Search Count: 60
All
Selected
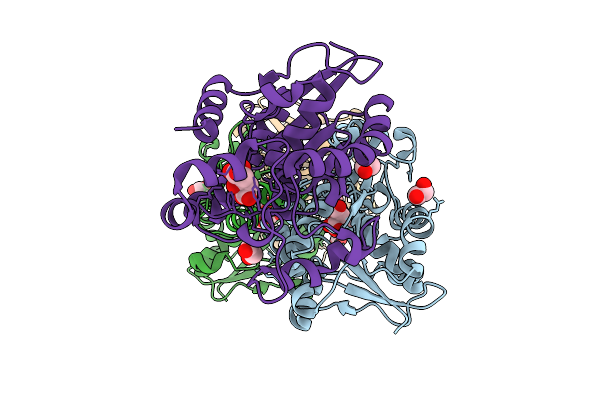 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: CIT, EDO, ACT, GOL |
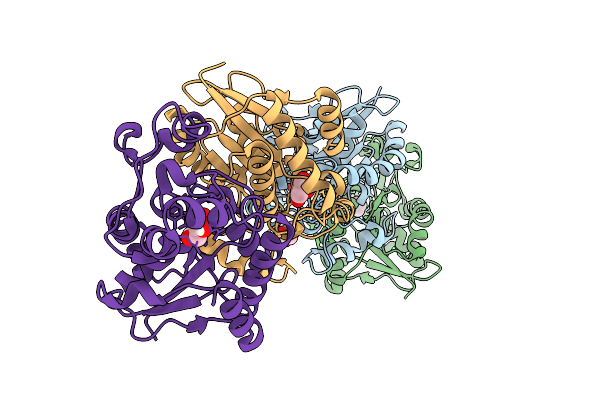 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: GOL, EDO |
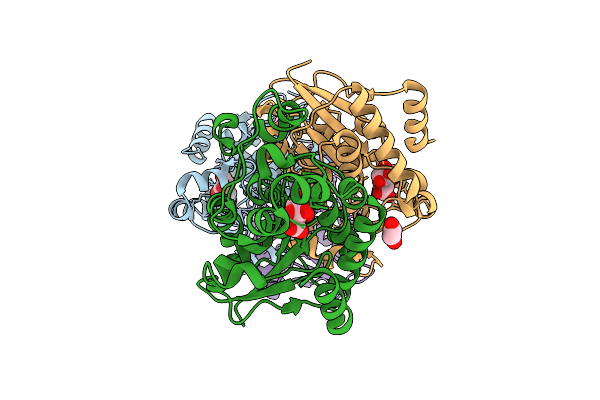 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: CIT, EDO |
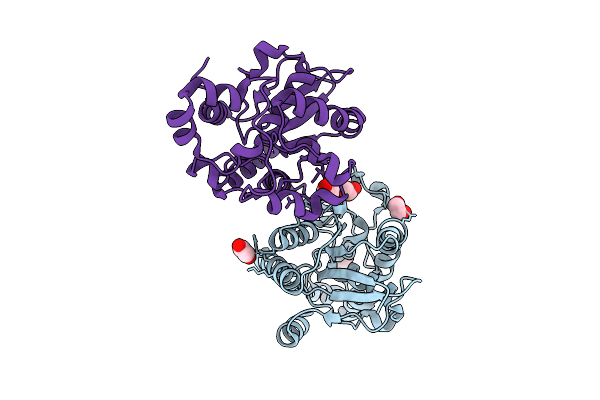 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: GOL, EDO |
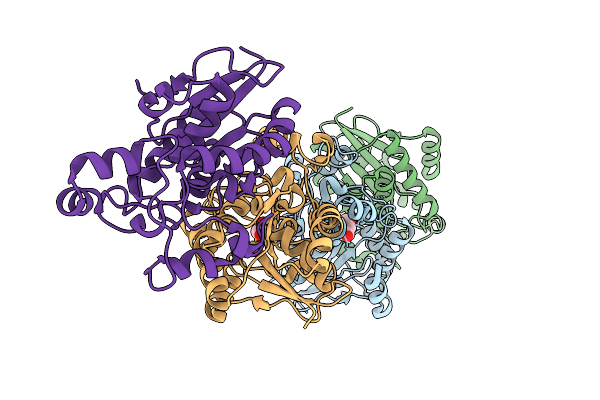 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: PEG, EDO |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: RIBOSOME Ligands: ZN, MG |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN |
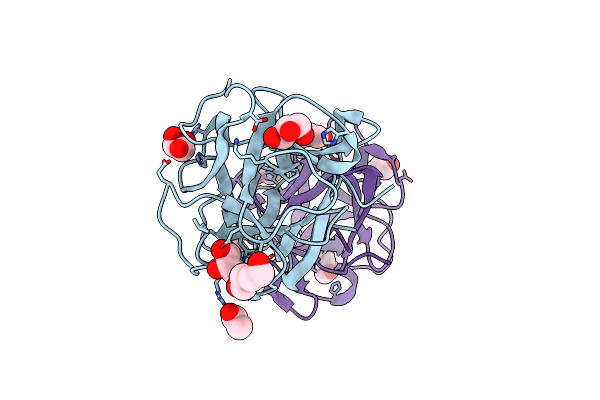 |
Organism: Mytilus californianus
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2024-12-18 Classification: SUGAR BINDING PROTEIN Ligands: GOL, ACT, PEG |
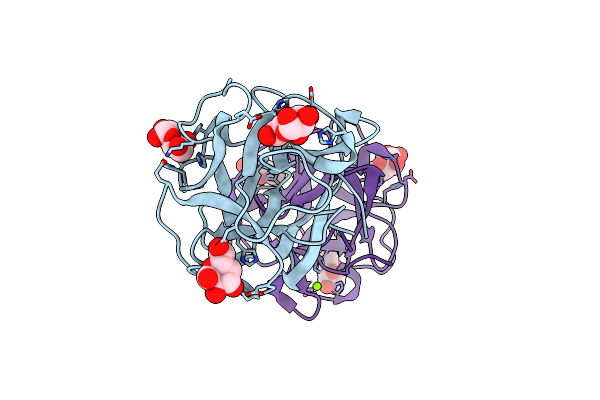 |
Organism: Mytilus californianus
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2024-12-18 Classification: SUGAR BINDING PROTEIN Ligands: GLA, MG |
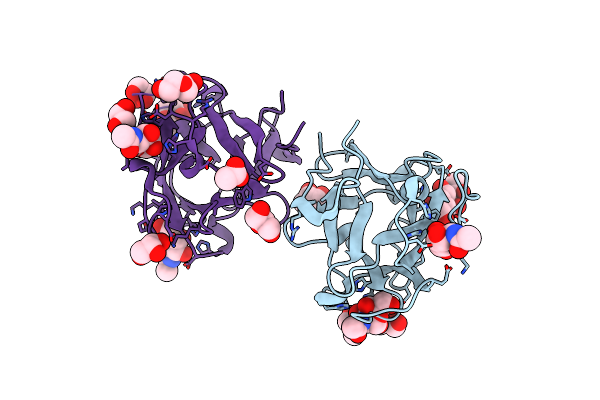 |
Organism: Mytilus californianus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2024-12-18 Classification: SUGAR BINDING PROTEIN Ligands: A2G, GOL, A1AAJ, ACT, PEG |
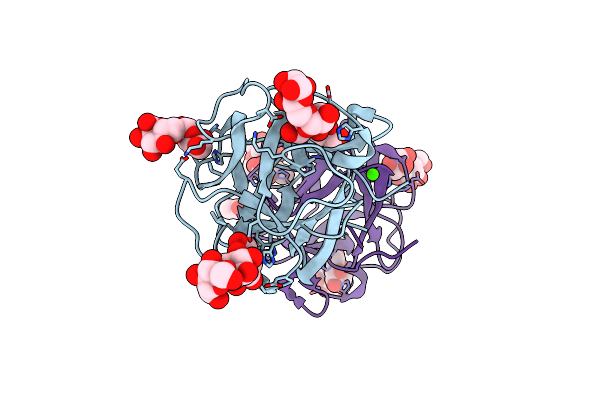 |
Organism: Mytilus californianus
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2024-12-18 Classification: SUGAR BINDING PROTEIN Ligands: CA, ACT, GOL |
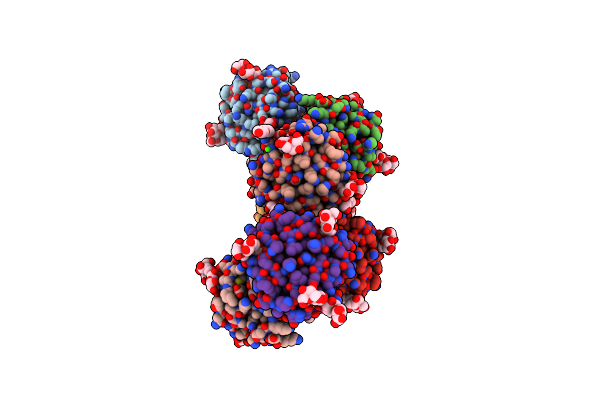 |
Organism: Mytilus californianus
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2024-12-18 Classification: SUGAR BINDING PROTEIN Ligands: GLA, EPE, PG4, CA, ACT, GOL |
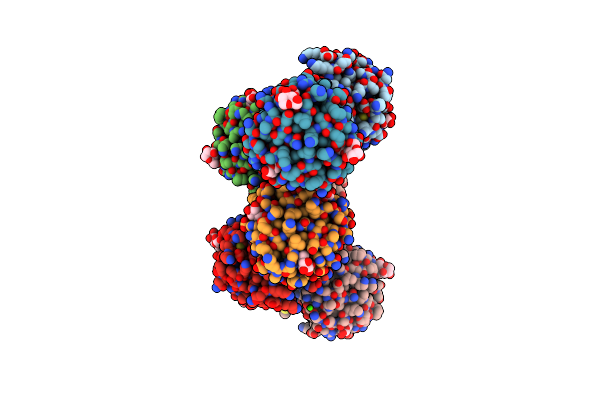 |
Organism: Mytilus californianus
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2024-12-18 Classification: SUGAR BINDING PROTEIN Ligands: GAL, CA |
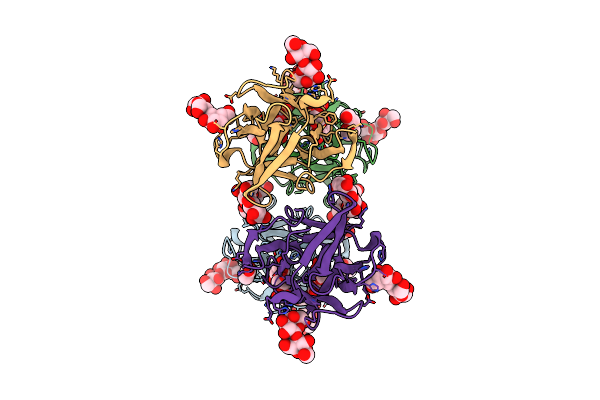 |
Organism: Mytilus californianus
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-12-18 Classification: SUGAR BINDING PROTEIN Ligands: GOL, ACT, PEG, NI |
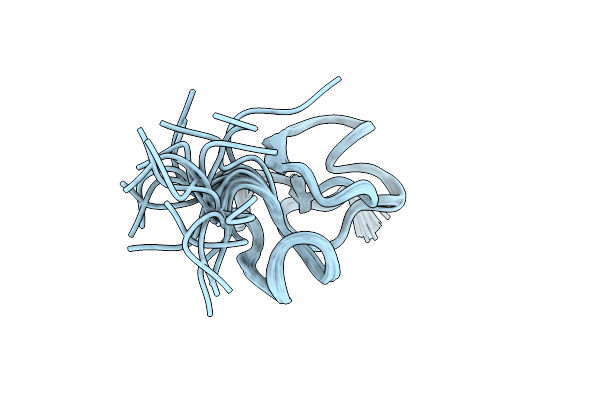 |
Organism: Triticum aestivum
Method: SOLUTION NMR Release Date: 2024-06-05 Classification: SUGAR BINDING PROTEIN |
 |
Organism: Triticum aestivum
Method: SOLUTION NMR Release Date: 2024-06-05 Classification: SUGAR BINDING PROTEIN |
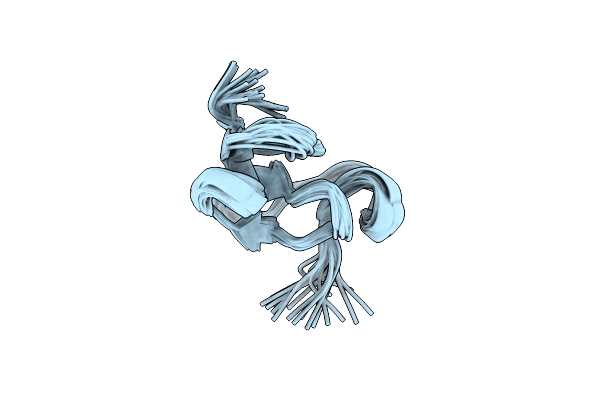 |
Organism: Triticum aestivum
Method: SOLUTION NMR Release Date: 2024-06-05 Classification: SUGAR BINDING PROTEIN |
 |
Organism: Moraxella
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-08-23 Classification: HYDROLASE Ligands: MLI |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: DE NOVO PROTEIN |

