Search Count: 3,008
All
Selected
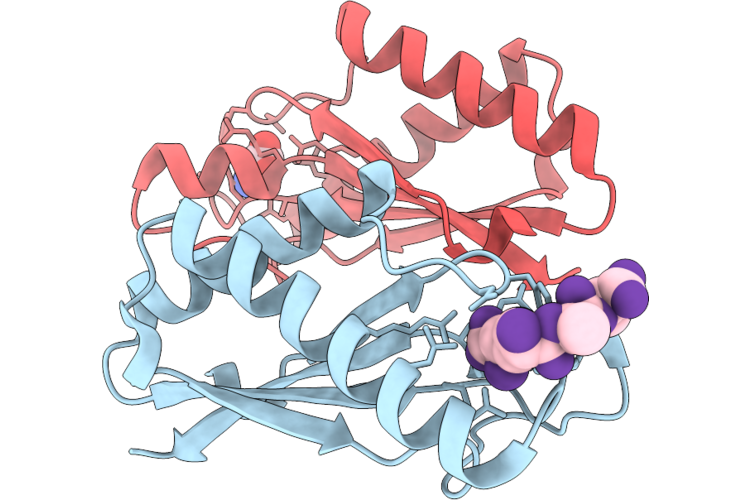 |
Crystal Structure Of The Periplasmic Domain Of Cadf From Campylobacter Jejuni In Complex With A Peptidoglycan Peptide
Organism: Campylobacter jejuni, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-05-06 Classification: PEPTIDE BINDING PROTEIN Ligands: API |
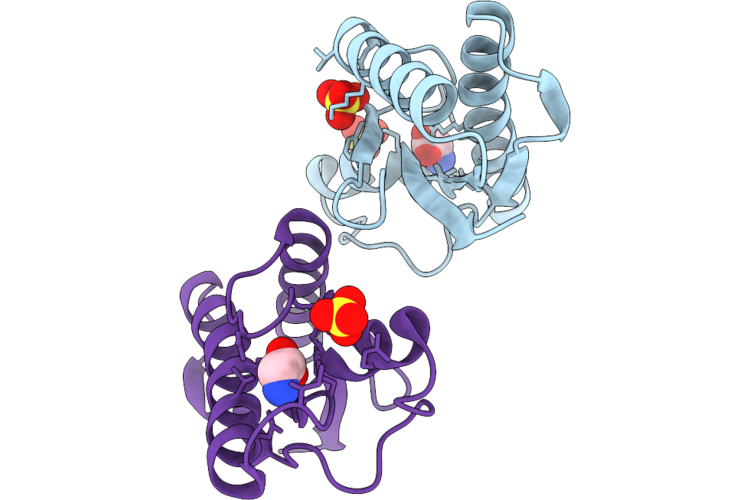 |
Crystal Structure Of The Periplasmic Domain Of Cadf From Campylobacter Jejuni In Complex With Glycine
Organism: Campylobacter jejuni
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-05-06 Classification: PEPTIDE BINDING PROTEIN Ligands: SO4, GLY |
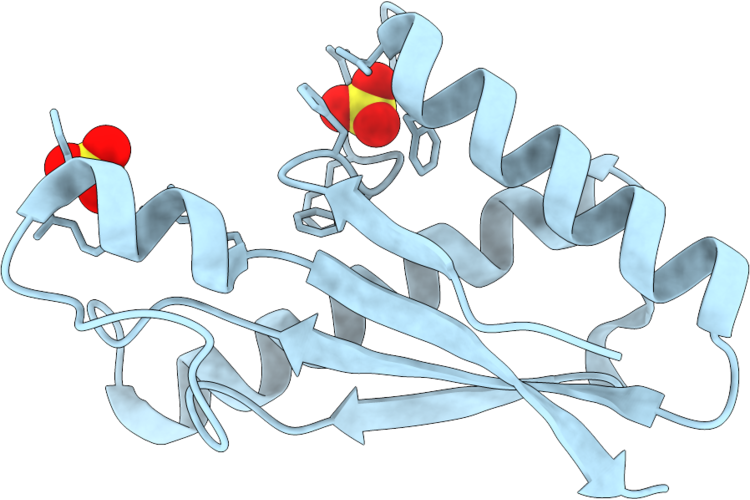 |
Crystal Structure Of The Periplasmic Domain Of Campylobacter Jejuni Cadf R268E
Organism: Campylobacter jejuni
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-06 Classification: PEPTIDE BINDING PROTEIN Ligands: SO4 |
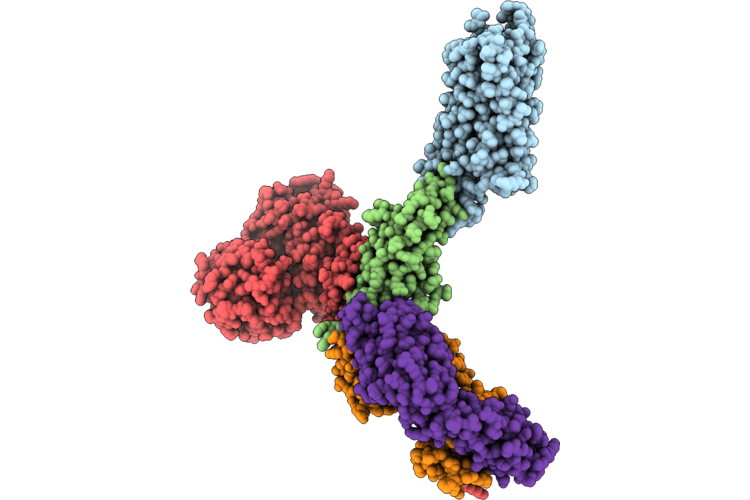 |
Organism: Escherichia coli o157:h7, Staphylococcus aureus, Staphylococcus aureus subsp. aureus mu50, Streptococcus sp. 'group g', Homo sapiens, Spodoptera frugiperda
Method: ELECTRON MICROSCOPY Resolution:3.08 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN |
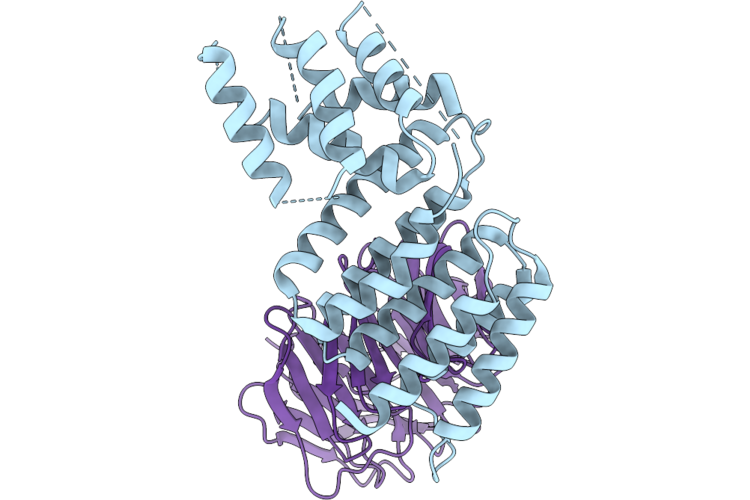 |
Stabilized Complex Of Chlamydia Trachomatic Efector Ct622 In Complex With Human Wd40 Domain Of Atg16L1
Organism: Escherichia coli o157:h7, Chlamydia trachomatis, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: PROTEIN TRANSPORT |
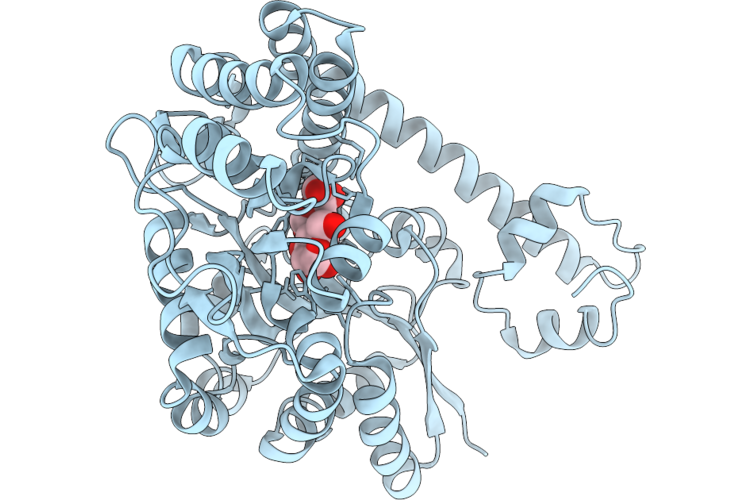 |
The Structure Of Egalitarian In Complex With The K10 Mrna Localization Signal Reveals A Modular Binding Surface Required For Function
Organism: Escherichia coli (strain k12), Drosophila melanogaster
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-04-29 Classification: RNA BINDING PROTEIN |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.35 Å Release Date: 2026-04-29 Classification: SUGAR BINDING PROTEIN |
 |
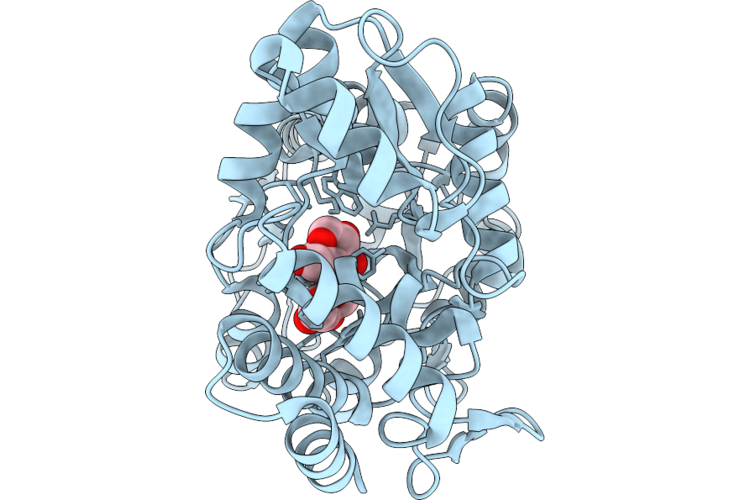
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Major State, Active Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.15 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.45 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Major State, Inactive Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.12 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.08 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
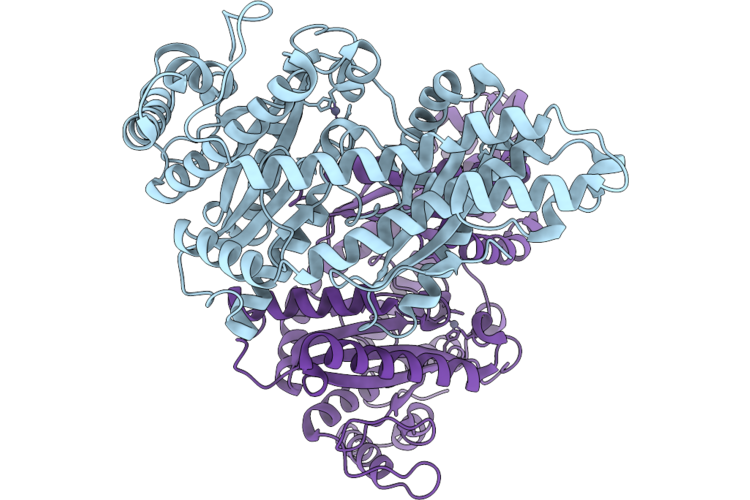 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 1
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.18 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
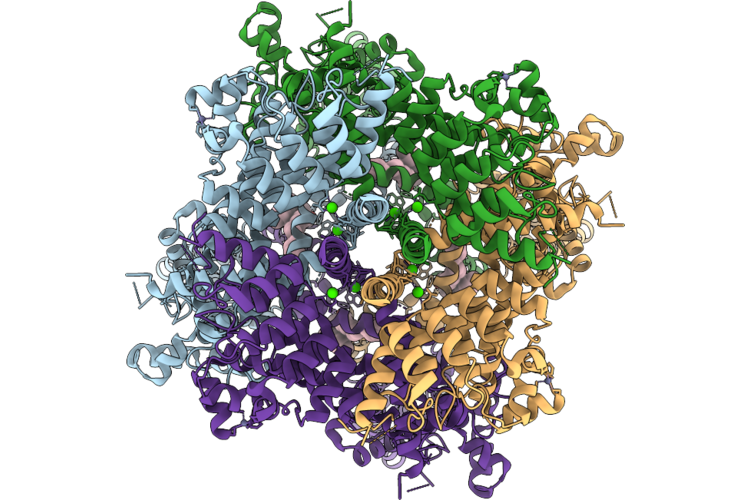
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 2
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.22 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
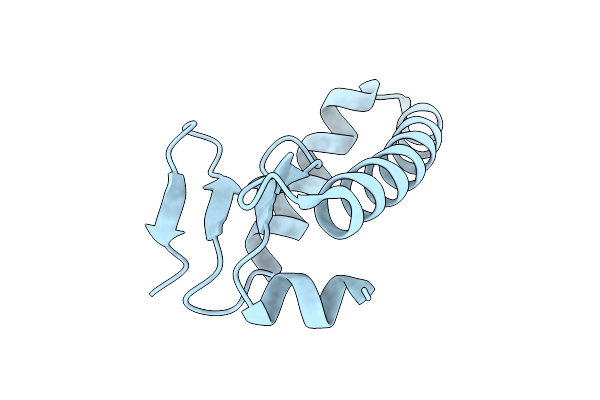
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: METAL TRANSPORT Ligands: CA, ZN, A1L5I |
 |
Organism: Escherichia coli bw25113
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-08 Classification: UNKNOWN FUNCTION |
 |
Organism: Nitratidesulfovibrio vulgaris str. hildenborough
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: SF4, 402, CMO |
 |
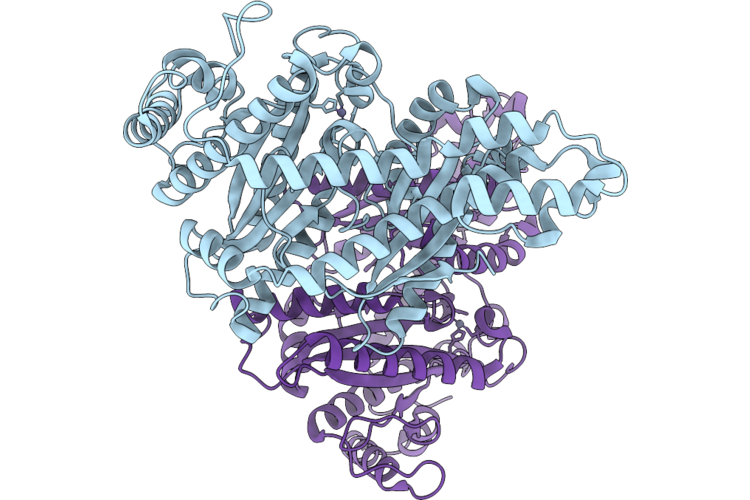
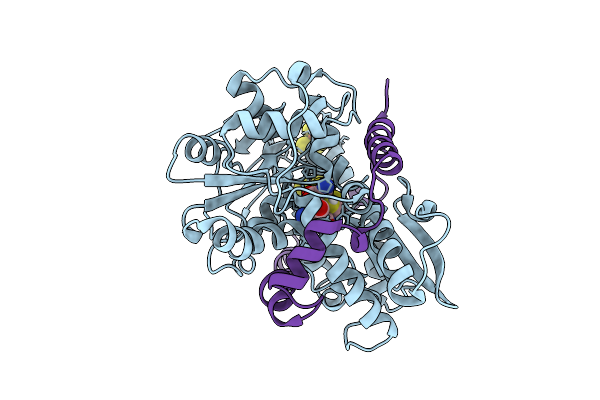
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: PEG, EDO, HEM, TRP, CYN, NA |

