Search Count: 558
All
Selected
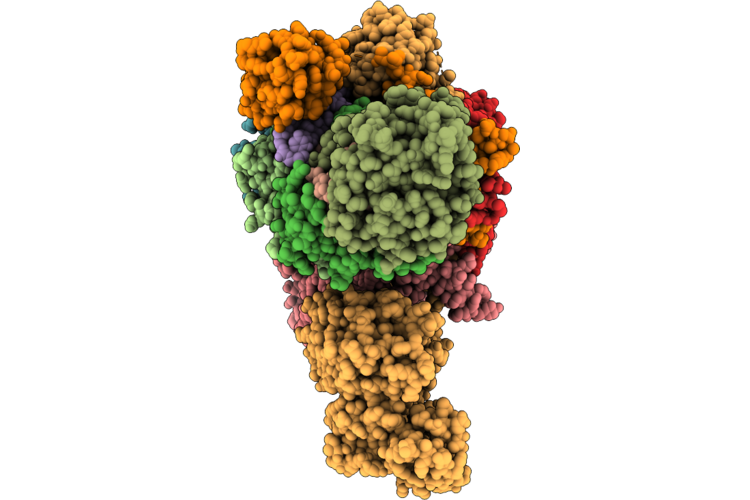 |
Cryo-Em Structure Of Type Vii Crispr-Cas Complex At The Target Engagement State
Organism: Metagenome
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: ANTIVIRAL PROTEIN/RNA Ligands: ZN |
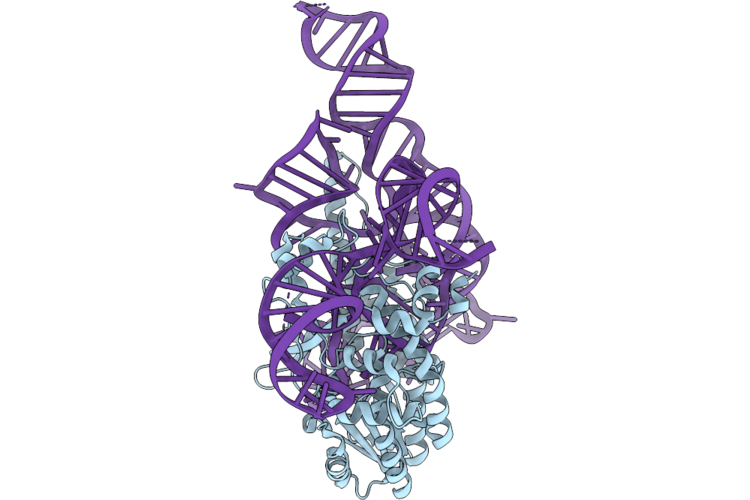 |
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Resolution:3.49 Å Release Date: 2026-04-22 Classification: ANTIVIRAL PROTEIN Ligands: MG |
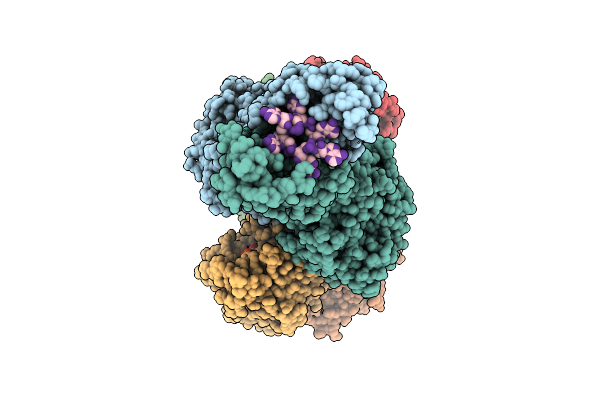 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 3
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, MG, ZN |
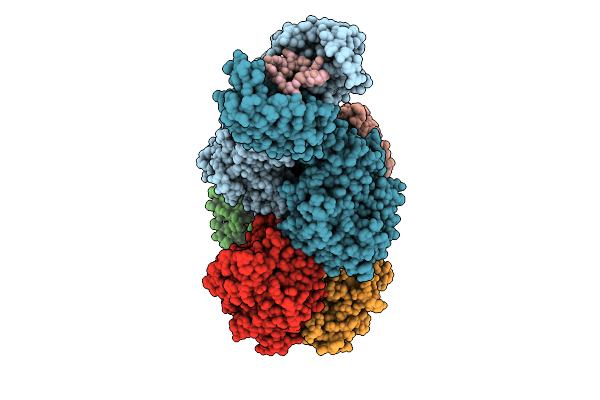 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 4
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
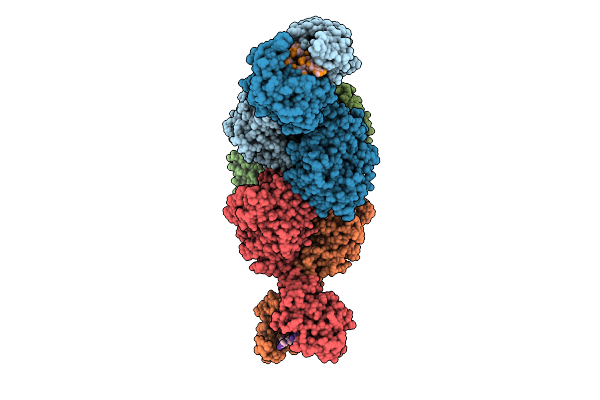 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 5
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
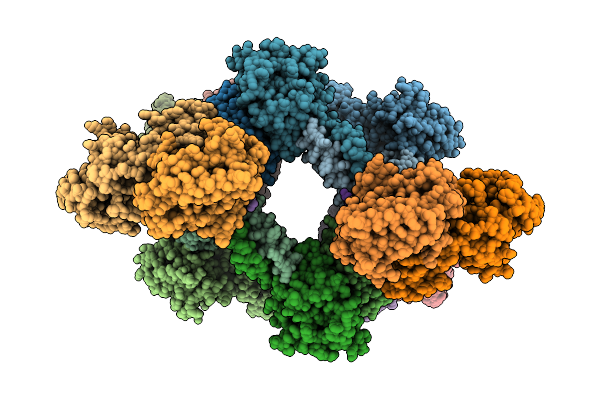 |
Organism: Enterococcus faecalis
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN Ligands: CA |
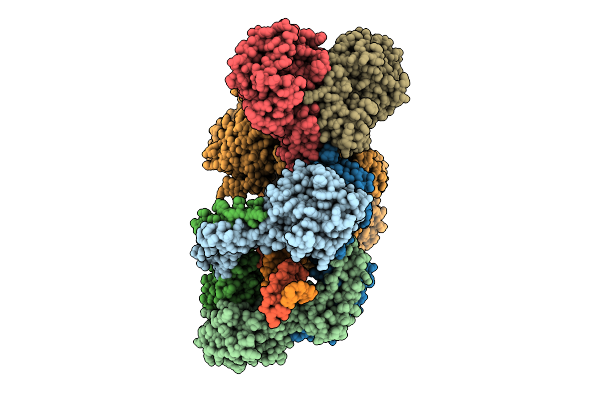 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
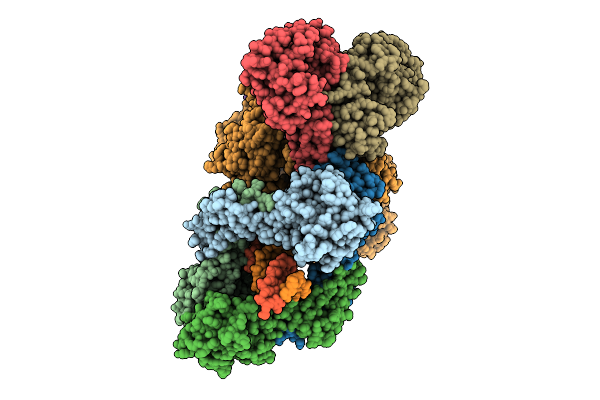 |
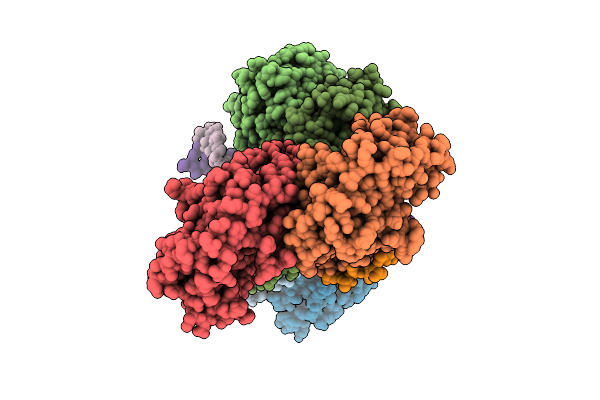
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.76 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA Ligands: CA |
 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVAL PROTEIN/RNA/DNA |
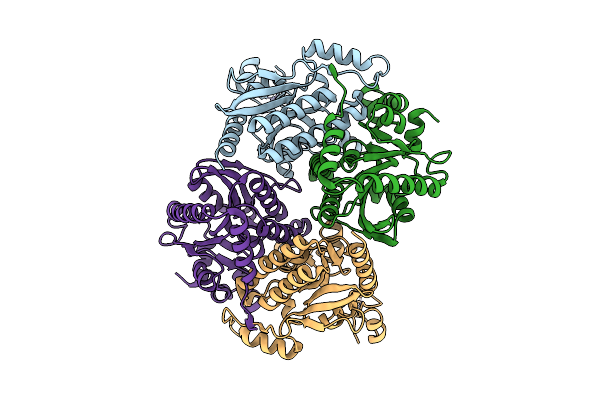 |
Crystal Structure Of The Carboxyltransferase Subunit Of Acetyl-Coa Carboxylase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2026-03-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN |
 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN |
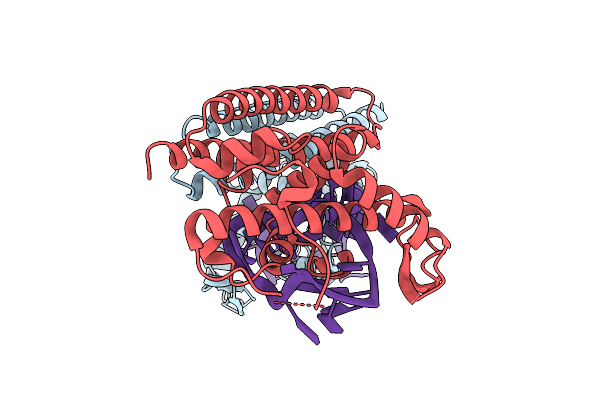 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN |
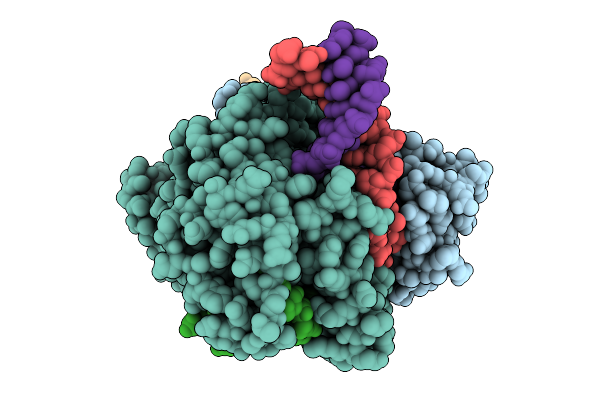 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN Ligands: MG |
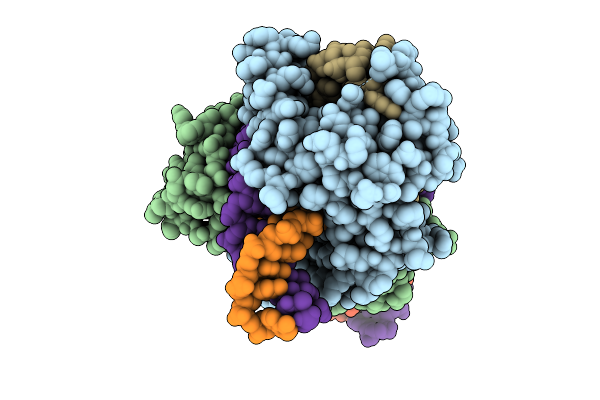 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN Ligands: MG |
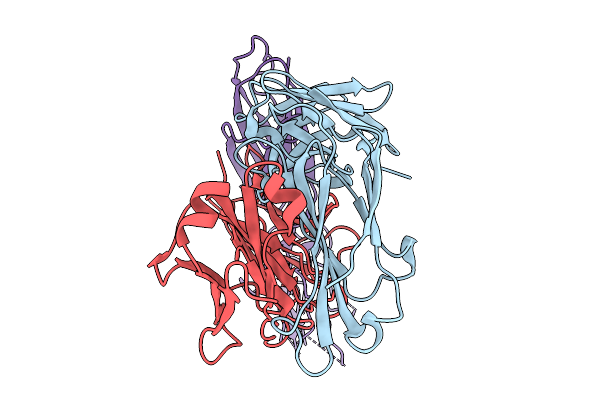 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: ANTITUMOR PROTEIN/IMMUNE SYSTEM |
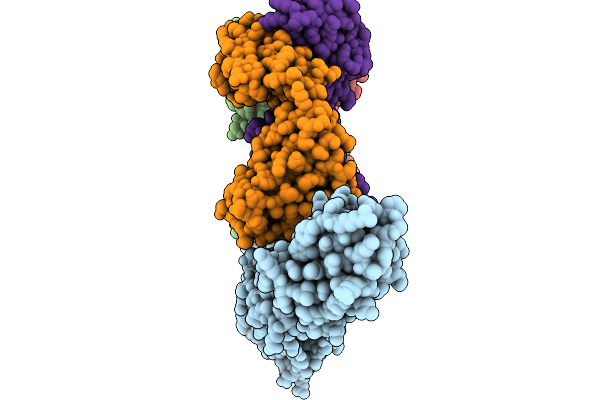 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: ANTITUMOR PROTEIN/IMMUNE SYSTEM |
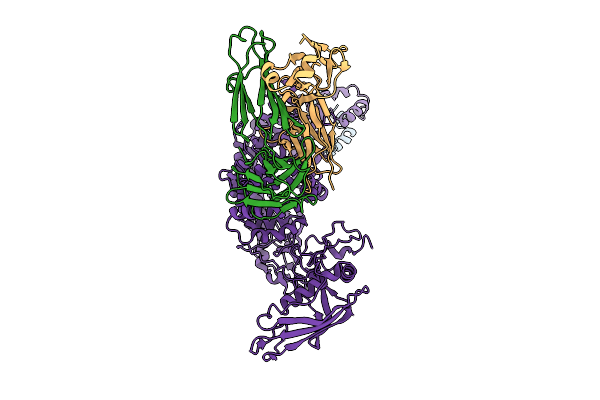 |
P116 From Mycoplasma Pneumoniae In Complex With Mild Growth Suppressor Monoclonal Antibody
Organism: Mus musculus, Mycoplasmoides pneumoniae m129
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: PROTEIN TRANSPORT |
 |
Crystal Structure Of Nodd-Ebd (Effector Binding Domain) In Complex With Hesperetin From Rhizobium Leguminosarum Bv. Vicae 3841
Organism: Rhizobium leguminosarum
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-12-24 Classification: TRANSCRIPTION Ligands: GOL, 6JP |

