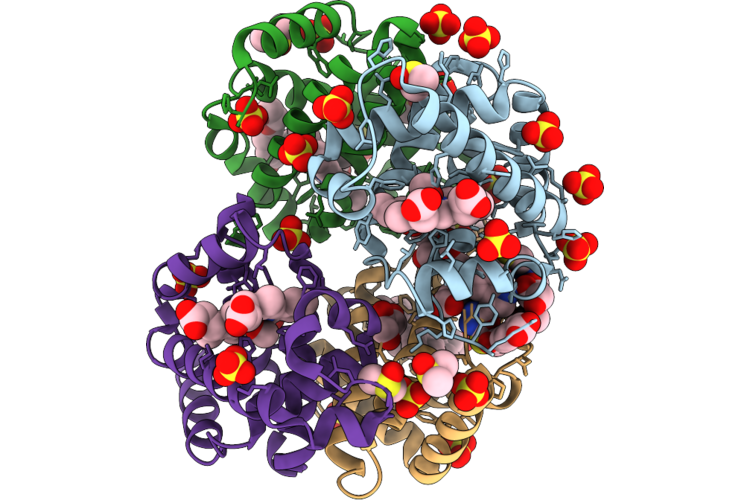
Crystal Structure Of Myst Histone Acetyltransferase Kat6A In Complex With Inhibitor Compound 9
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2026-05-06 Classification: TRANSCRIPTION Ligands: A1MGC |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: HEM, GOL, A1ITR, DMS, CMO, ACT, SO4, PGE |
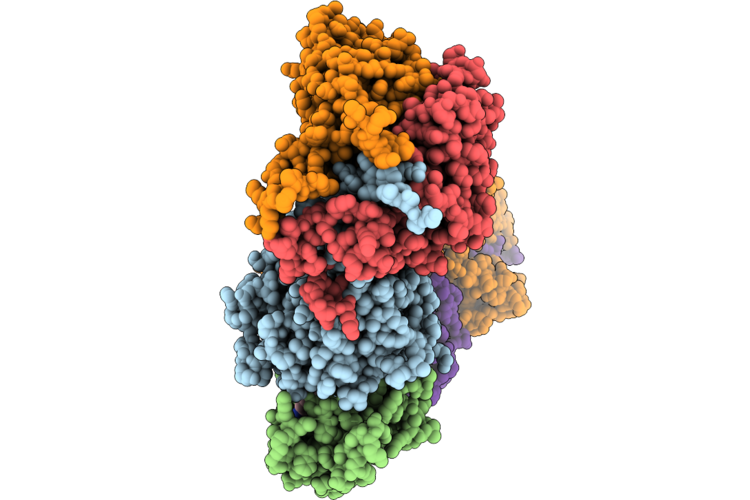 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10088
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
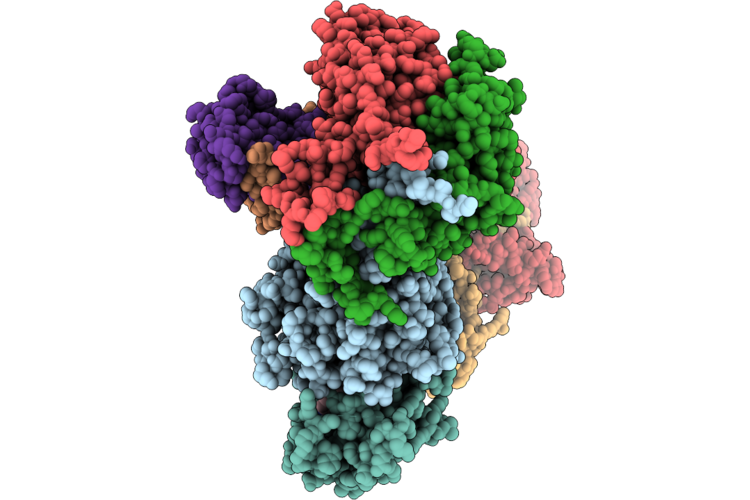 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2, Unc10088 And Bach1
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
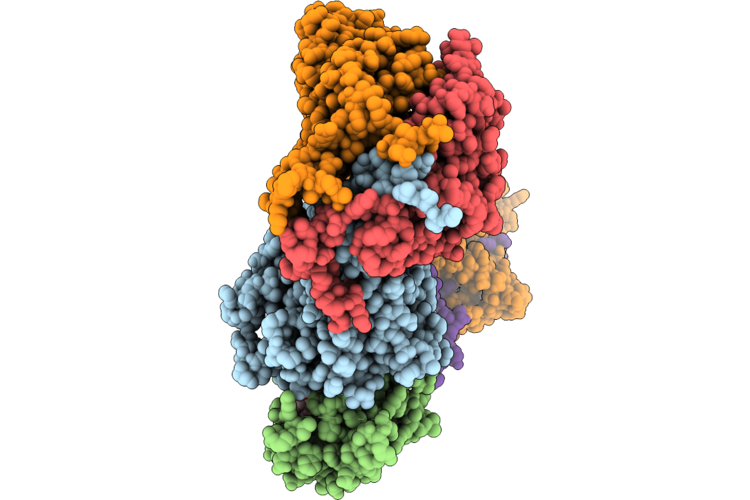 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10415667
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J21 |
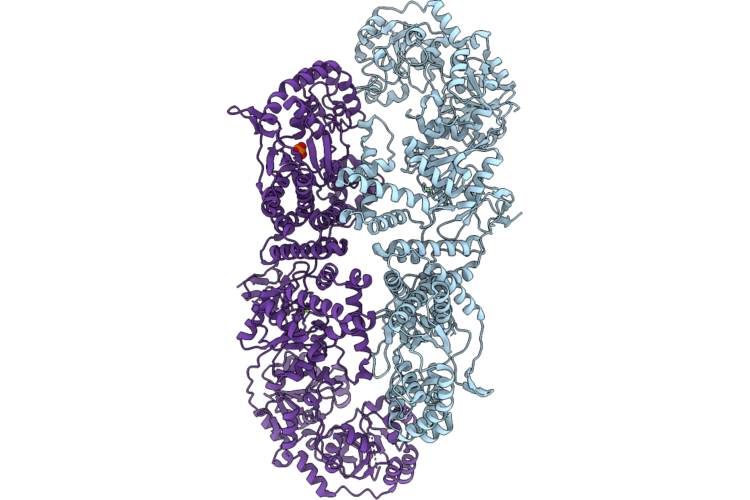 |
Organism: Haemophilus influenzae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: BIOSYNTHETIC PROTEIN Ligands: MG, PO4 |
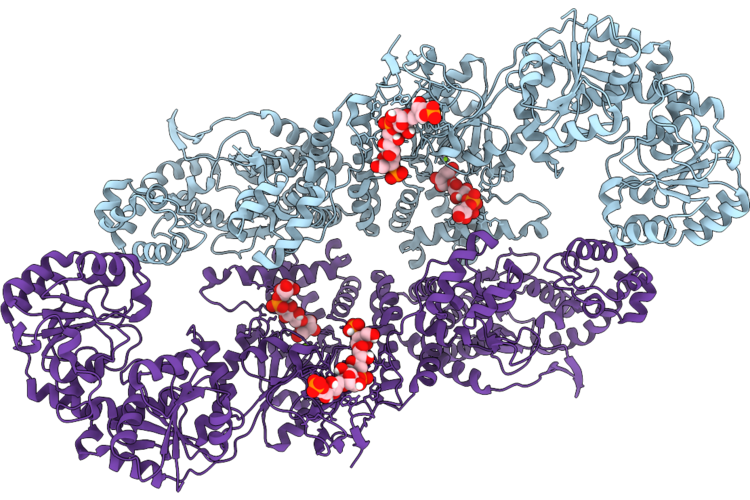 |
Haemophilus Influenzae Serotype B Capsule Polymerase Bcs3 In Complex With Acceptor Polymer
Organism: Haemophilus influenzae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: BIOSYNTHETIC PROTEIN Ligands: MG, KOF |
 |
Aetf, A Single-Component Flavin-Dependent Tryptophan Halogenase, In Complex With Tryptoline
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-05-06 Classification: FLAVOPROTEIN Ligands: FAD, WYH, PEG, CL |
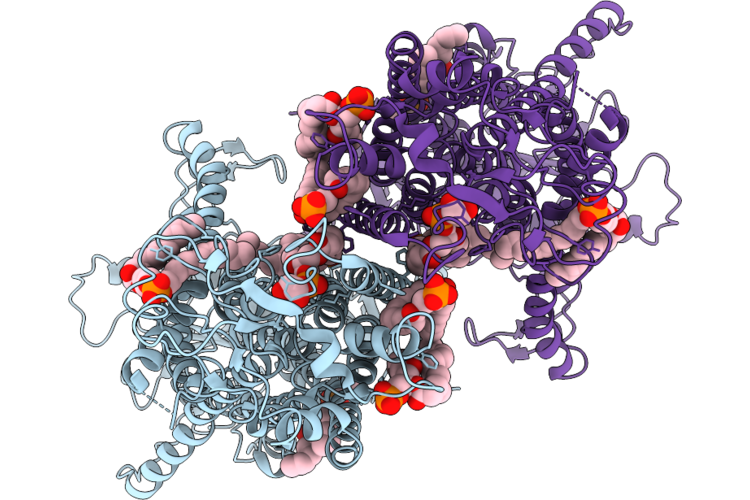 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIPID TRANSPORT Ligands: PGW |
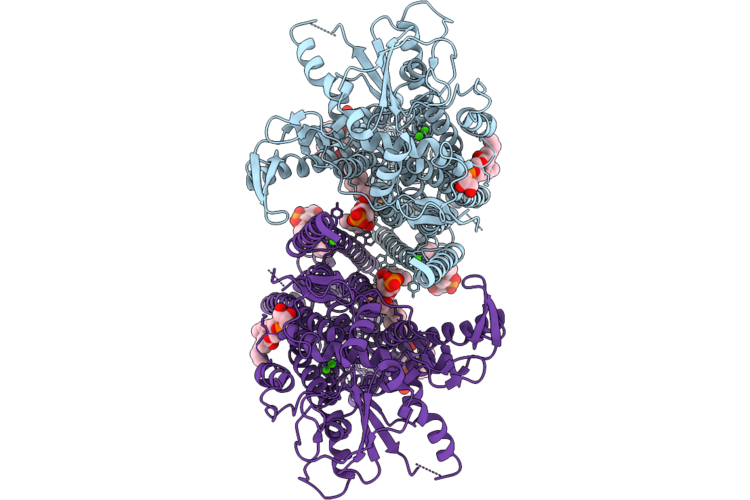 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIPID TRANSPORT Ligands: CA, PGW |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIPID TRANSPORT Ligands: CA |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIPID TRANSPORT Ligands: CA, PGW |

