Search Count: 5,702
All
Selected
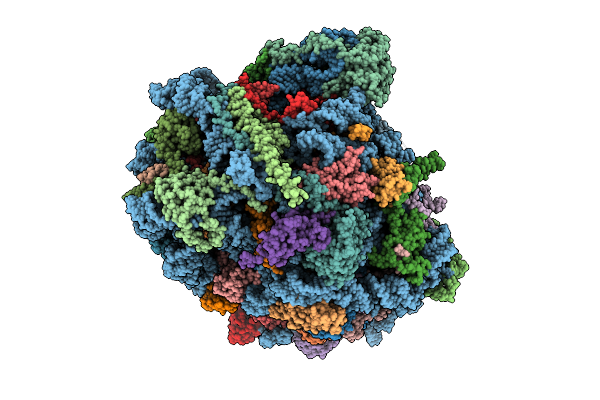 |
In Vitro Reconstituted Complex Of Purified S. Pombe Large Ribosomal Subunit And Snor
Organism: Schizosaccharomyces pombe 972h-
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: RIBOSOME Ligands: ZN |
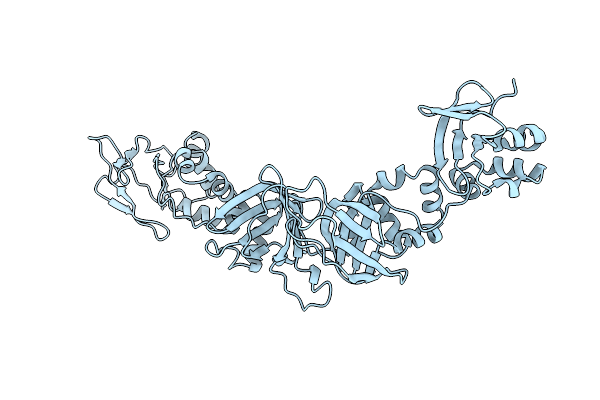 |
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-01 Classification: CELL ADHESION |
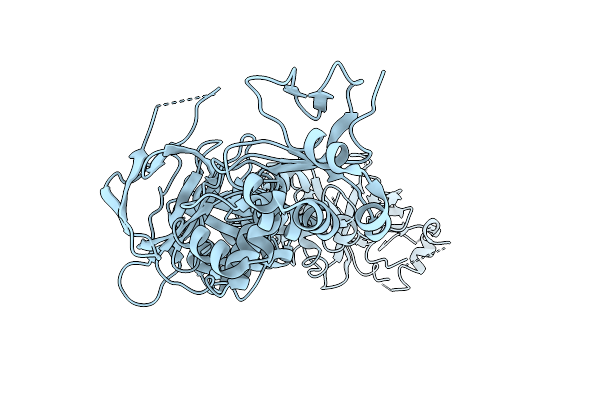 |
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-01 Classification: CELL ADHESION |
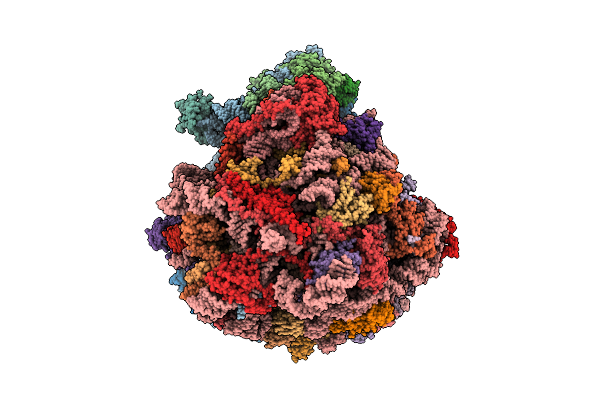 |
Organism: Schizosaccharomyces pombe
Method: ELECTRON MICROSCOPY Resolution:3.38 Å Release Date: 2026-04-01 Classification: RIBOSOME Ligands: ZN |
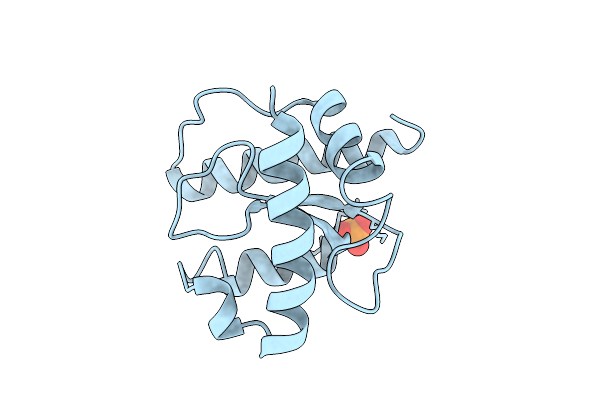 |
X-Ray Structure Of The C-Terminal Domain (Residues 366-485) Of S. Pombe Threonylcarbamoyladenosine Dehydratase At 1.55 A Resolution
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
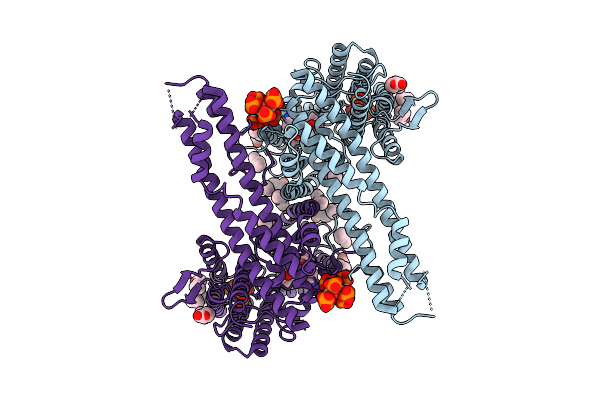
Organism: Schizosaccharomyces pombe (strain 972 / atcc 24843)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: TRANSPORT PROTEIN Ligands: POV, 8PE, PO4 |
 |
Organism: Schizosaccharomyces pombe 972h-
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: TRANSPORT PROTEIN Ligands: POV, 8PE, IHP, PO4 |
 |
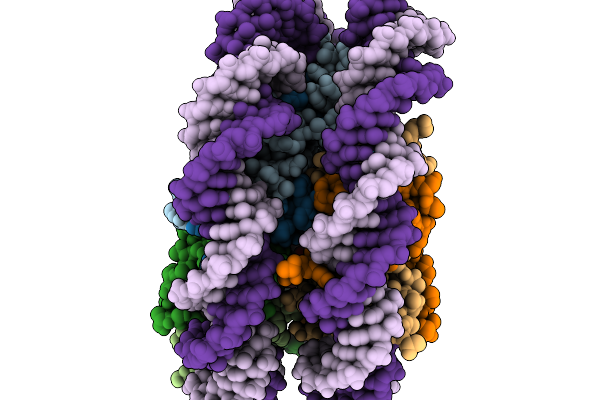
Organism: Schizosaccharomyces pombe dm3757
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: HYDROLASE |
 |
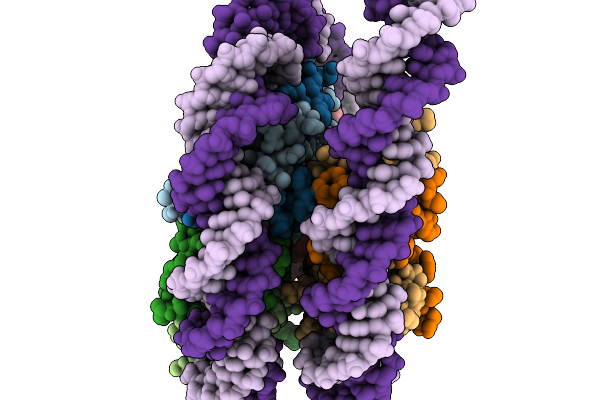
Organism: Schizosaccharomyces pombe (strain 972 / atcc 24843)
Method: ELECTRON MICROSCOPY Resolution:2.99 Å Release Date: 2026-02-04 Classification: NUCLEAR PROTEIN/DNA |
 |
Organism: Schizosaccharomyces pombe (strain 972 / atcc 24843)
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2026-02-04 Classification: NUCLEAR PROTEIN/DNA |
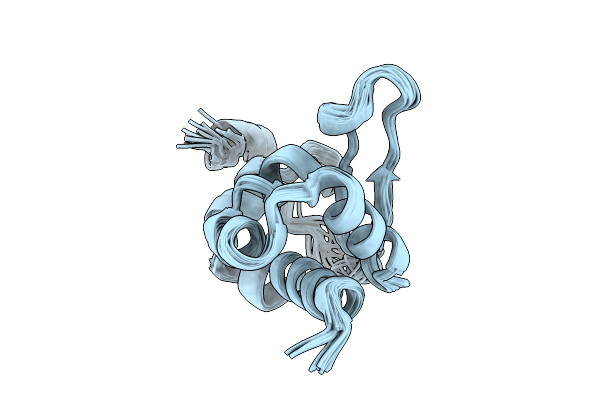 |
Organism: Schizosaccharomyces pombe 972h-
Method: SOLUTION NMR Release Date: 2025-12-24 Classification: RNA BINDING PROTEIN |
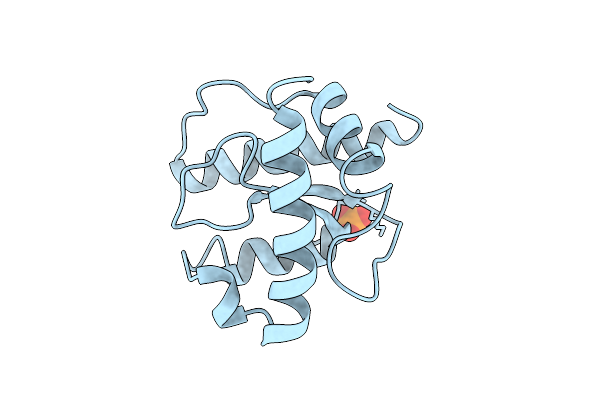 |
X-Ray Structure Of The C-Terminal Domain (Residues 366-485) Of S. Pombe Threonylcarbamoyladenosine Dehydratase
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-12-24 Classification: RNA BINDING PROTEIN Ligands: PO4 |
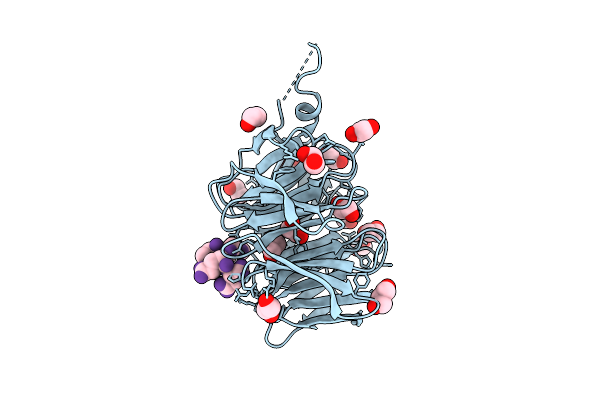 |
Organism: Schizosaccharomyces pombe, Murine hepatitis virus strain a59
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2025-10-22 Classification: PROTEIN TRANSPORT Ligands: EDO, GOL |
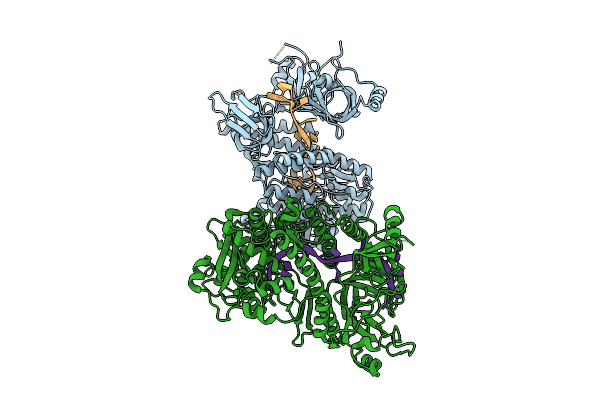 |
Organism: Schizosaccharomyces pombe (strain 972 / atcc 24843), Schizosaccharomyces pombe 927
Method: X-RAY DIFFRACTION Resolution:3.52 Å Release Date: 2025-08-06 Classification: HYDROLASE Ligands: MG |
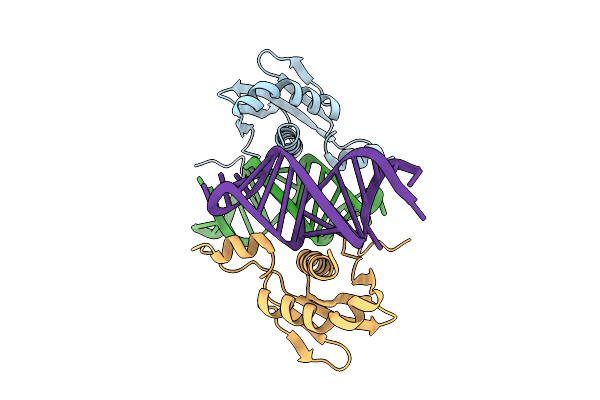 |
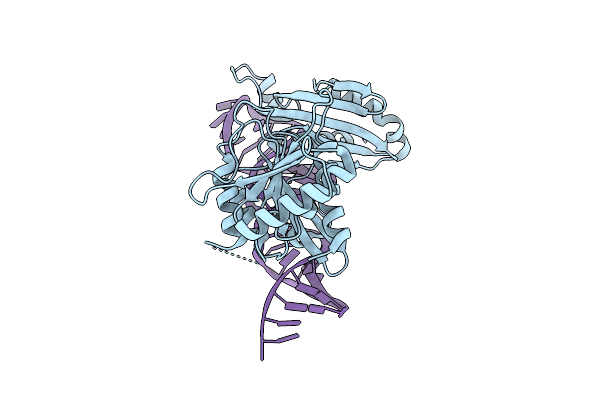
Crystal Structure Of Spmettl16 Kinase Associated 1 Domain In Complex With U6 Snrna Internal Stem Loop
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-08-06 Classification: TRANSFERASE/RNA |
 |
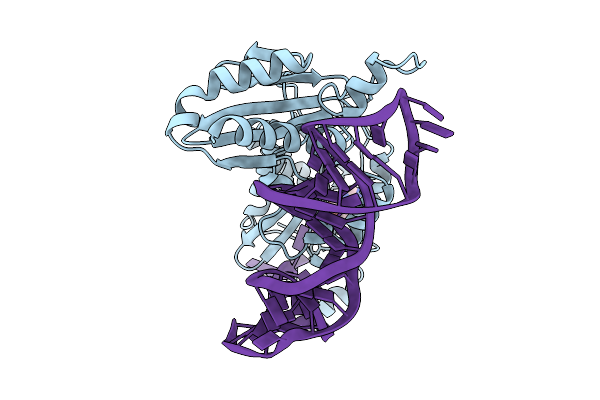
Organism: Schizosaccharomyces pombe
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: TRANSFERASE/RNA |
 |
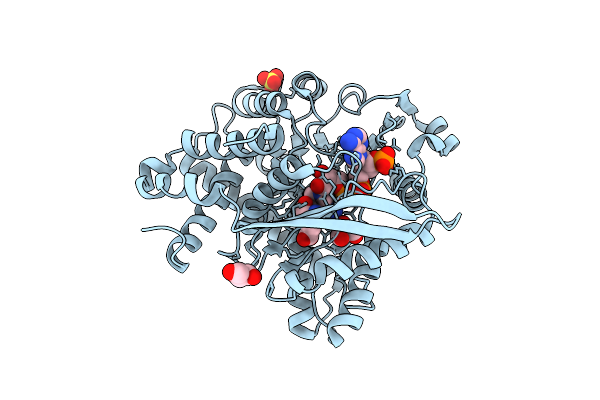
Organism: Schizosaccharomyces pombe
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: TRANSFERASE/RNA Ligands: SAM |
 |
Crystal Structure Of Nadp-Specific Glutamate Dehydrogenase Gdh1 From Schizosaccharomyces Pombe In Complex With Alpha-Iminoglutarate And Nadp+
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-07-30 Classification: OXIDOREDUCTASE Ligands: NAP, SO4, 2IT, GOL |
 |
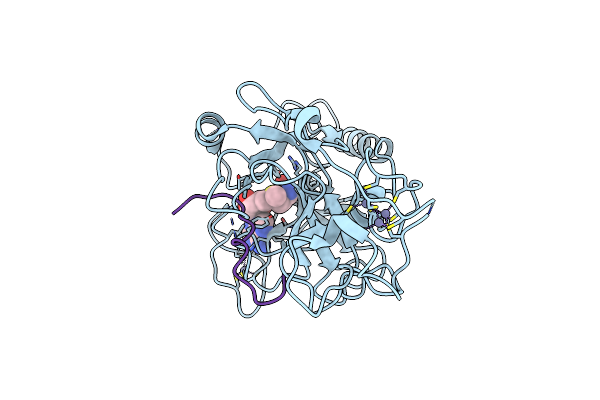
Organism: Schizosaccharomyces pombe 972h-, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-04-30 Classification: GENE REGULATION Ligands: SAM, ZN |
 |
Organism: Schizosaccharomyces pombe 972h-
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2025-04-30 Classification: GENE REGULATION Ligands: SAM, ZN |

