Search Count: 502
All
Selected
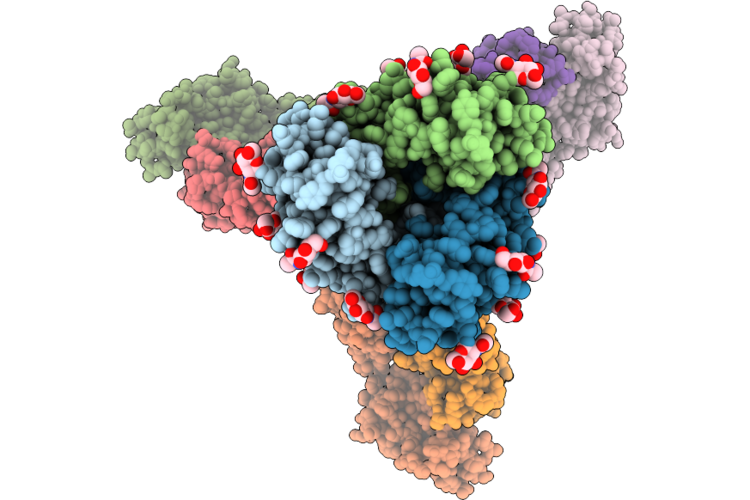 |
Cryo-Em Structure Of Sars-Cov-2 Wide-Type S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab (Focused Refinement Of The S2-76E1)
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
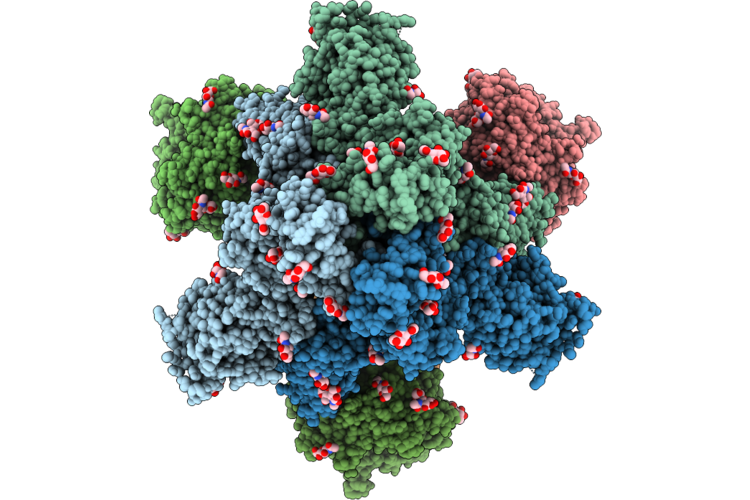 |
Cryo-Em Structure Of Sars-Cov-2 Wide-Type S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
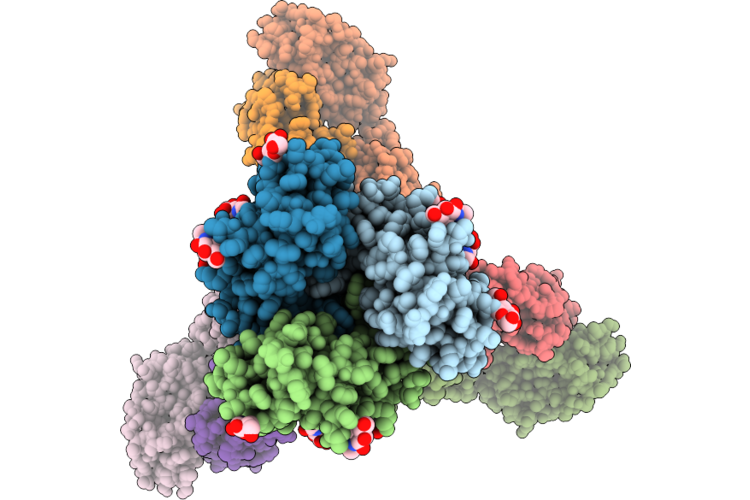 |
Cryo-Em Structure Of Sars-Cov-2 Xbb.1.5 S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab (Focused Refinement Of The S2-76E1)
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
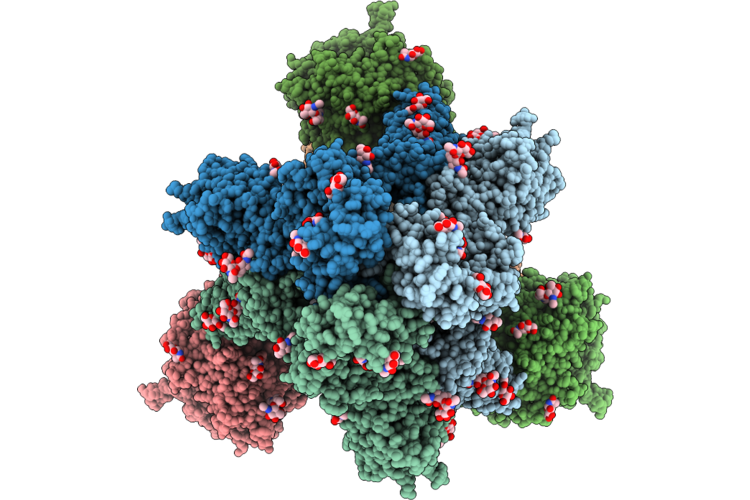 |
Cryo-Em Structure Of Sars-Cov-2 Xbb.1.5 S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
 |
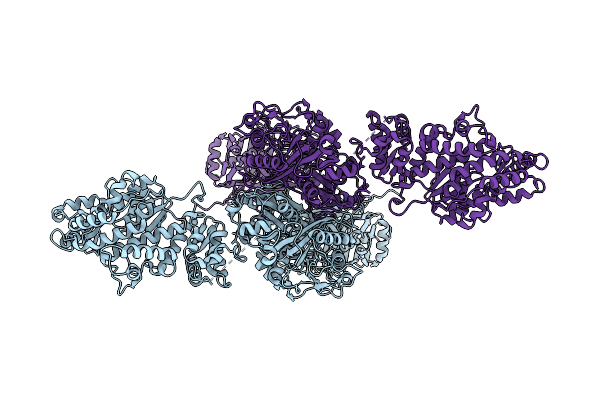
Organism: Escherichia phage t4
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN |
 |
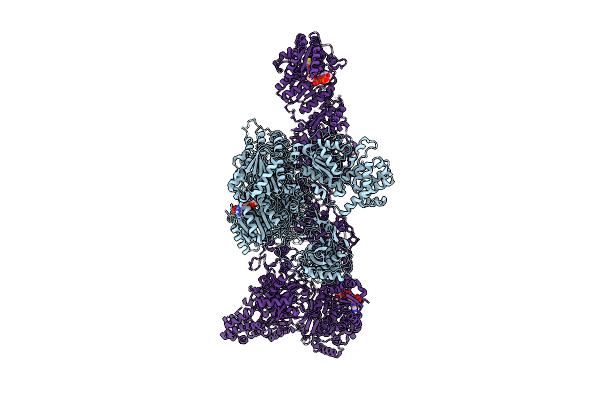
Organism: Aspergillus fumigatus (strain atcc mya-4609 / cbs 101355 / fgsc a1100 / af293)
Method: ELECTRON MICROSCOPY Resolution:2.99 Å Release Date: 2026-04-15 Classification: BIOSYNTHETIC PROTEIN |
 |
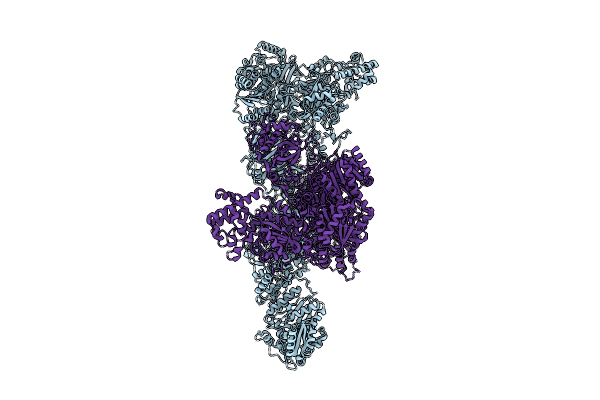
Organism: Aspergillus fumigatus af293
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2026-04-08 Classification: BIOSYNTHETIC PROTEIN Ligands: MLC |
 |
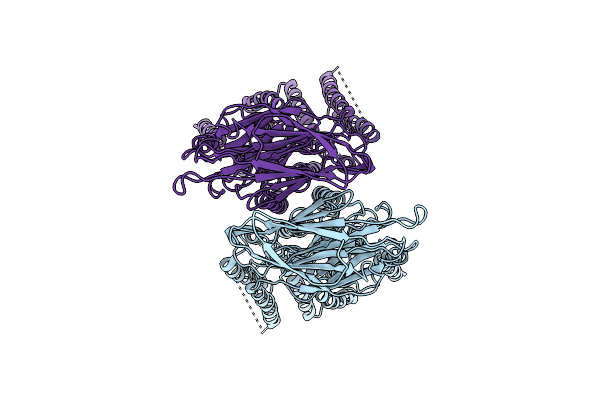
Organism: Aspergillus fumigatus af293
Method: ELECTRON MICROSCOPY Resolution:3.36 Å Release Date: 2026-04-08 Classification: BIOSYNTHETIC PROTEIN Ligands: COZ, NDP |
 |
Organism: Aspergillus fumigatus af293
Method: ELECTRON MICROSCOPY Resolution:3.39 Å Release Date: 2026-04-08 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.34 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: A1ETX, K |
 |
Organism: Homo sapiens, Apis mellifera
Method: ELECTRON MICROSCOPY Resolution:3.23 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: K, POV |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.35 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: POV, CLR, CA, A1B92, K |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: K, Y7Z |
 |
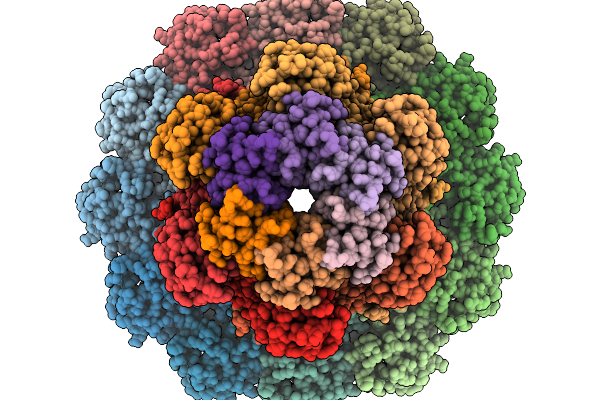
Structure Of Bacteriophage T4 Neck Protein Gp13 And Gp14 Assembled In Vitro In C6 Symmetry
Organism: Escherichia phage t4
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
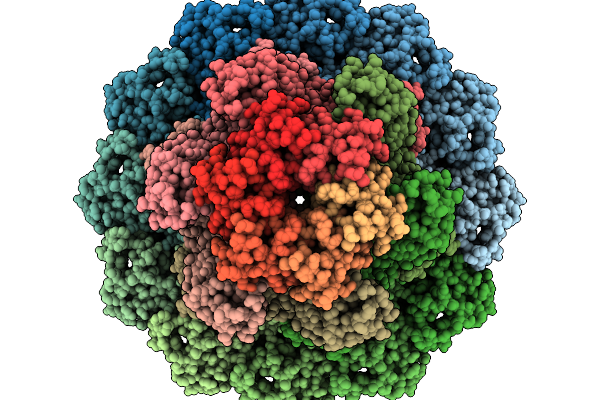
Structure Of Bacteriophage T4 Neck Protein Gp13 And Gp14 And Hfq Assembled In Vitro In C6 Symmetry
Organism: Escherichia phage t4
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
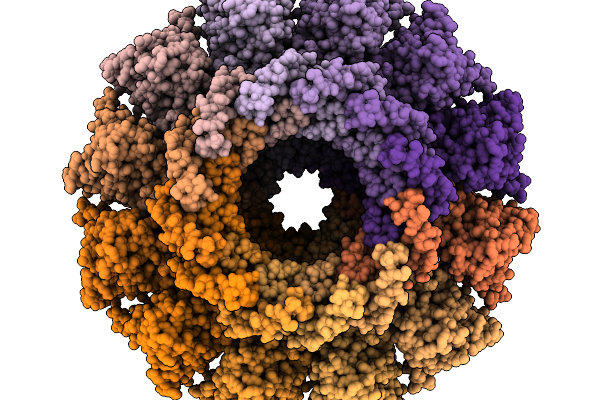
Structure Of Bacteriophage T4 Protal-Neck Protein Gp20-Gp13-Gp14-Hfq Assembled In Vitro In C6 Symmetry
Organism: Escherichia phage t4
Method: ELECTRON MICROSCOPY Resolution:2.91 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
Structure Of Bacteriophage T4 Protal-Neck Mismatch Complex Gp20-Gp14-Gp13 Assembled In Vitro In C6 Symmetry
Organism: Escherichia phage t4
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: POV, AV0, ERG |

