Search Count: 258
All
Selected
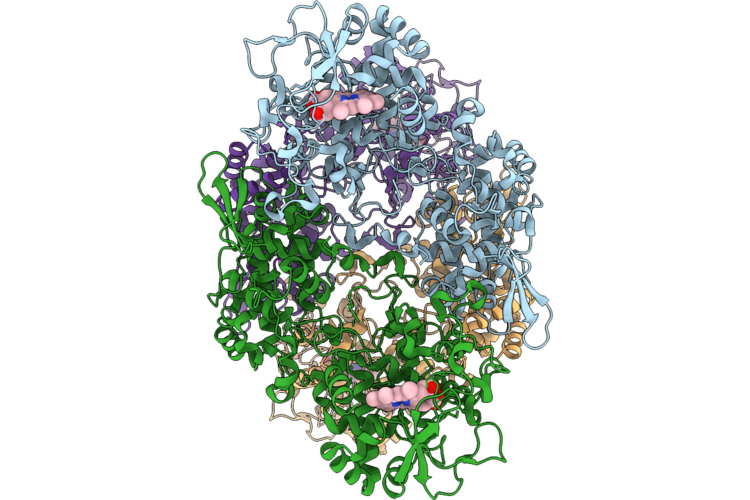 |
Cryoem Structure Of Eckatg S-Trp105 At 2.22 Angstrom Resolution Revealing An Asymmetric Sulfur Center In O=S-Trp
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
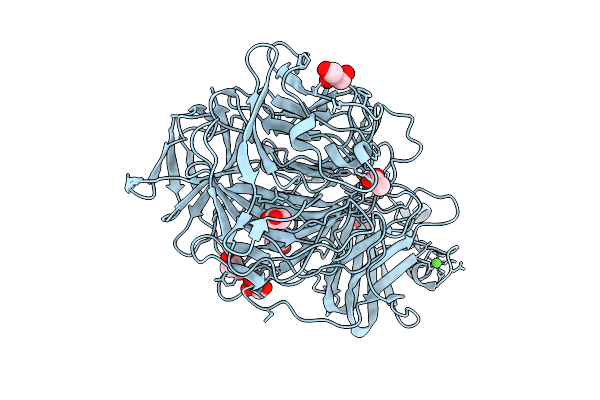 |
The 1.22 Angstrom Crystal Structure Of Galactose Oxidase Variant With Genetically Incorporated F2-Tyr495
Organism: Fusarium austroamericanum
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: CU, CA, ACT, GOL |
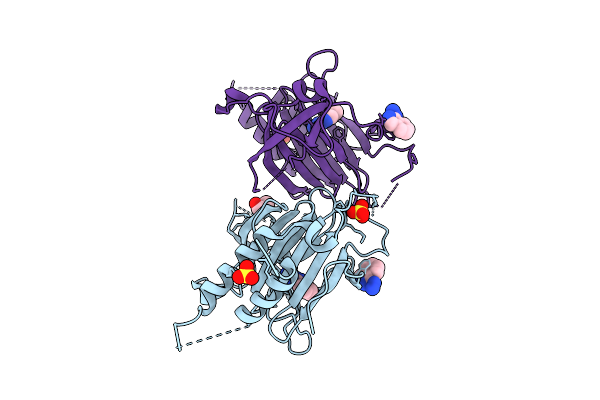 |
Crystal Structure Of Ferric Human Ado C18S/C239S Variant In Complex With Hydralazine At 1.98 Angstrom Resolution
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: FE, HLZ, SO4, GOL |
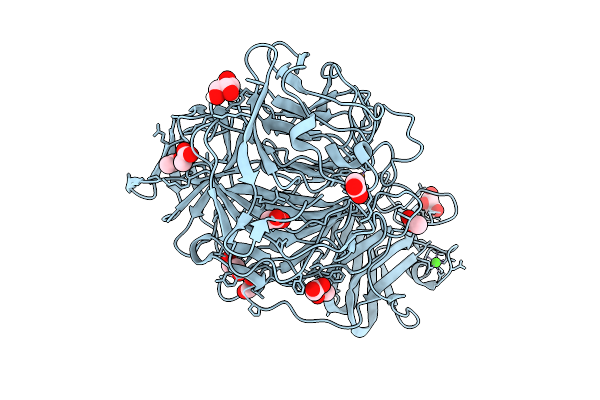 |
The 1.48 Angstrom Crystal Structure Of Galactose Oxidase Variant With Genetically Incorporated Cl2-Tyr495
Organism: Fusarium austroamericanum
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: CA, CU, ACT, GOL |
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-01-21 Classification: SIGNALING PROTEIN |
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-01-21 Classification: SIGNALING PROTEIN |
 |
Organism: Methanococcoides burtonii
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-11-19 Classification: LIGASE Ligands: MG |
 |
Crystal Structure Of Rubisco In Complex With Cabp From Methanococcoides Burtonii
Organism: Methanococcoides burtonii
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2025-11-19 Classification: LIGASE Ligands: CAP, MG |
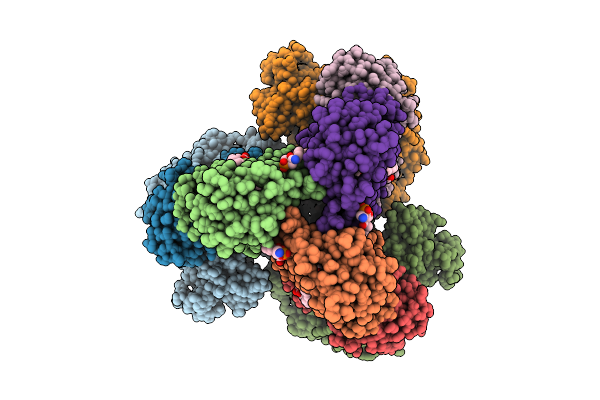 |
Cryo-Em Structure Of The Apo-Form Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
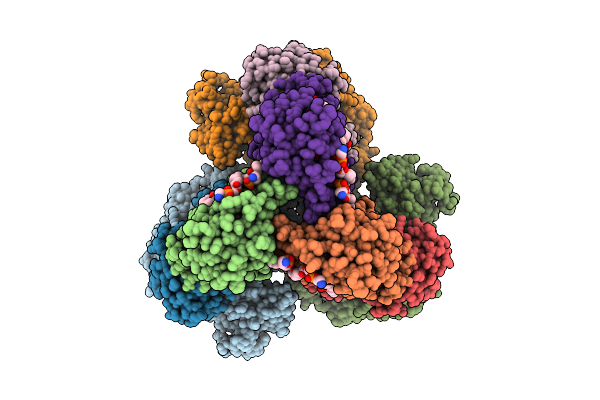 |
Cryo-Em Structure Of The Lipid-Bound Succiante Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
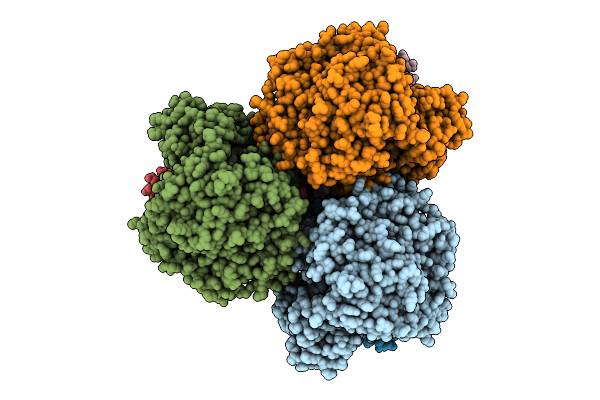 |
Cryo-Em Structure Of The Mk7-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, MQ7 |
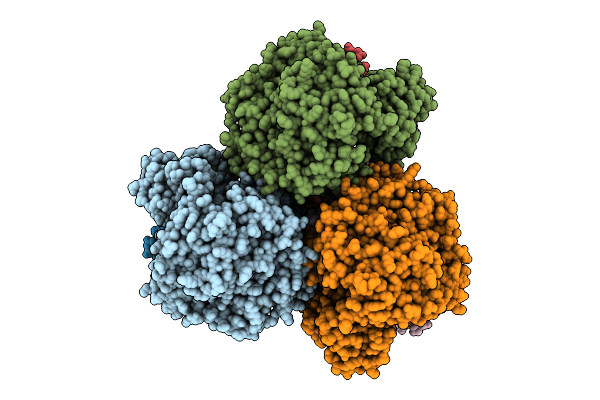 |
Cryo-Em Structure Of The Mk4-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, 1L3 |
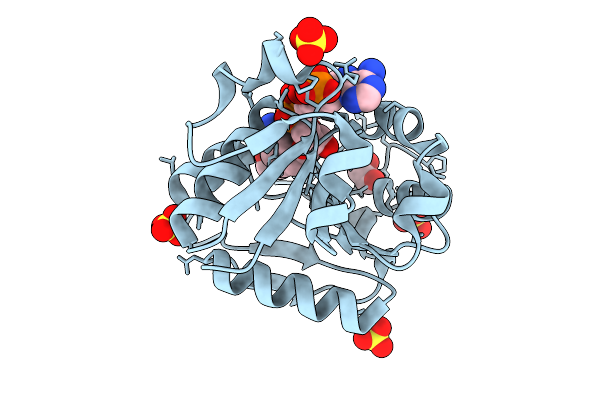 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2025-10-29 Classification: OXIDOREDUCTASE Ligands: NDP, EDO, SO4 |
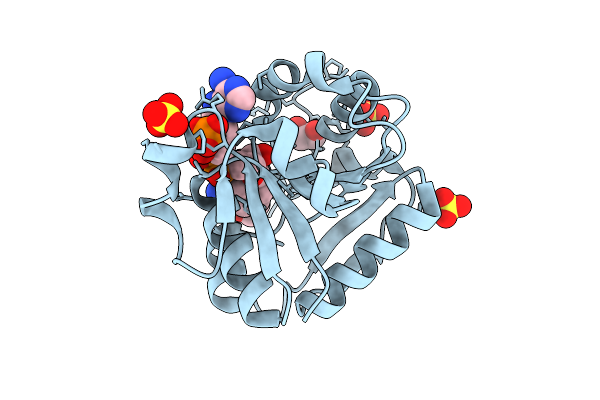 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2025-10-29 Classification: OXIDOREDUCTASE Ligands: EDO, NDP, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-10-29 Classification: OXIDOREDUCTASE Ligands: NDP, EDO, SO4 |
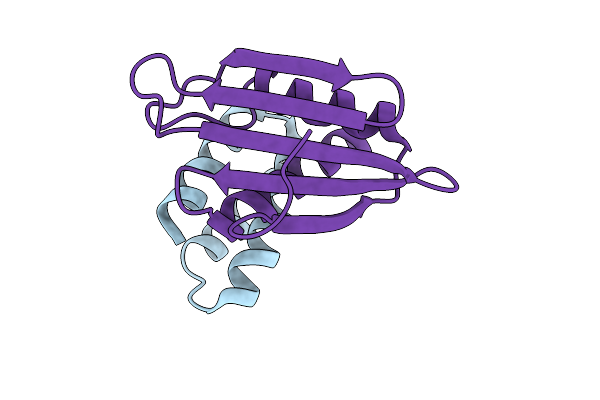 |
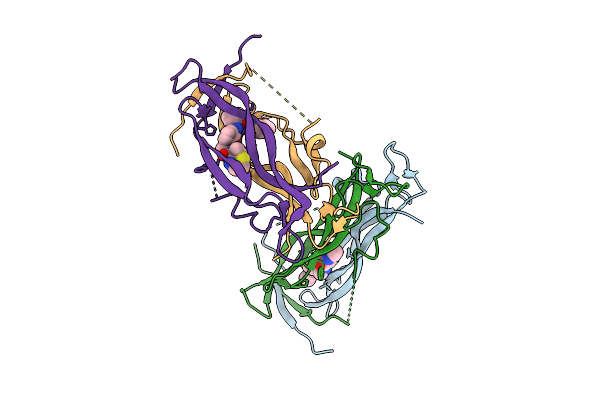
Organism: Francisella tularensis, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.37 Å Release Date: 2025-10-22 Classification: APOPTOSIS Ligands: IOD |
 |
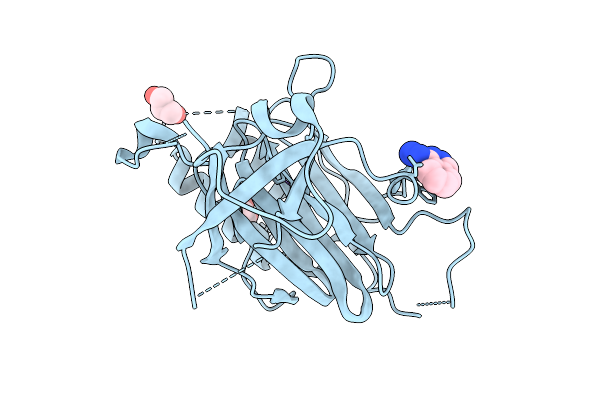
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2025-10-15 Classification: CYTOKINE Ligands: A1CXF |
 |
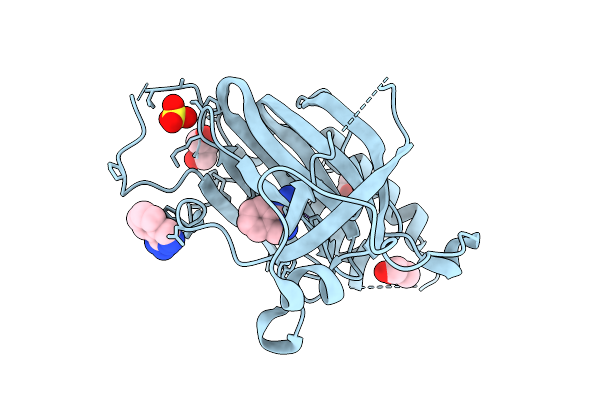
Crystal Structure Of Iron-Bound Human Ado C18S/C239S Variant Soaked In Hydralazine At 2.39 Angstrom Resolution
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2025-09-24 Classification: OXIDOREDUCTASE Ligands: FE2, HLZ, GOL |
 |
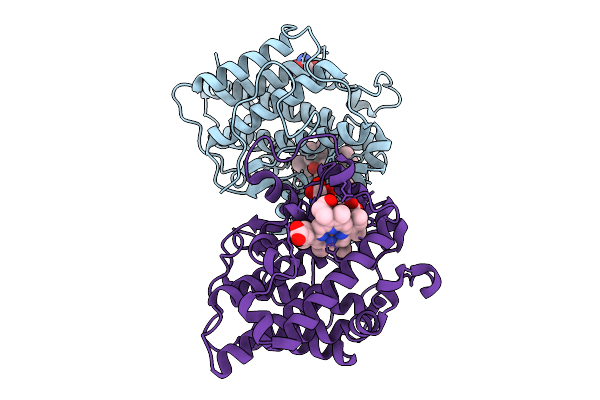
Crystal Structure Of Cobalt-Bound Human Ado C18S/C239S Variant In Complex With Hydralazine At 1.89 Angstrom Resolution
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-09-10 Classification: OXIDOREDUCTASE Ligands: CO, HLZ, GOL, SO4 |
 |
A Crystal Structure Of Heme-Dependent Tyrosine Hydroxylase Complexed With A Substrate Analog, 3-(4-Hydroxyphenyl)Propionic Acid
Organism: Streptomyces sclerotialus
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE/INHIBITOR Ligands: HPP, HEM, TRS |

