Search Count: 774
All
Selected
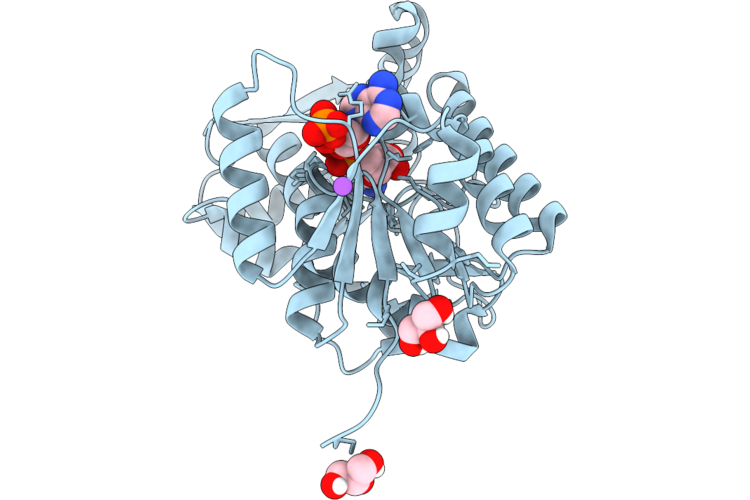 |
Organism: Illicium verum
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2026-05-20 Classification: PLANT PROTEIN Ligands: GOL, NDP, NA |
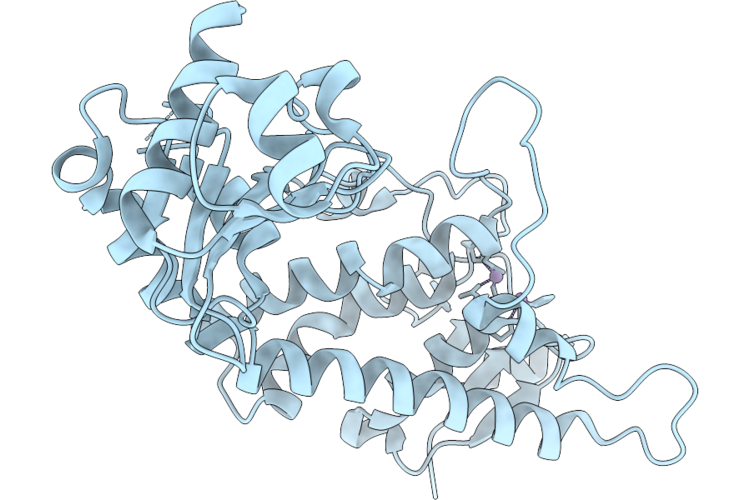 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MN |
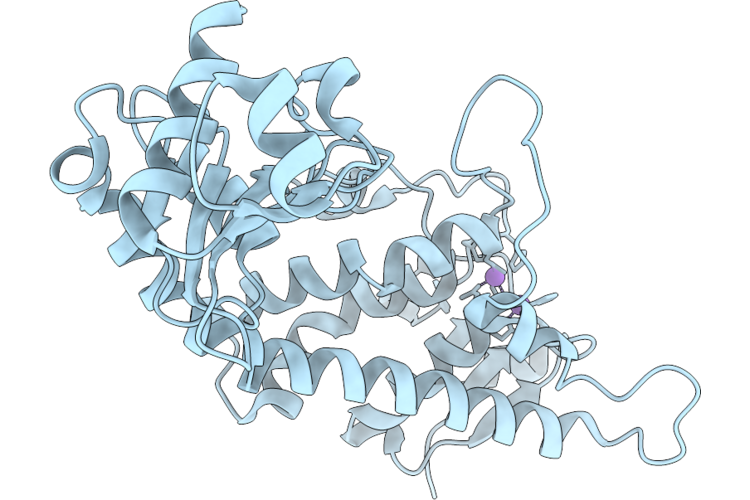 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-22 Classification: HYDROLASE |
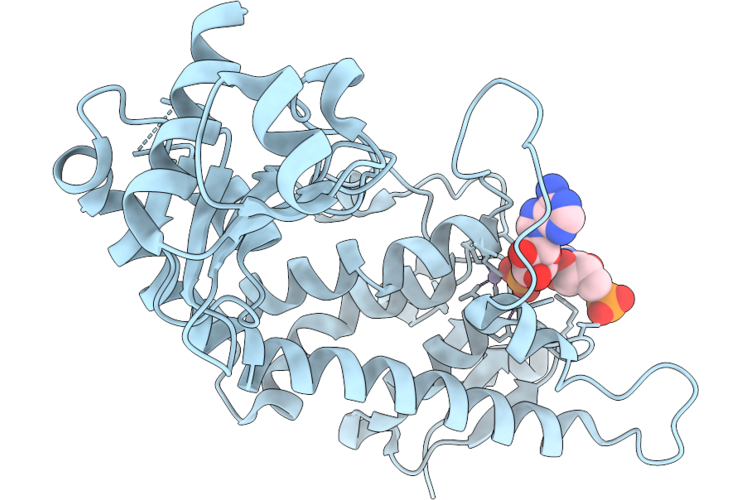 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: AMP, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: 5GP, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: U5P, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: C5P, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: DNR, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: TMP, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: AMP, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: AMP, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: AMP, MN |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-22 Classification: HYDROLASE/DNA Ligands: MN |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: HYDROLASE/DNA Ligands: MN |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-22 Classification: HYDROLASE/RNA Ligands: MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: D5M, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: DGP, MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: AMP, MN |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-04-22 Classification: HYDROLASE/RNA Ligands: MN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: AMP, MN |

