Search Count: 154
All
Selected
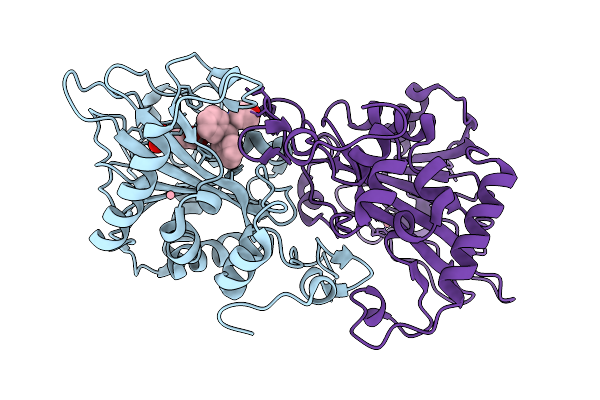 |
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: AKG, CO, A1EM0 |
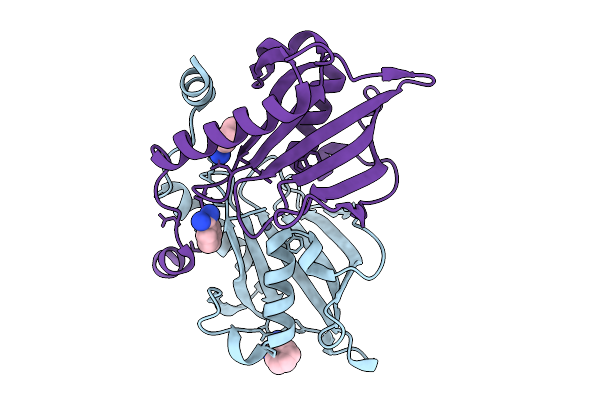 |
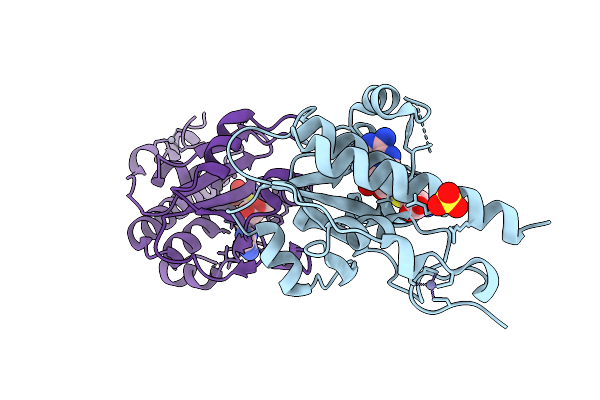
Crystal Structure Of Adp-Ribosylated Endolytic Muramidase Tdem From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa (strain atcc 15692 / dsm 22644 / cip 104116 / jcm 14847 / lmg 12228 / 1c / prs 101 / pao1)
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-21 Classification: DE NOVO PROTEIN Ligands: BEN, ZN |
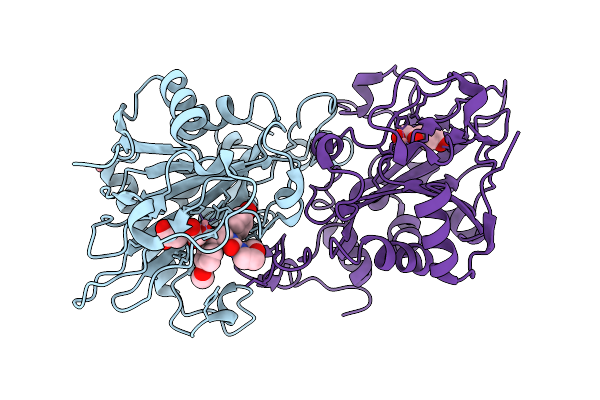 |
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EKG, AKG, CO |
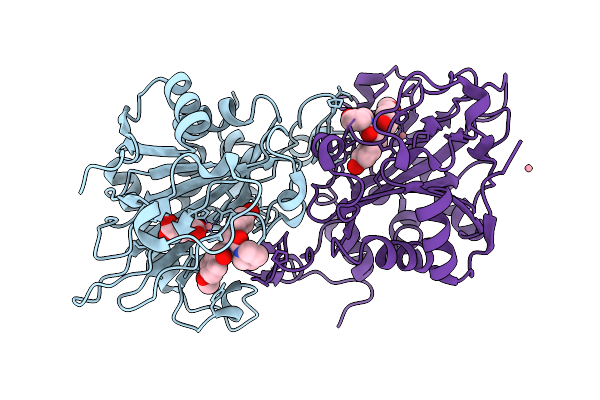 |
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:1.69 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EKF, SIN, CO |
 |
Organism: Escherichia coli (strain b / bl21-de3)
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-12-17 Classification: TRANSFERASE Ligands: SAM, ZN, SO4 |
 |
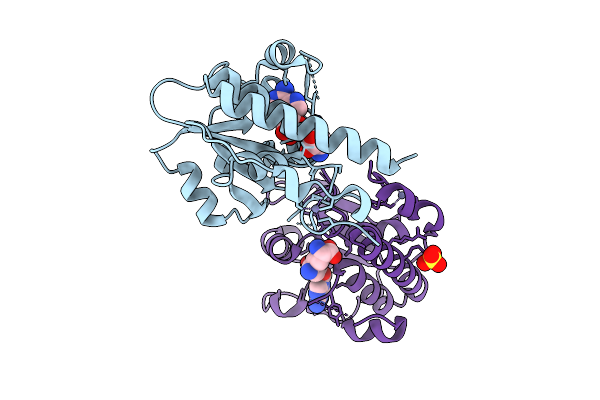
Organism: Escherichia coli (strain b / bl21-de3)
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2025-12-17 Classification: TRANSFERASE Ligands: MTA, ZN, SO4 |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2025-12-17 Classification: TRANSFERASE Ligands: SFG, ZN, SO4 |
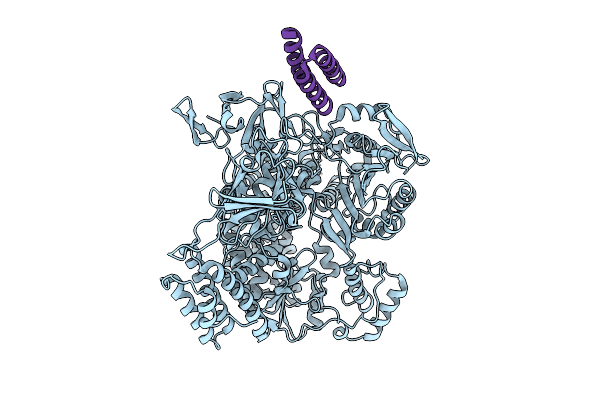 |
De Novo Designed Minibinder Complexed With Clostridioides Difficile Toxin B
Organism: Clostridioides difficile, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: TOXIN |
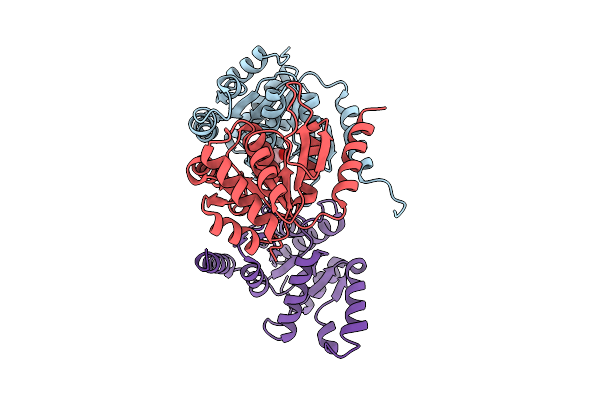 |
Organism: Neisseria meningitidis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-08-06 Classification: HYDROLASE Ligands: CIT |
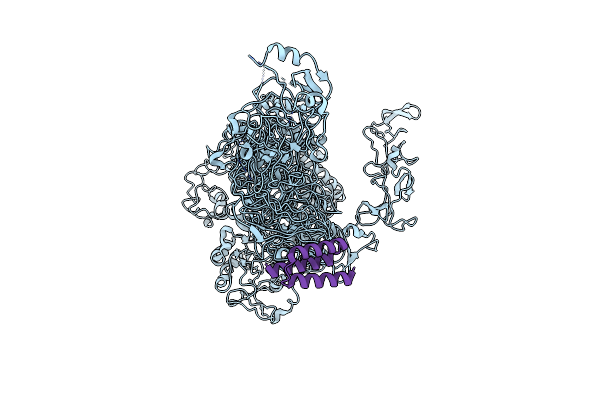 |
Organism: Synthetic construct, Clostridioides difficile
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: TOXIN |
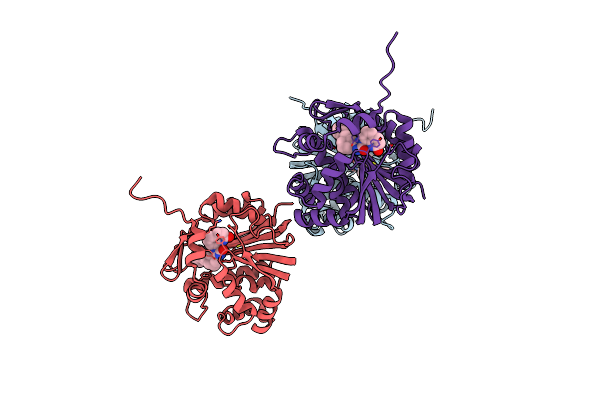 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2025-03-05 Classification: TRANSFERASE Ligands: A1AF6 |
 |
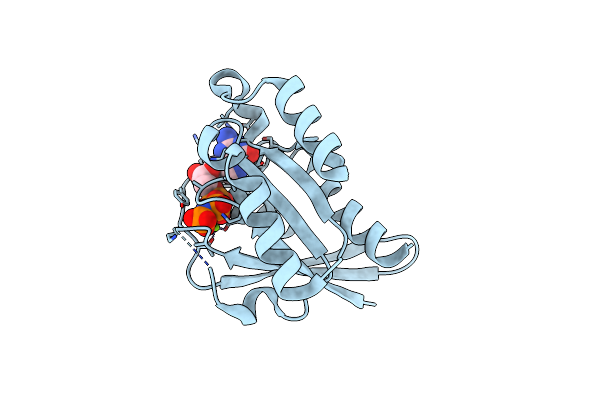
Crystal Structure Of Flt3 In Complex With A Pyrazinamide Macrocycle Derivative
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2024-12-11 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: ZX3 |
 |
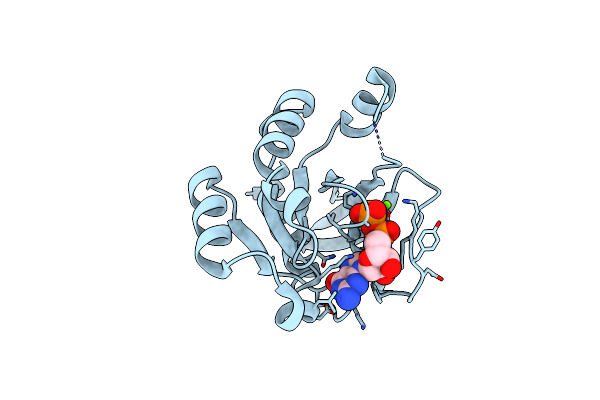
Crystal Structure Of Human Rab23 In Complex With Gmppnp (1.35 Angstroms Resolution)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2024-12-04 Classification: HYDROLASE Ligands: GNP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2024-12-04 Classification: HYDROLASE Ligands: GDP, MG |
 |
Crystal Structure Of Human Rab23 In Complex With Gmppnp (1.8 Angstroms Resolution)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-12-04 Classification: HYDROLASE Ligands: GNP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2024-12-04 Classification: HYDROLASE Ligands: GDP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2024-12-04 Classification: HYDROLASE Ligands: MG, GNP |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2024-12-04 Classification: METAL BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-27 Classification: METAL BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-27 Classification: METAL BINDING PROTEIN Ligands: CU |

