Search Count: 979
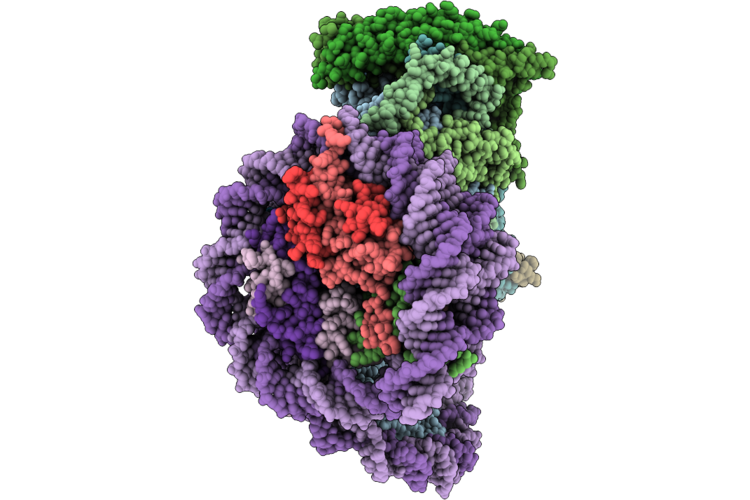 |
Cryo-Em Structure Of The Histone Deacetylase Complex Rpd3L In Complex With Di-Nucleosome
Organism: Saccharomyces cerevisiae s288c, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: GENE REGULATION/DNA Ligands: ZN |
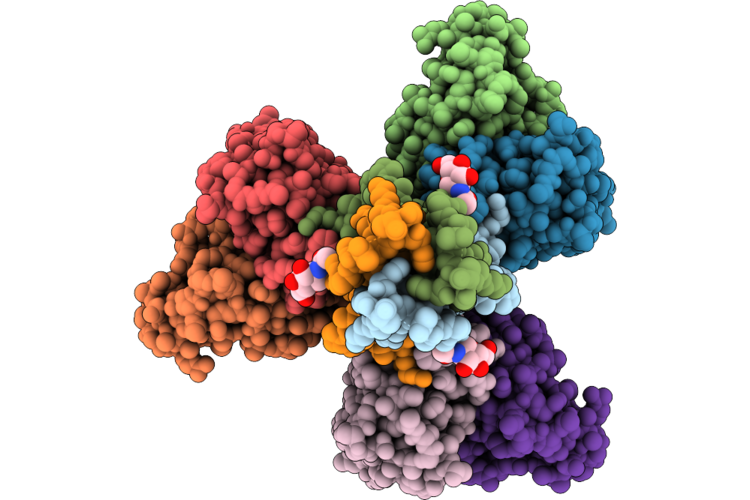 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To The Vn01H1 Fab (Fab Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
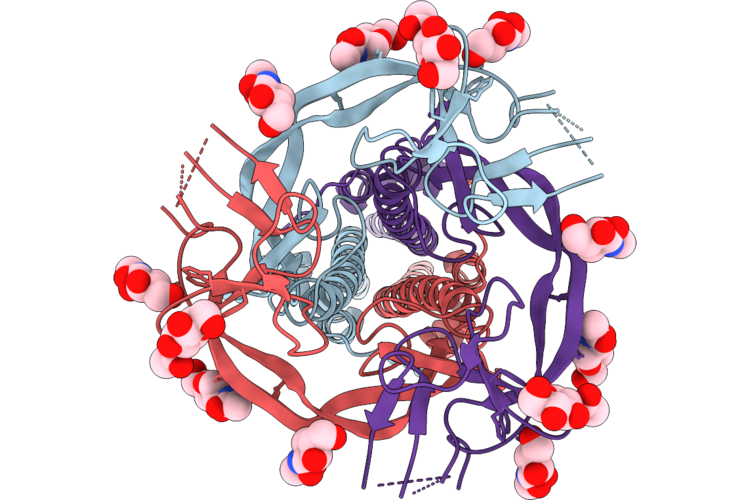 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To The Vn01H1 Fab (S2 Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
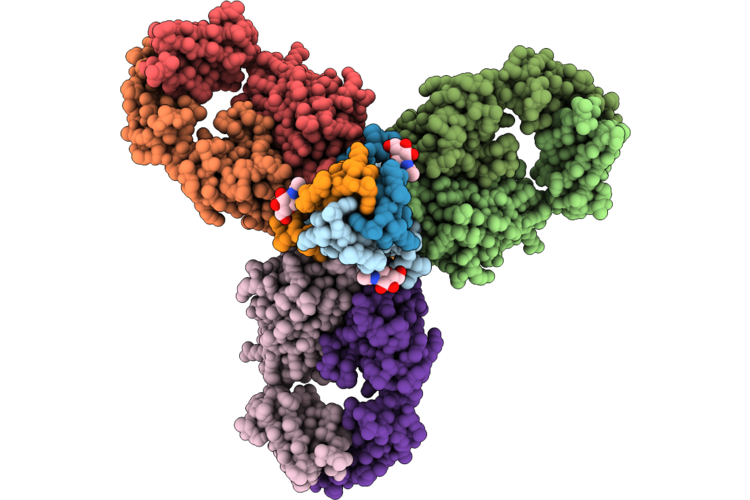 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To C77G12 (Fab Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cm1-Activated Gturc In Complex With Nascent Alpha-E254D Mutant Microtubules
Organism: Saccharomyces cerevisiae s288c, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: STRUCTURAL PROTEIN Ligands: GTP |
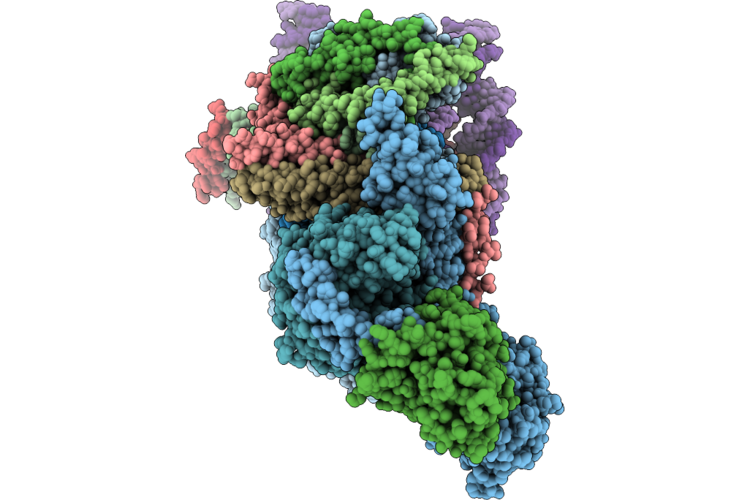 |
Cryo-Em Structure Of The Histone Deacetylase Complex Rpd3L In Complex With Mono-Nucleosome
Organism: Saccharomyces cerevisiae s288c, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: GENE REGULATION/DNA Ligands: ZN |
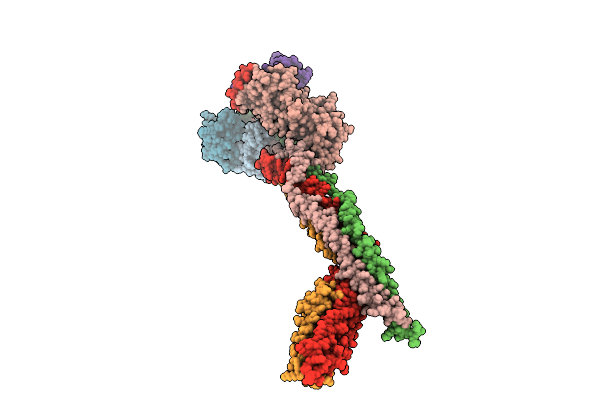 |
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Lacking The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
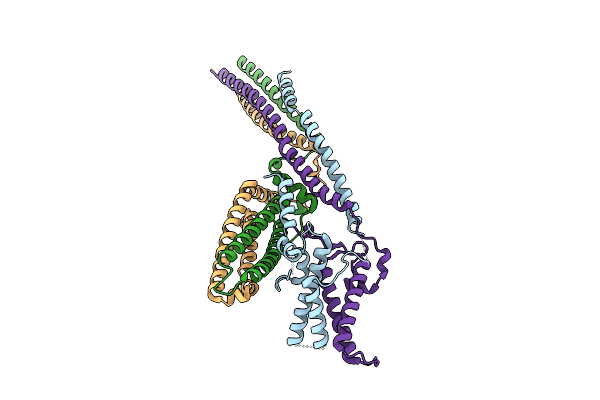 |
Cryo-Em Structure Of The Base Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
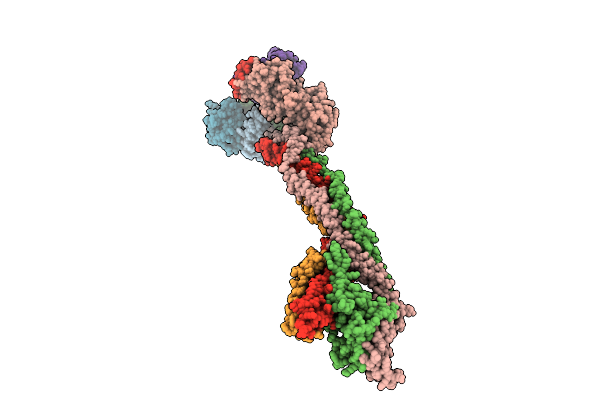 |
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae, Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: Y01, XKP, C14, D12, D10, A1EOT, NAG |
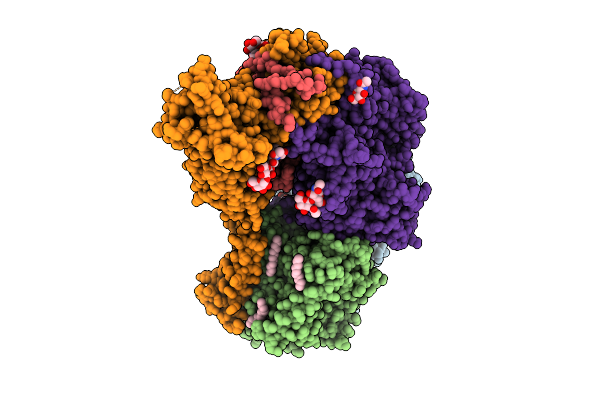
 |
Organism: Human alphaherpesvirus 1, Saccharomyces cerevisiae s288c, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: REPLICATION Ligands: A1BXD, ZN |
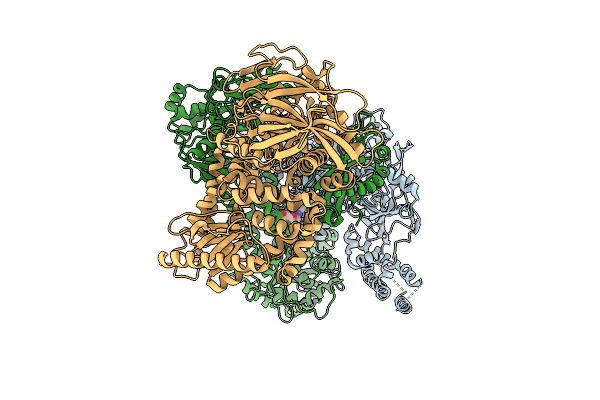
 |
The Helicase Module Of The Helicase-Primase Complex From Hhv1 Bound With Ssdna And Amenamevir
Organism: Human alphaherpesvirus 1, Saccharomyces cerevisiae s288c, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: REPLICATION Ligands: A1BXD, ZN |
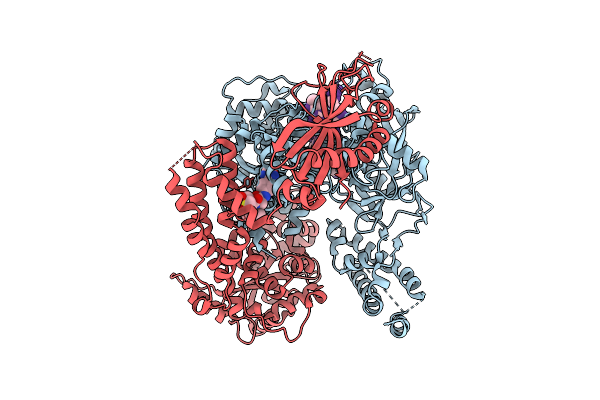
 |
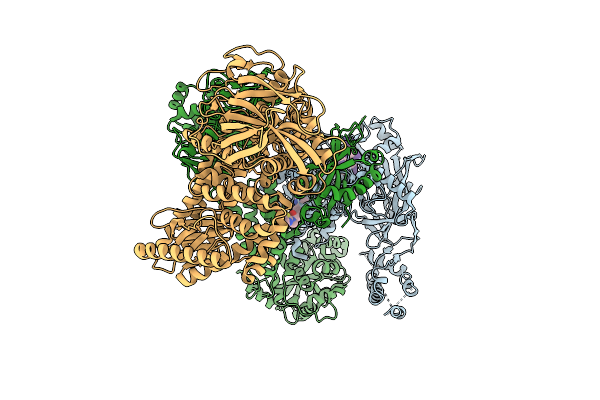
The Primase Module Of The Helicase-Primase Complex From Hhv1 Bound With Ssdna And Amenamevir
Organism: Saccharomyces cerevisiae s288c, Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: REPLICATION |
 |
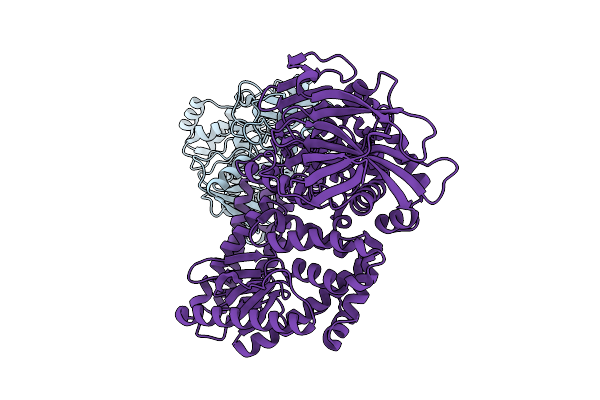
Organism: Human alphaherpesvirus 1, Saccharomyces cerevisiae s288c, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: REPLICATION Ligands: A1BXB, ZN |
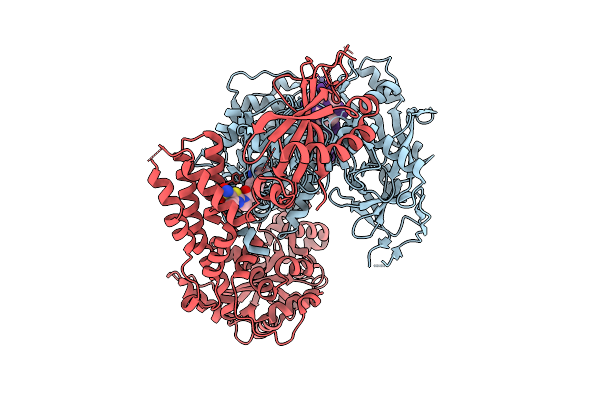 |
The Helicase Module Of The Helicase-Primase Complex From Hhv1 Bound With Ssdna And Pritelivir
Organism: Human alphaherpesvirus 1, Saccharomyces cerevisiae s288c, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: REPLICATION Ligands: A1BXB, ZN |
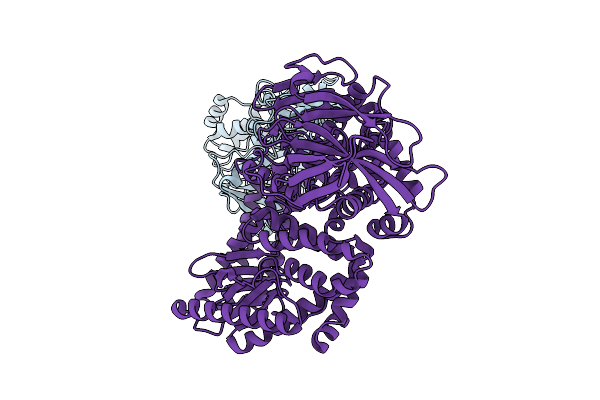 |
The Primase Module Of The Helicase-Primase Complex From Hhv1 Bound With Ssdna And Pritelivir
Organism: Saccharomyces cerevisiae s288c, Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: REPLICATION |
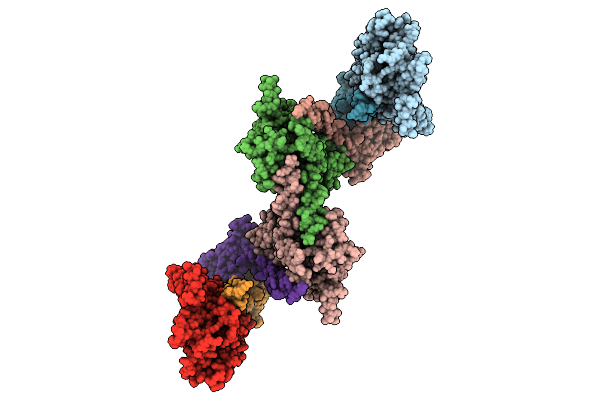 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSFERASE |
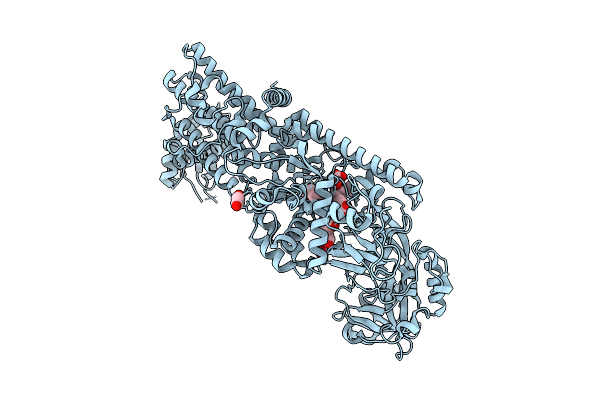 |
Crystal Structure Of Saccharomyces Cerevisiae Isoleucyl-Trna Synthetase In Complex With Reveromycin A And Isoleucine
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-04-08 Classification: LIGASE Ligands: GVU, ILE, EDO |
 |
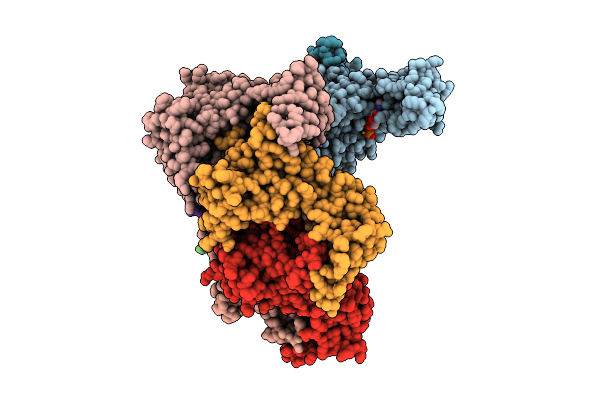
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To C77G12 (S2 Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Heptamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |

