Search Count: 803
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: CYTOSOLIC PROTEIN |
 |
Crystal Structure Of Rubredoxin From Piezophilic Hyperthermophilic Archaeon Pyrococcus Yayanosii
Organism: Pyrococcus yayanosii ch1
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2026-05-06 Classification: ELECTRON TRANSPORT Ligands: FE, NA |
 |
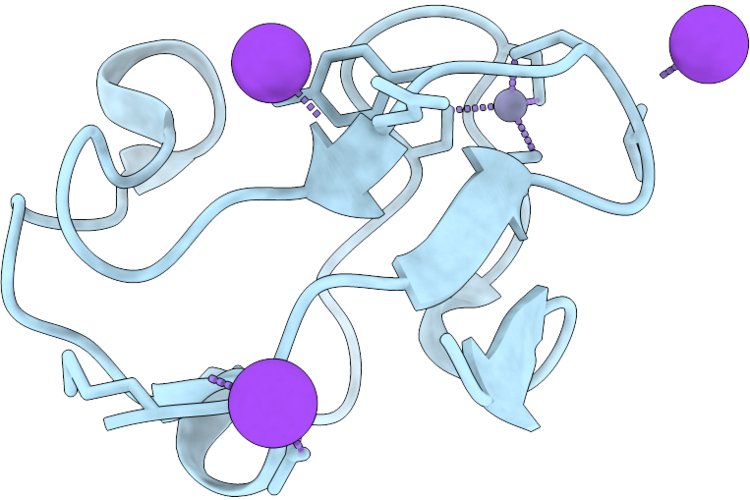
Crystal Structure Of Zn-Substituted Rubredoxin From Hyperthermophilic Bacterium Thermotoga Maritima Msb8
Organism: Thermotoga maritima msb8
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-05-06 Classification: ELECTRON TRANSPORT Ligands: ZN |
 |
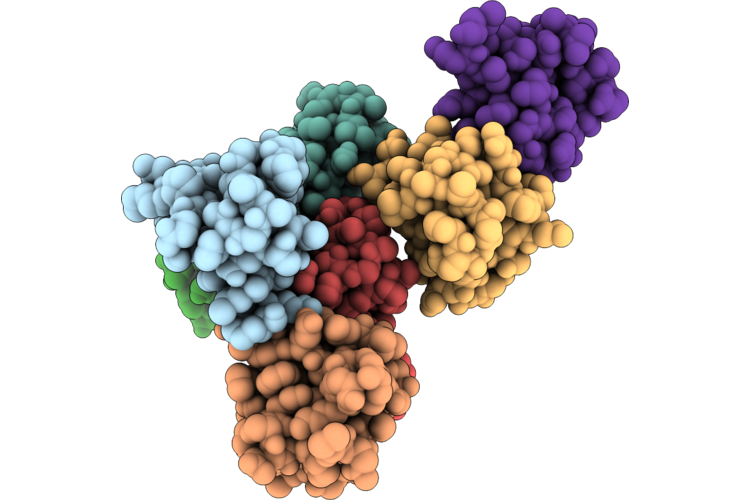
Crystal Structure Of Zn-Substituted Rubredoxin From Psychrophilic Clostridia Clostridium Psychrophilum
Organism: Clostridium psychrophilum
Method: X-RAY DIFFRACTION Resolution:0.84 Å Release Date: 2026-05-06 Classification: ELECTRON TRANSPORT Ligands: ZN, K |
 |
Crystal Structure Of Rubredoxin From Psychrophilic Bacterium Polaromonas Glacialis
Organism: Polaromonas glacialis
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-05-06 Classification: ELECTRON TRANSPORT Ligands: FE, NA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: CYTOSOLIC PROTEIN |
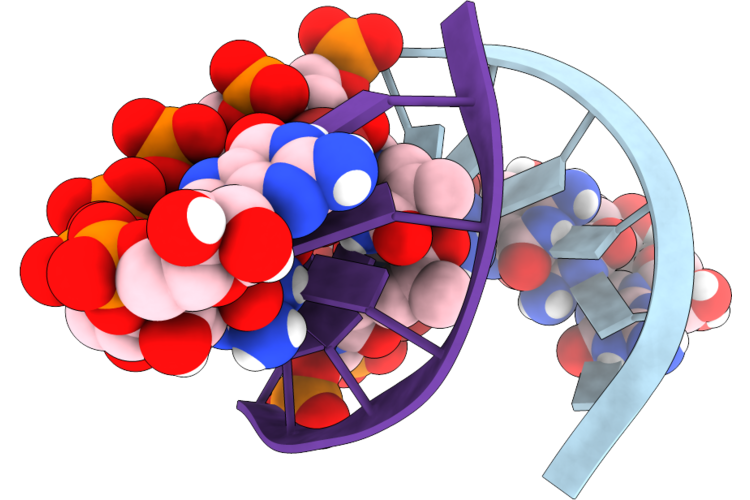 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-04-22 Classification: RNA Ligands: A1B9T, 5BU, 5GP, C5P, C |
 |
Cryo-Em Structure Of The Vo Domain Of V/A-Atpase In Liposomes Under No Pmf Condition,State2
Organism: Thermus thermophilus hb8
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MOTOR PROTEIN |
 |
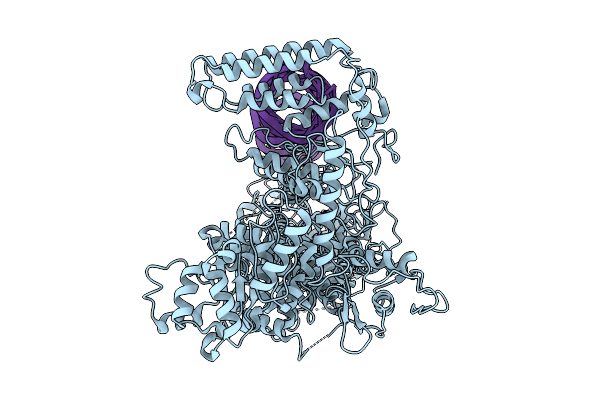
Cryo-Em Structure Of The Vo Domain Of V/A-Atpase In Liposomes Under No Pmf Condition,State3
Organism: Thermus thermophilus hb8
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MOTOR PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-03-11 Classification: HYDROLASE/RNA |
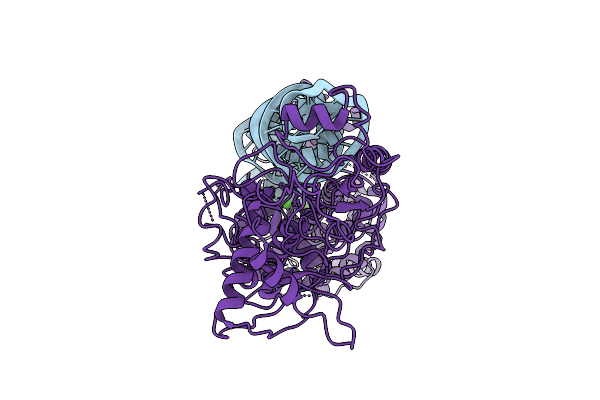 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.21 Å Release Date: 2026-03-11 Classification: HYDROLASE/RNA Ligands: CA |
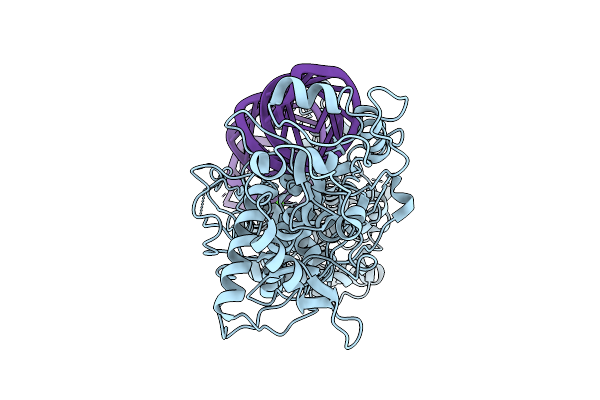 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-03-11 Classification: HYDROLASE/RNA Ligands: CA |
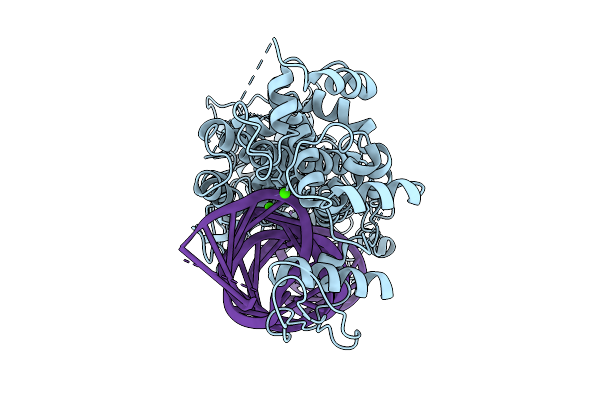 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: HYDROLASE Ligands: CA |
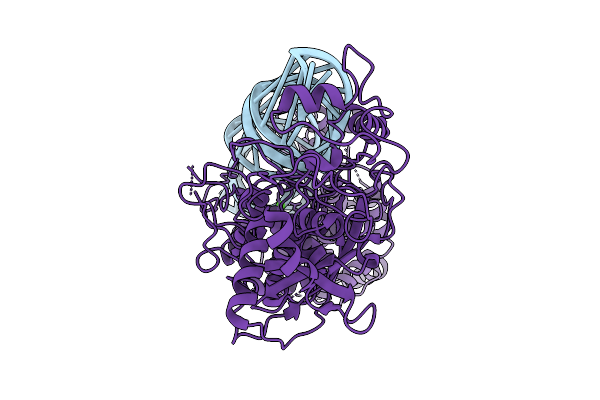 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.34 Å Release Date: 2026-03-11 Classification: HYDROLASE Ligands: CA |
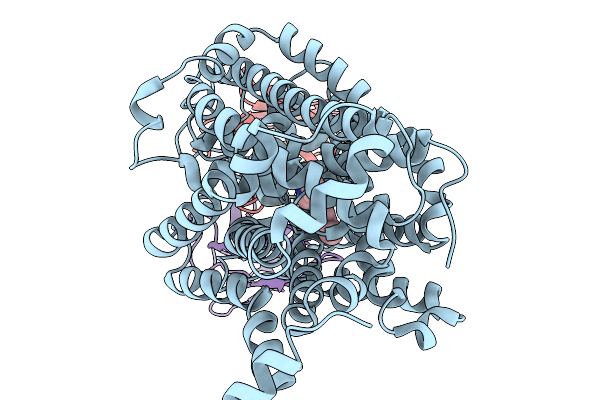 |
Structure Of Human Serotonin Transporter Bound To Small Molecule Zpzd In Lipid Nanodisc And Nacl
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN/IMMUNE SYSTEM/INHIBITOR Ligands: CL, A1CIZ |
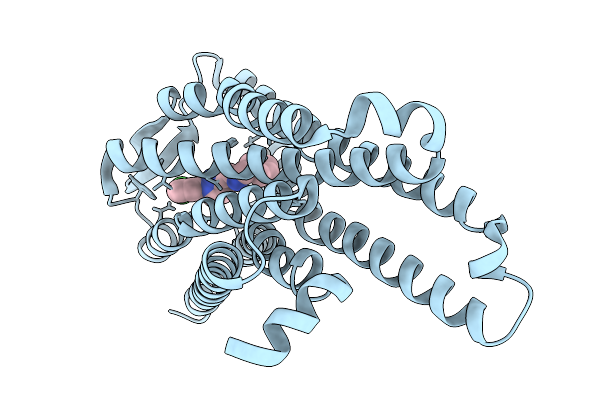 |
Locally-Refined Mu-Opioid Receptor Bound With Novel Compound 0505 (3-[({[(1P)-1-(3-Chlorophenyl)-1H-Pyrazol-3-Yl]Methyl}Amino)Methyl]Phenol)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: A1CIX |
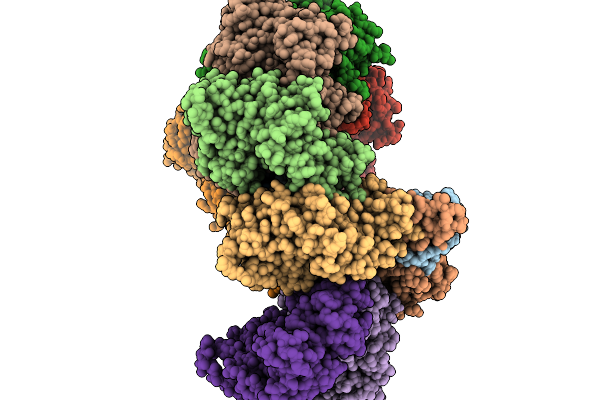 |
Target Dna-Bound Type I-F3 Tniq-Cascade Of Vibrio Parahaemolyticus In Partial R-Loop State
Organism: Vibrio parahaemolyticus rimd 2210633, Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Resolution:2.68 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: MG |
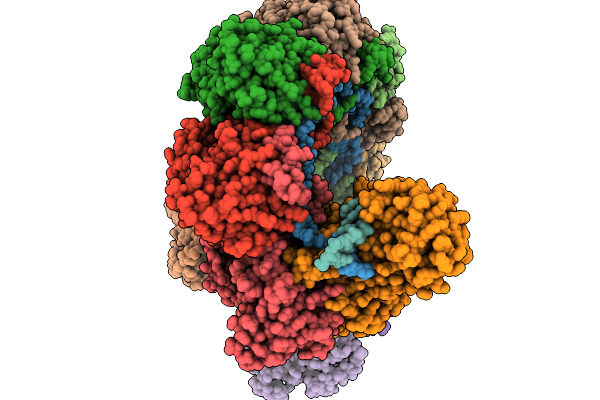 |
Target Dna-Bound Type I-F3 Tniq-Cascade Of Vibrio Parahaemolyticus In Full R-Loop State 1
Organism: Vibrio parahaemolyticus rimd 2210633
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: MG |
 |
Target Dna-Bound Type I-F3 Tniq-Cascade Of Vibrio Parahaemolyticus In Full R-Loop State 2
Organism: Vibrio parahaemolyticus rimd 2210633
Method: ELECTRON MICROSCOPY Resolution:2.89 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN Ligands: MG |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Amppnp From Thermococcus Kodakarensis
Organism: Thermococcus kodakarensis
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-02-04 Classification: TRANSLOCASE Ligands: ANP |

