Search Count: 39
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: GDP, A1CRR, MG, CIT |
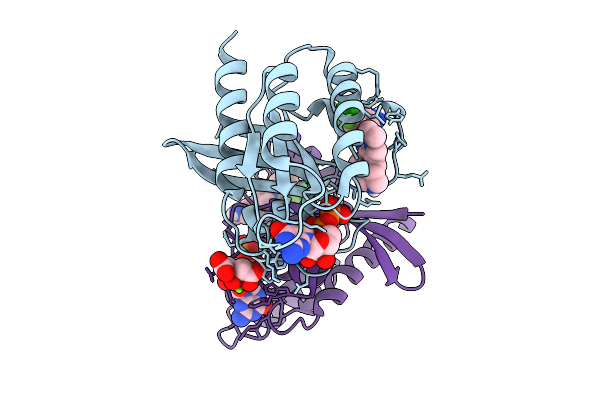 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: GDP, MG, A1CRV, CIT |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: GDP, A1CRW, MG, CIT |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: GDP, MG, A1CR1, CIT |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: GDP, MG, A1CR2, CIT |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: GDP, MG, A1CSD, CIT |
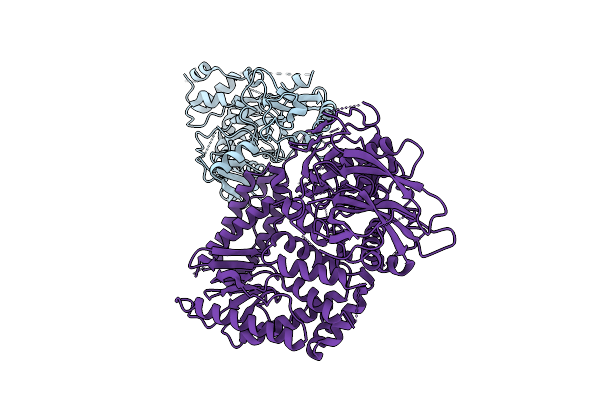 |
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna And Atp-Gamma-S
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
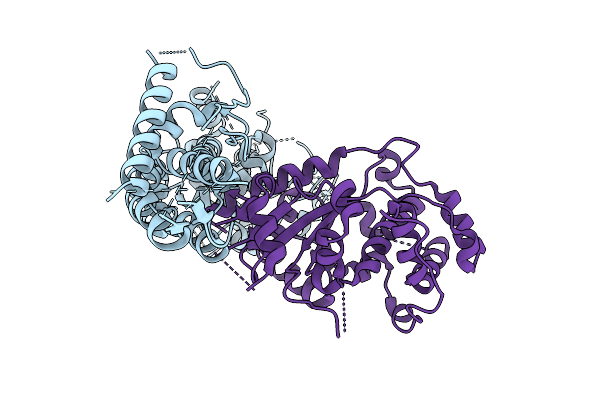 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna And Atp-Gamma-S
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
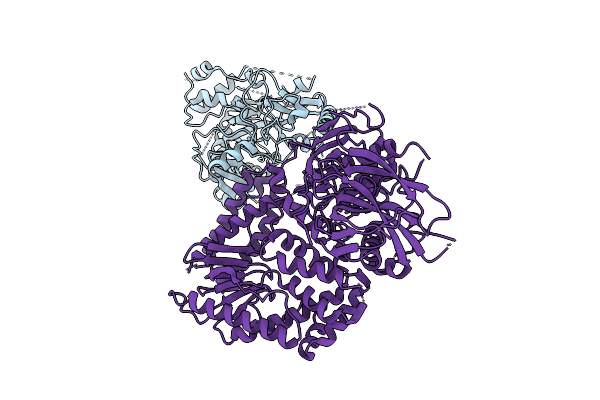 |
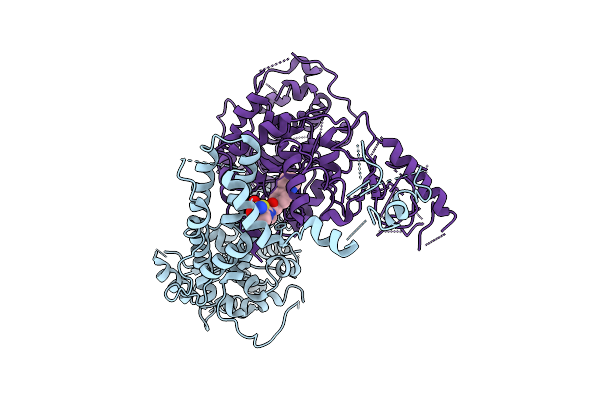
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna, Atp-Gamma-S And Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
 |
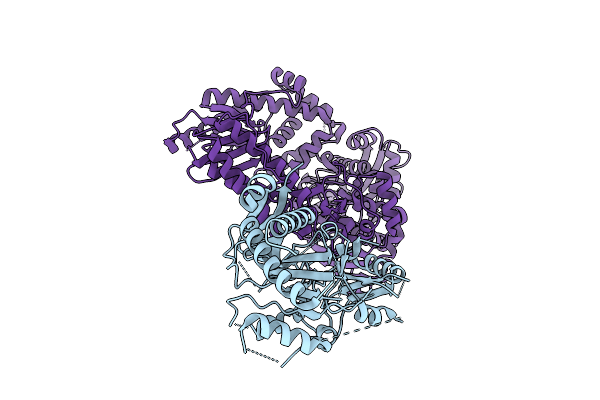
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna, Atp-Gamma-S And Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXB |
 |
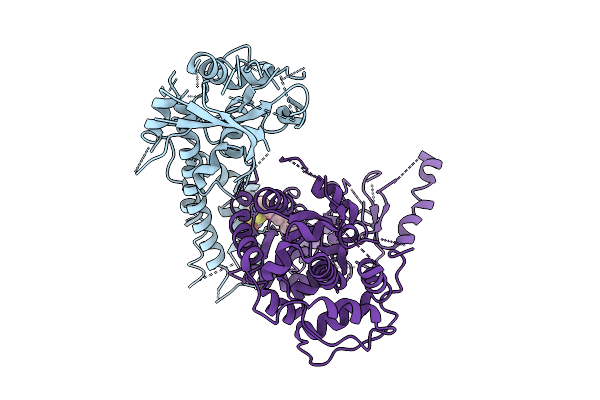
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR |
 |
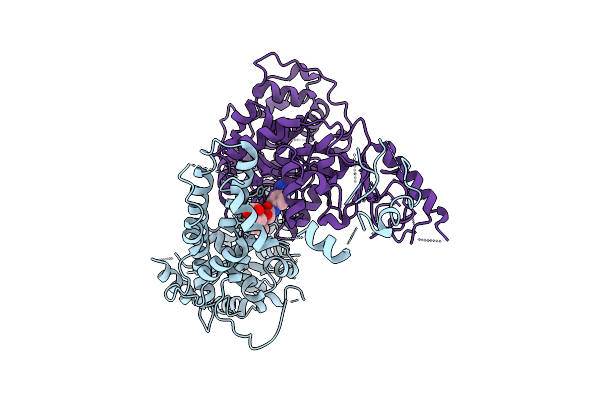
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXB |
 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Amenamevir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXD |
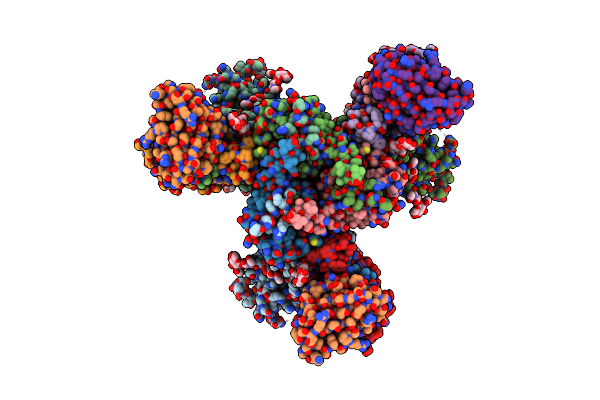 |
Cryo-Em Structure Of Hiv-1 Bg505Ds-Sosip.664 Env Trimer Bound To Dj85-B.01 Fab
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
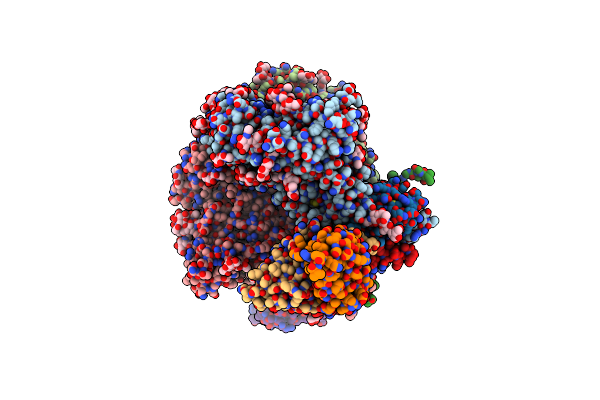 |
Cryo-Em Structure Of Hiv-1 Bg505Ds-Sosip.664 Env Trimer Bound To Dj85-C.01 Fab
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
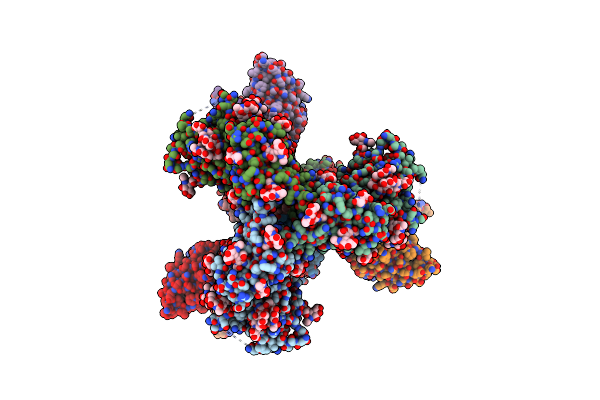 |
Cryo-Em Structure Of Hiv-1 Bg505Ds-Sosip.664 Env Trimer Bound To Dj85-D.01 Fab
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
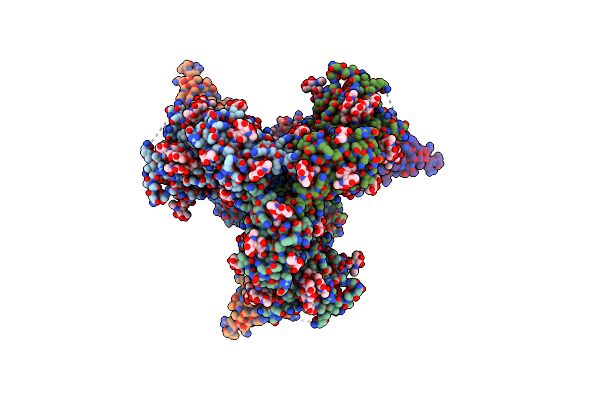 |
Cryo-Em Structure Of Hiv-1 Bg505Ds-Sosip.664 Env Trimer Bound To Dj85-E.01 Fab
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Cryo-Em Structure Of Hiv-1 Bg505Ds-Sosip.664 Env Trimer Bound To Herh-C.01 Fab
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Cryo-Em Structure Of Trnm-B*01 Fab In Complex With Hiv-1 Env Trimer Bg505.Ds Sosip
Organism: Macaca mulatta, Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2024-08-07 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Cryo-Em Structure Of Herh-B*01 Fab In Complex With Hiv-1 Env Trimer Bg505.Ds Sosip
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2024-08-07 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |

