Search Count: 305
All
Selected
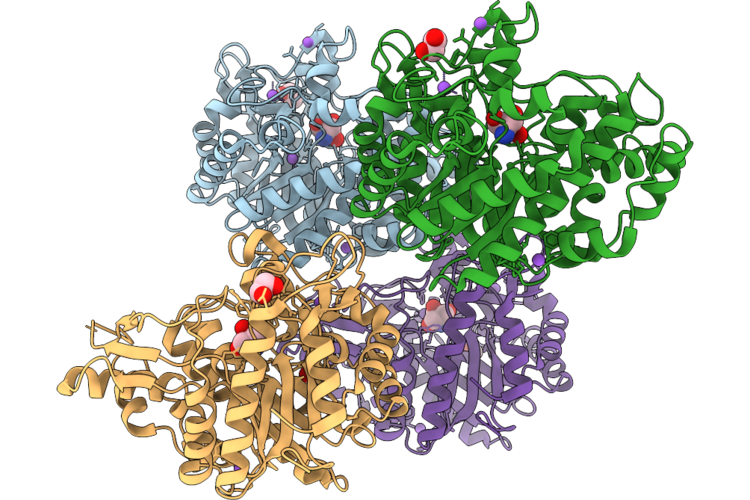 |
Organism: Thermoanaerobacterium saccharolyticum
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: TRS, EDO, NA |
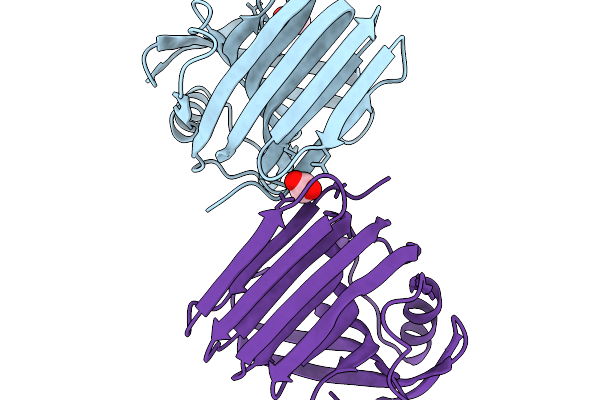 |
Organism: Thermoanaerobacterium saccharolyticum
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: ACT |
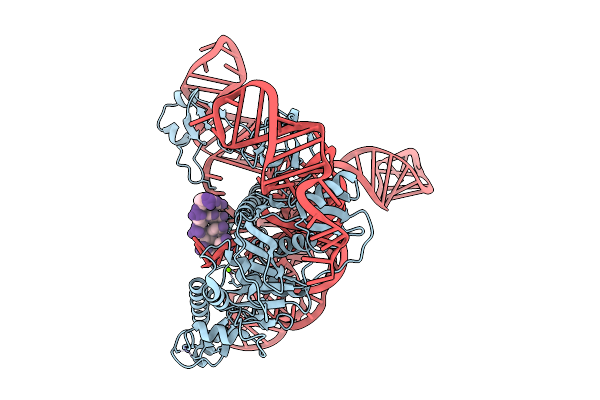 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2026-01-21 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
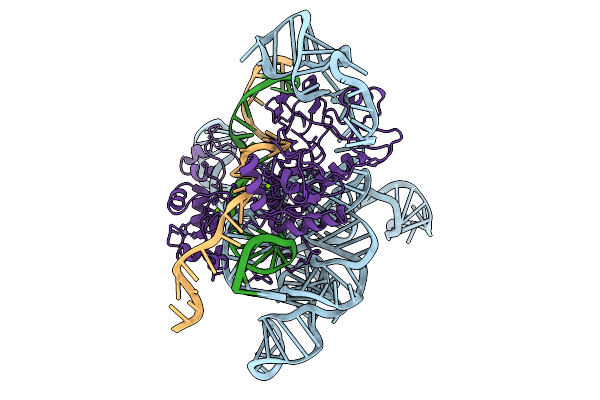 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-01-21 Classification: DNA BINDING PROTEIN Ligands: MG, ZN |
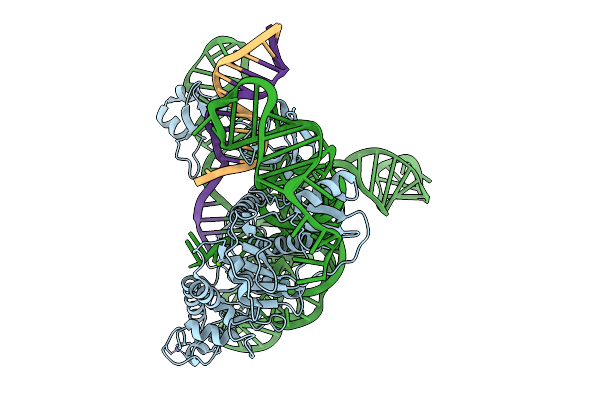 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.39 Å Release Date: 2026-01-21 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
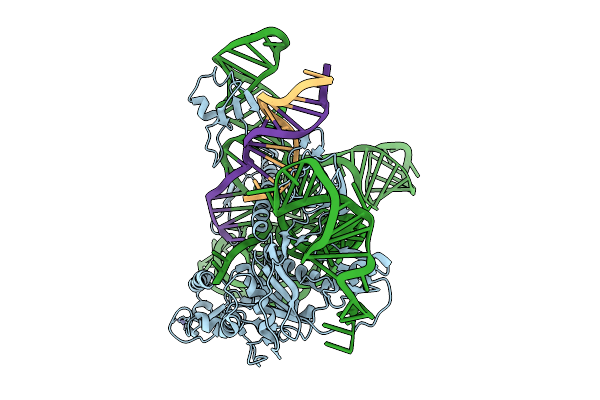 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.09 Å Release Date: 2026-01-21 Classification: DNA BINDING PROTEIN Ligands: ZN, MG |
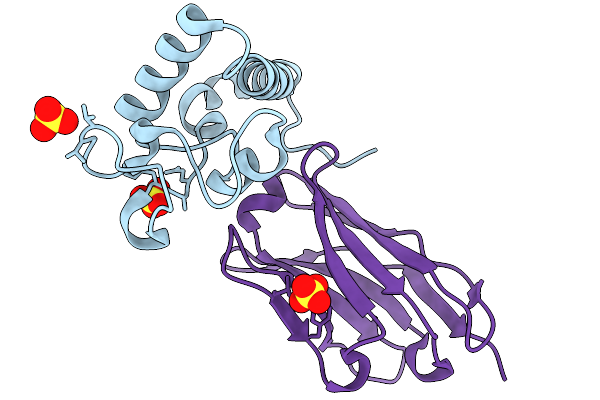 |
Crystal Structure Of Tetraspanin Cd63Mutant Large Extracellular Loop (Lel) In Complex With Sybody Sb3
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-11-19 Classification: CELL ADHESION Ligands: SO4 |
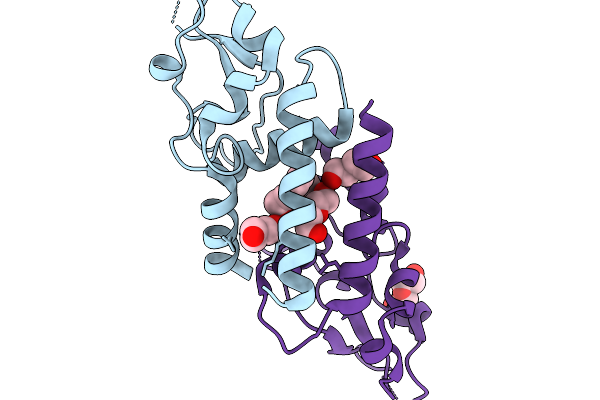 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-11-19 Classification: ENDOCYTOSIS Ligands: PEG |
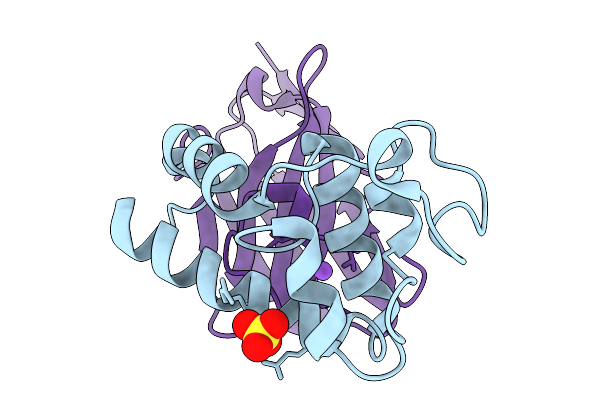 |
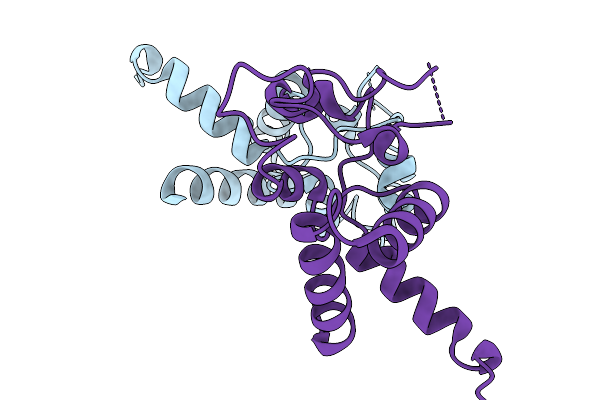
Crystal Structure Of Tetraspanin Cd63Mutant Large Extracellular Loop (Lel) In Complex With Sybody La4
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-11-19 Classification: CELL ADHESION Ligands: SO4, NA |
 |
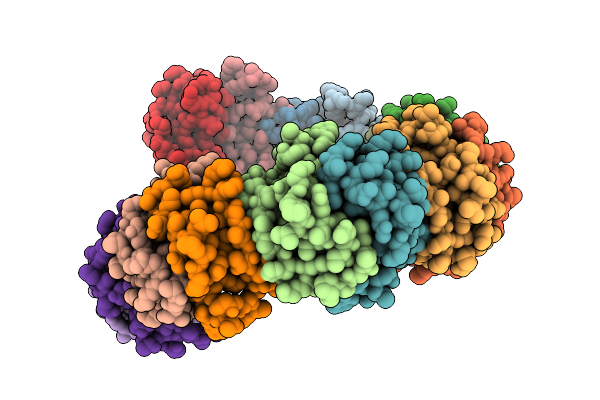
Crystal Structure Of Tetraspanin Tspan10Mutant Large Extracellular Loop (Lel)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-11-19 Classification: SIGNALING PROTEIN |
 |
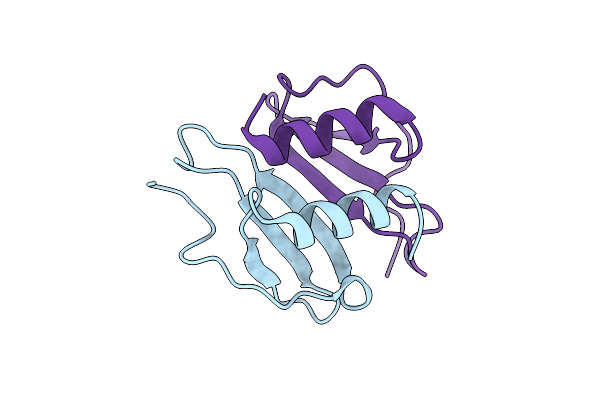
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.60 Å Release Date: 2025-11-12 Classification: CYTOKINE Ligands: NO3 |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2025-11-12 Classification: CYTOKINE |
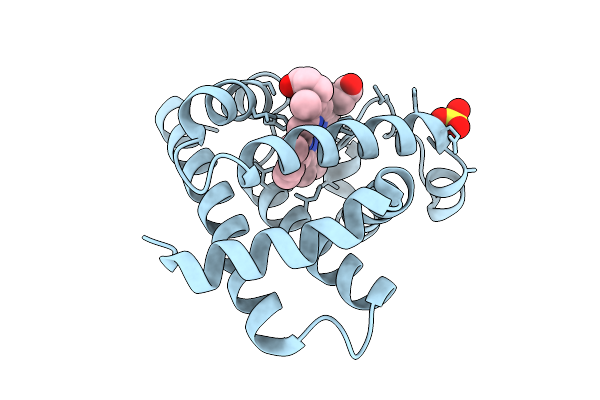 |
Serial Femtosecond Crystallography Structure Of Myoglobin From Equine Skeletal Muscle
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-11-12 Classification: OXYGEN STORAGE Ligands: SO4, HEM |
 |
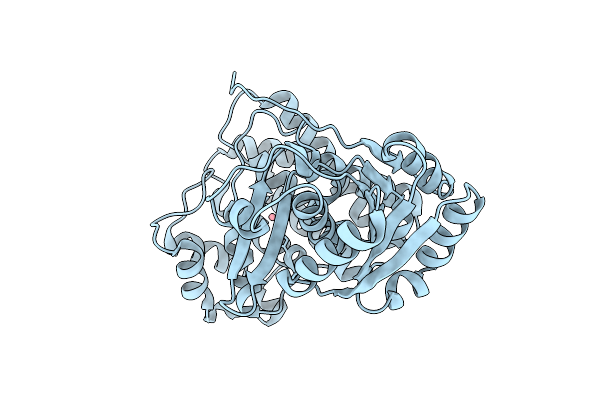
Room Temperature Structure Of Glucose Isomerase By Serial Femtosecond Crystallography
Organism: Streptomyces pseudogriseolus
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-11-12 Classification: ISOMERASE Ligands: MG |
 |
Organism: Fusobacterium nucleatum
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-10-29 Classification: BIOSYNTHETIC PROTEIN Ligands: CO |
 |
Organism: Fusobacterium nucleatum
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2025-10-29 Classification: BIOSYNTHETIC PROTEIN Ligands: CO, ADP |
 |
Organism: Thermoascus crustaceus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-07-02 Classification: HYDROLASE |
 |
Organism: Cellulomonas biazotea
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-05-28 Classification: HYDROLASE Ligands: GOL |
 |
Crystal Structure Of Beta-Glucosidase Cba3 From Cellulomonas Biazotea In Complex With Glucose
Organism: Cellulomonas biazotea
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-05-28 Classification: HYDROLASE Ligands: BGC |
 |
Organism: Montipora sp. 20
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-04-23 Classification: FLUORESCENT PROTEIN |

