Search Count: 202
All
Selected
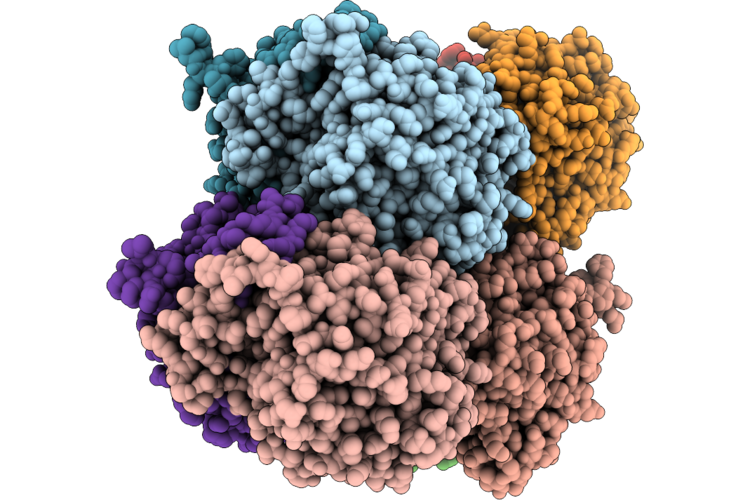 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-29 Classification: LIPID BINDING PROTEIN |
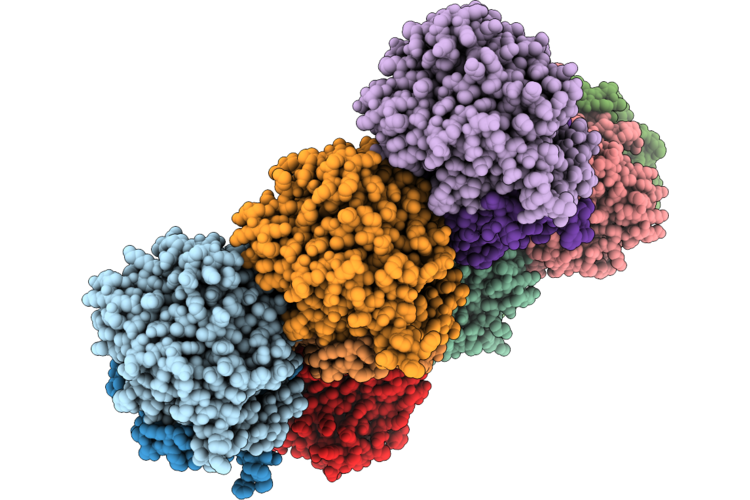 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.76 Å Release Date: 2026-04-29 Classification: LIPID BINDING PROTEIN Ligands: 4BW |
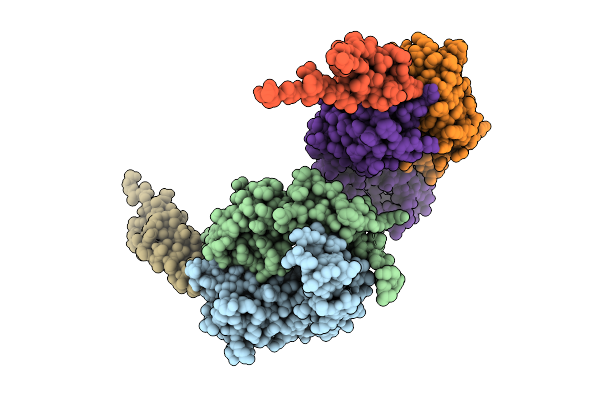 |
Organism: Plasmodium (plasmodium), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-03-18 Classification: IMMUNE SYSTEM |
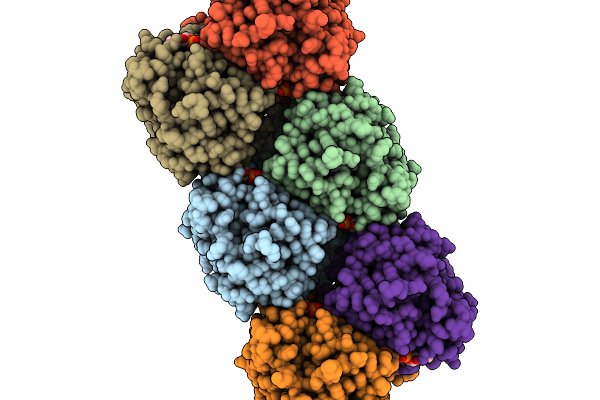 |
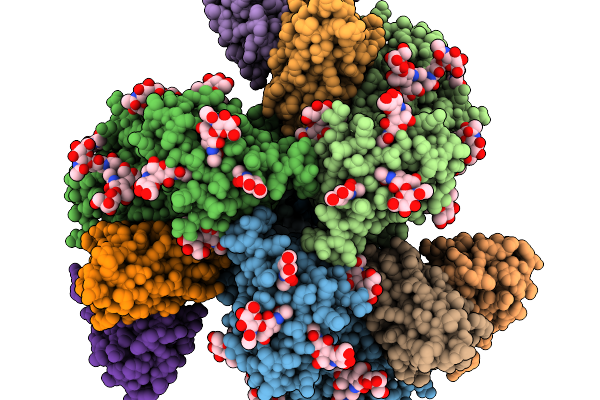
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: HYDROLASE Ligands: 4BW, PEE |
 |
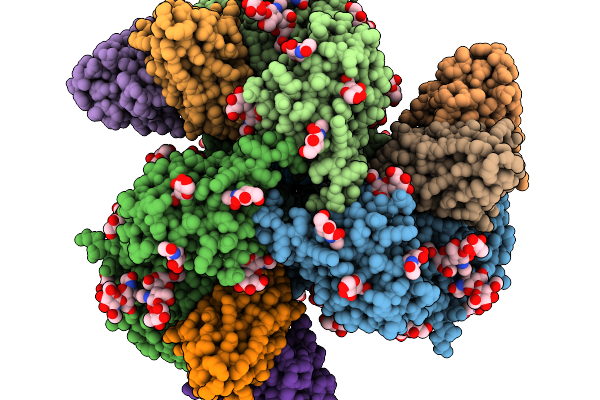
Organism: Homo sapiens, Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
 |
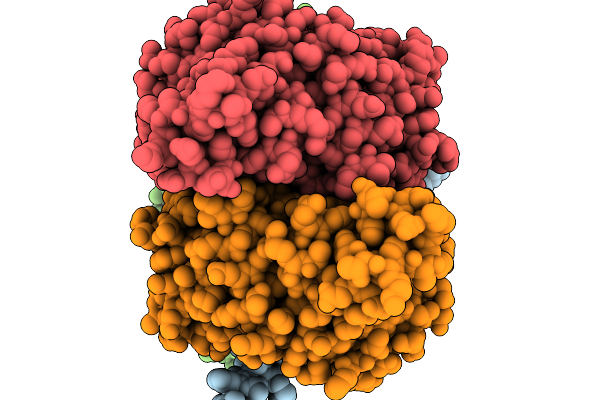
Organism: Macaca mulatta, Homo sapiens, Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
 |
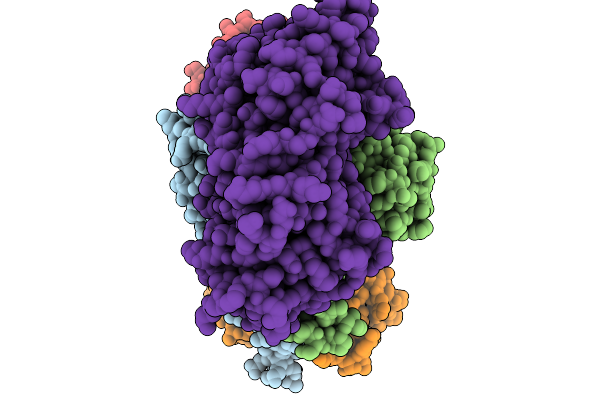
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: METAL TRANSPORT Ligands: PO4, ATP, MG |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: METAL TRANSPORT Ligands: PO4 |
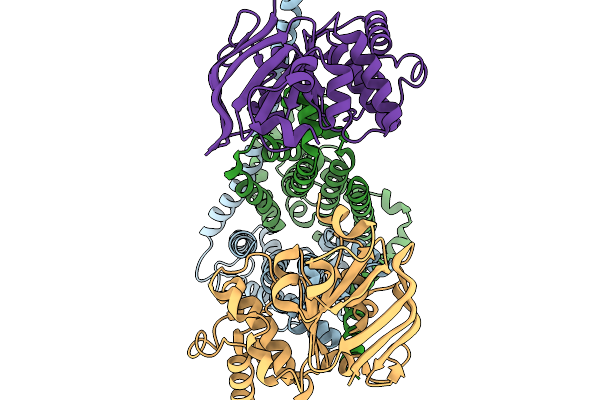 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: METAL TRANSPORT |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
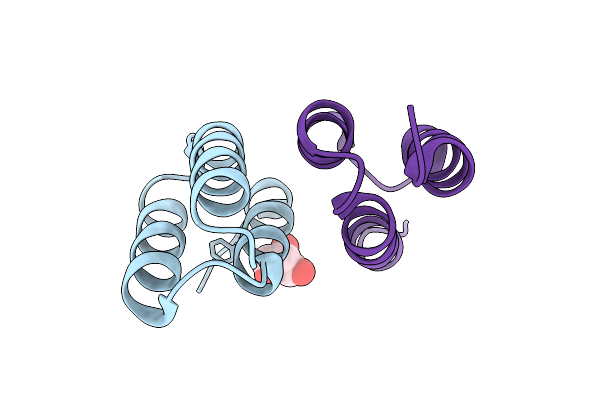 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING Ligands: GOL |
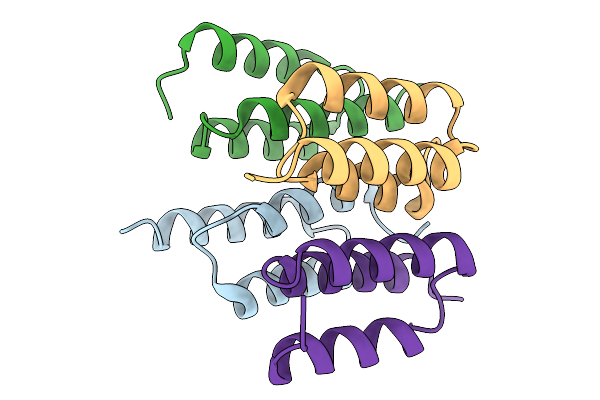 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING Ligands: SO4 |
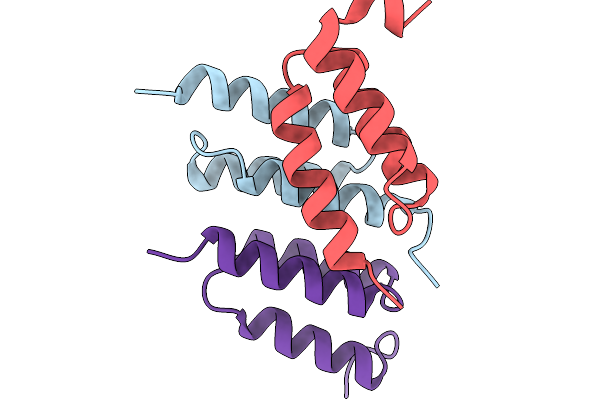 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING Ligands: EPE |
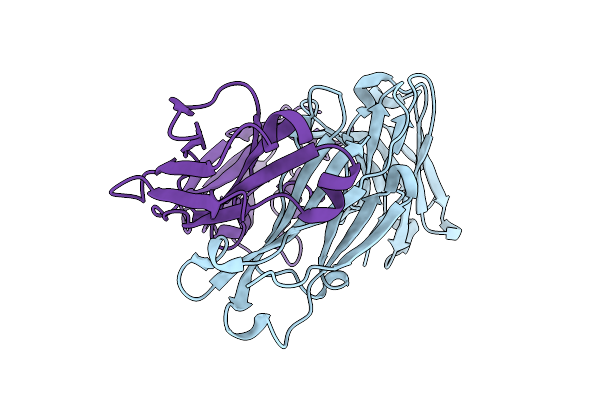 |
Crystal Structure Of Rm014, A Macaque-Derived Hiv V2-Apex-Targeting Antibody From Apexgt6 Immunization
Organism: Macaca
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-12-24 Classification: IMMUNE SYSTEM |
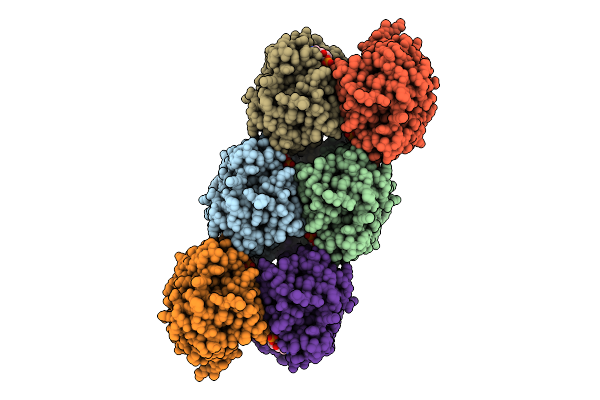 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: HYDROLASE Ligands: 4BW |
 |
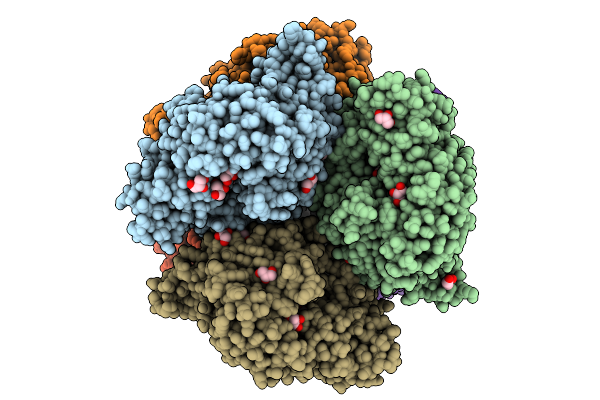
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GLC, GOL, NA, G1P |
 |
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GOL, GLC, PEG |

