Search Count: 368
All
Selected
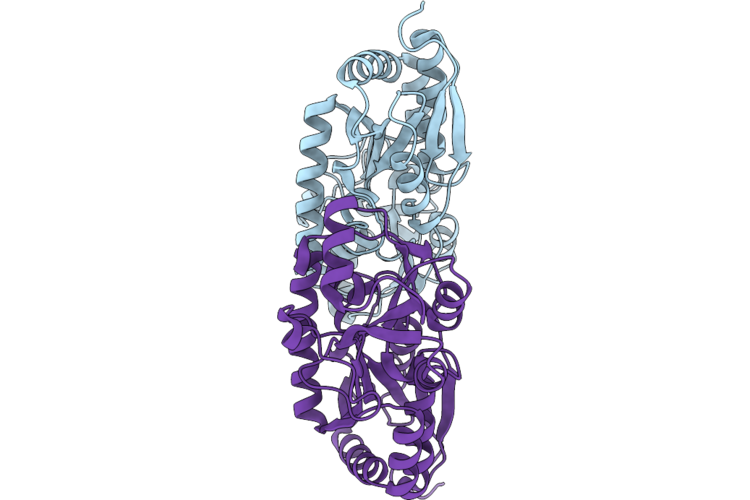 |
Crystal Structure Of A Tctc Solute Binding Protein From Vibrio Sp. C42, No Ligand
Organism: Vibrio atlanticus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING |
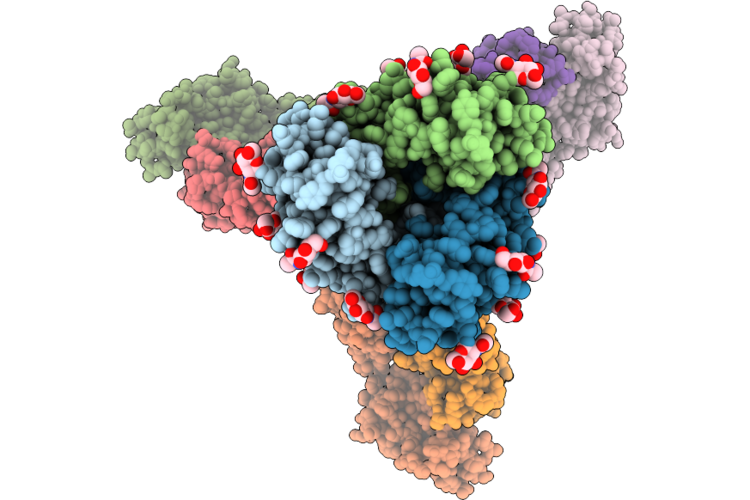 |
Cryo-Em Structure Of Sars-Cov-2 Wide-Type S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab (Focused Refinement Of The S2-76E1)
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
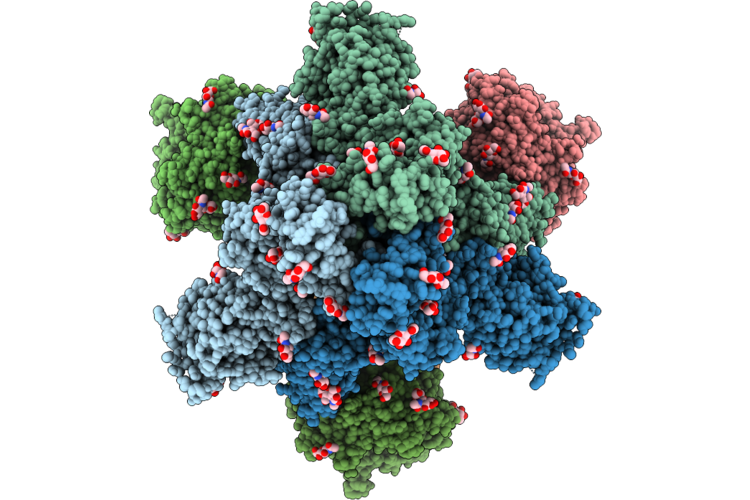 |
Cryo-Em Structure Of Sars-Cov-2 Wide-Type S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
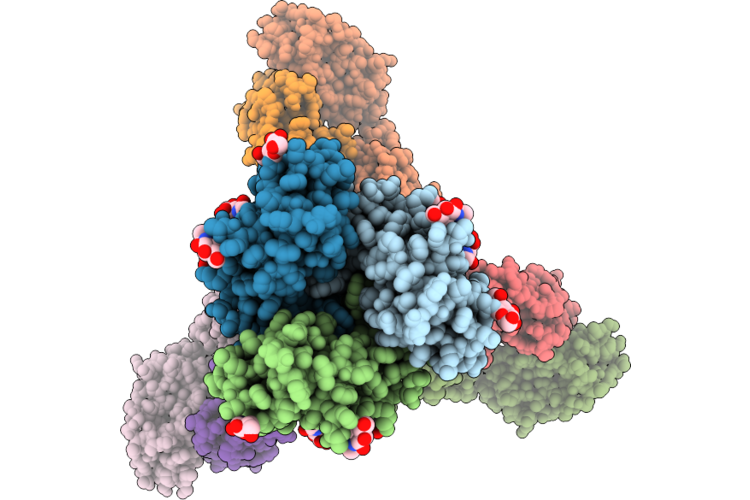 |
Cryo-Em Structure Of Sars-Cov-2 Xbb.1.5 S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab (Focused Refinement Of The S2-76E1)
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
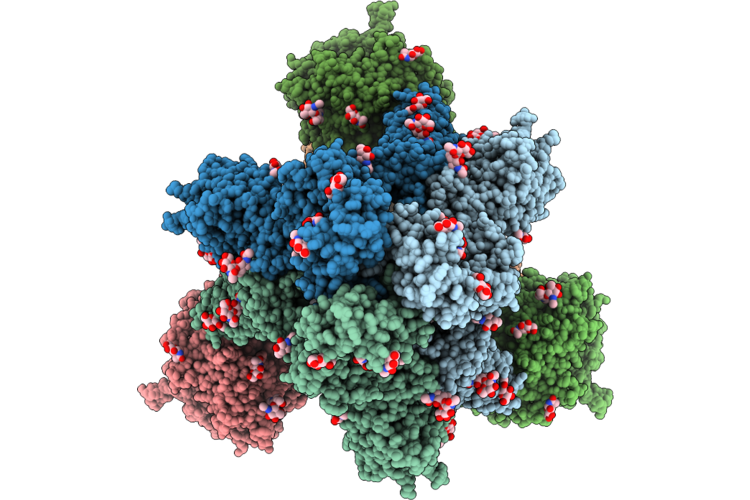 |
Cryo-Em Structure Of Sars-Cov-2 Xbb.1.5 S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
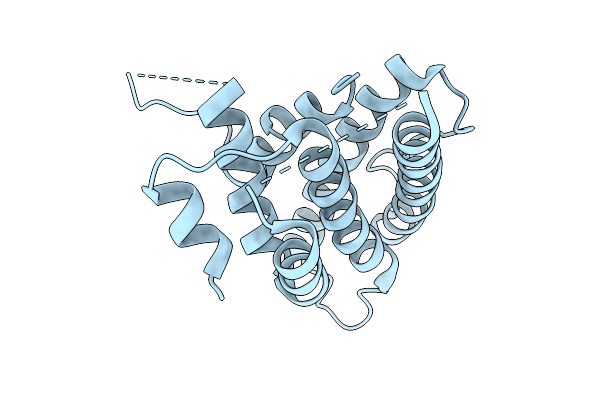 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-15 Classification: PROTEIN BINDING |
 |
Organism: Aquimarina sp. 2-a2
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-18 Classification: SUGAR BINDING PROTEIN |
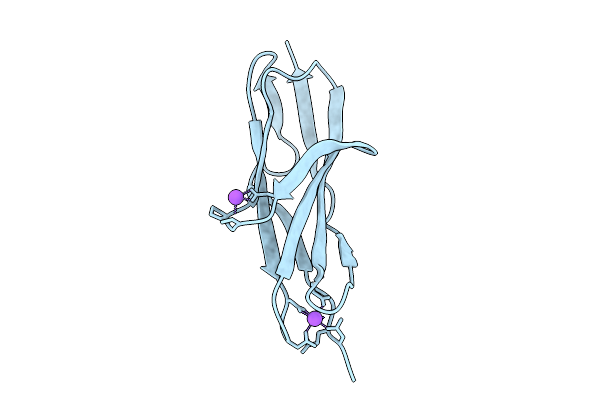 |
Organism: Aquimarina sp. 2-a2
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: SUGAR BINDING PROTEIN |
 |
Gii.23/24/25 Noroviruses Recognize Glycans Via A Conventional Glycan-Binding Site
Organism: Norovirus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-04 Classification: VIRAL PROTEIN |
 |
Gii.23/24/25 Noroviruses Recognize Glycans Via A Conventional Glycan-Binding Site
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN |
 |
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN |
 |
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN Ligands: P |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-11 Classification: NUCLEAR PROTEIN |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-02-11 Classification: NUCLEAR PROTEIN |
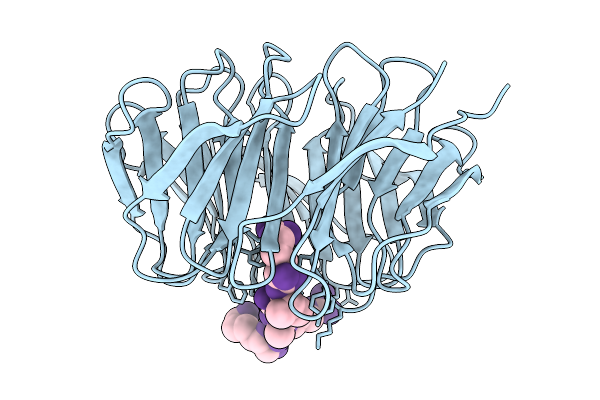 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2026-02-11 Classification: NUCLEAR PROTEIN |
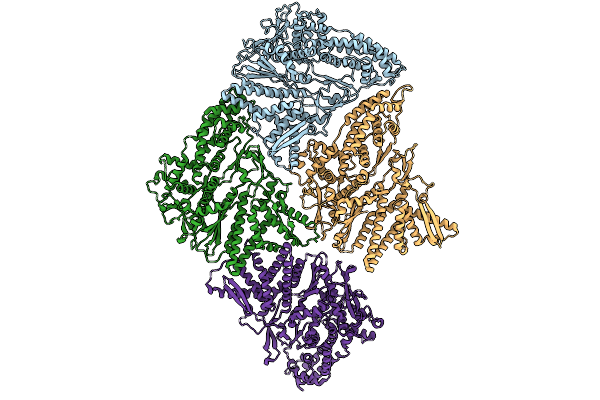 |
Inner Layer Protein P1 Chains In Transcribing Particles Of Bacteriophage Phi6
Organism: Cystovirus phi6
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRAL PROTEIN |
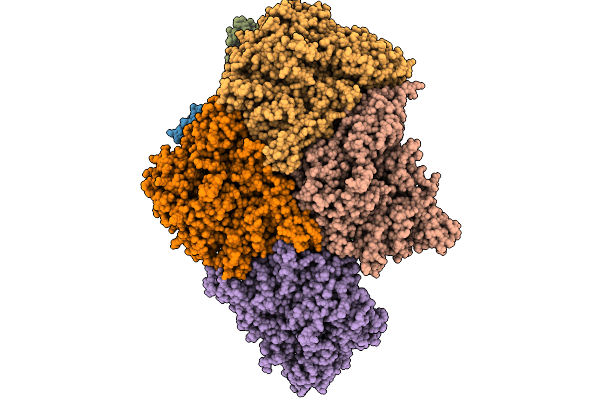 |
Extended And Wrapped Protein P7 Dimers Of Dimers, The P1 Layer And The Rna-Dependent Rna Polymerase P2 In Transcribing Particles Of Bacteriophage Phi6
Organism: Synthetic construct, Cystovirus phi6
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRAL PROTEIN |
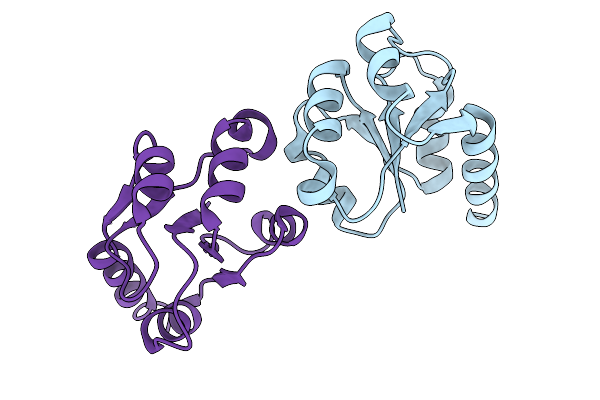 |
Organism: Cystovirus phi6
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRUS |
 |
Asymmetric Unit From Transcribing Double-Layered Particle Of Bacteriophage Phi6
Organism: Cystovirus phi6
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRUS |
 |
Asymmetric Unit From Transcribing Single-Layered Particle Of Bacteriophage Phi6
Organism: Cystovirus phi6
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRUS |

