Search Count: 2,901
All
Selected
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PEG, DSN, CA |
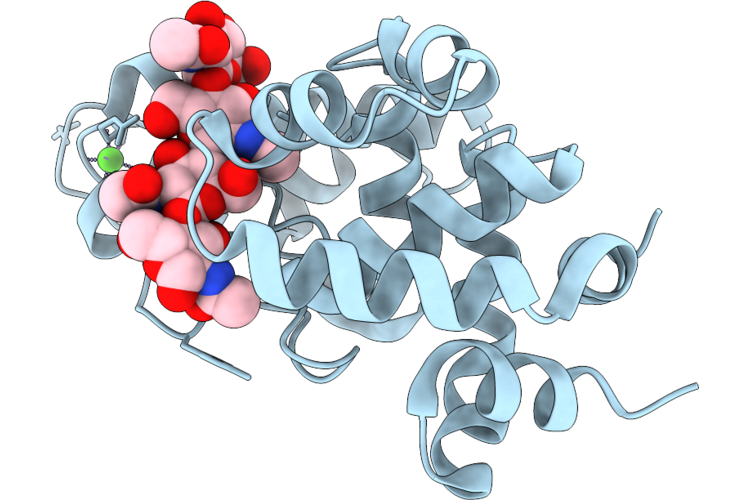 |
Crystal Structure Of Gh19 E228Q Domain Of D29-Lysa In Complex With (Glcnac)4
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CA |
 |
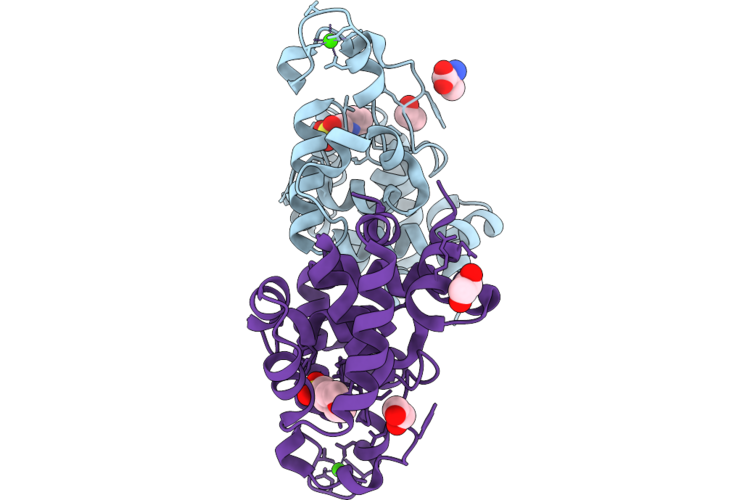
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, MPD |
 |
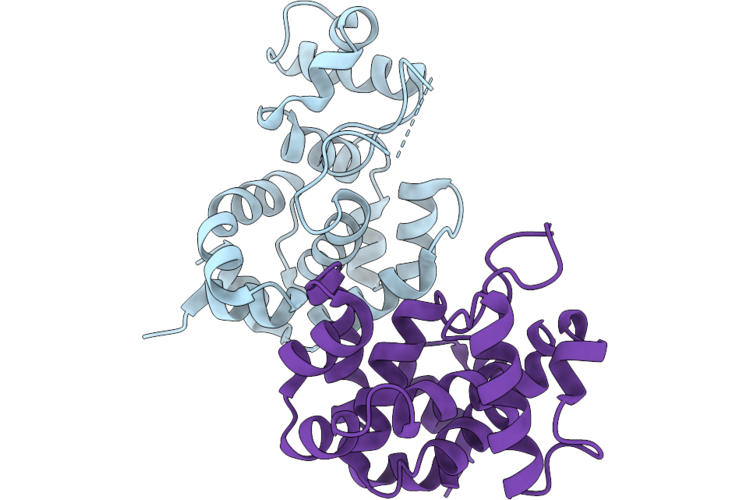
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, EDO, ALA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
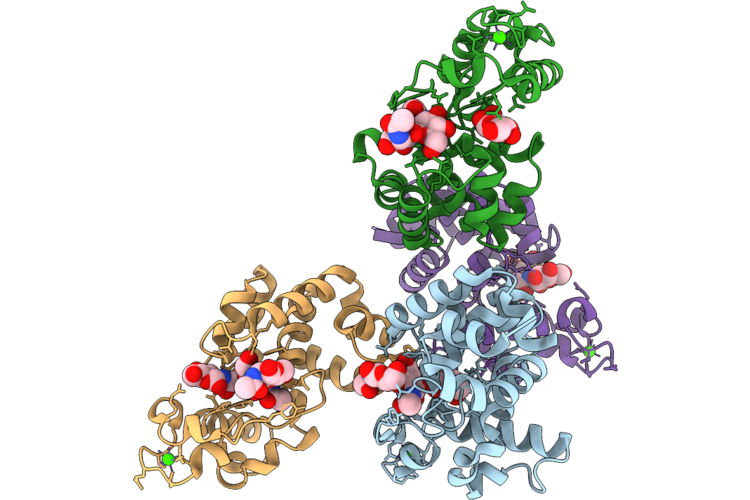
Crystal Structure Of Gh19 Domain Of D29-Lysa In Complex With Product (Glcnac)
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: NAG |
 |
Crystal Structure Of Gh19 Domain Of D29-Lysa In Complex With Products (Glcnac Y Glcnac2)
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: NAG, CA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Organism: Gallus gallus
Method: ELECTRON CRYSTALLOGRAPHY Resolution:0.83 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ACT, CL |
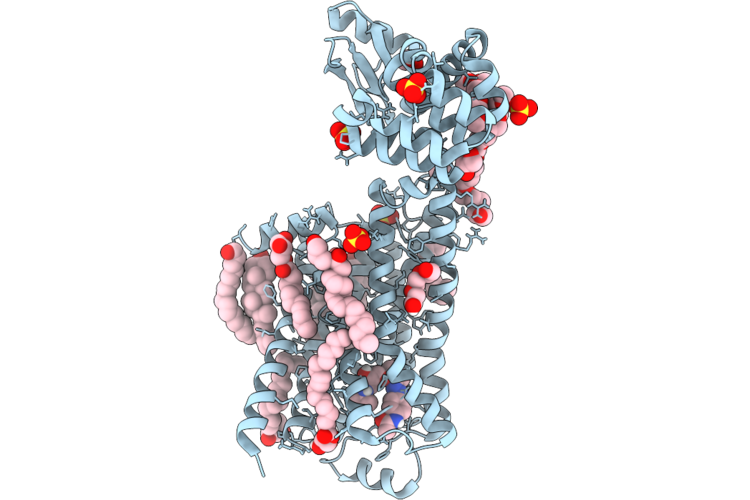 |
Dark Structure Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State Recorded At Swissfel
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, CLR, 12P, OLC, 1PE, EDO, GOL, A1JHU, PLM |
 |
Mixed Model Refinement Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State And Light State, 10 Seconds After Light Activation, Recorded At Swissfel
Organism: Homo sapiens, Tequatrovirus t4
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, ACM, CLR, PLM, 12P, OLC, 1PE, EDO, GOL, A1JHU |
 |
Dark Structure Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State Recorded At Lcls
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, CLR, PLM, 12P, A1JHU, OLC, EDO, GOL |
 |
Mixed Model Refinement Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State And Light State, 17 Nanoseconds After Light Activation, Recorded At Lcls
Organism: Homo sapiens, Tequatrovirus t4
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, ACM, CLR, PLM, 12P, OLC, EDO, GOL, 1PE, A1JHU |
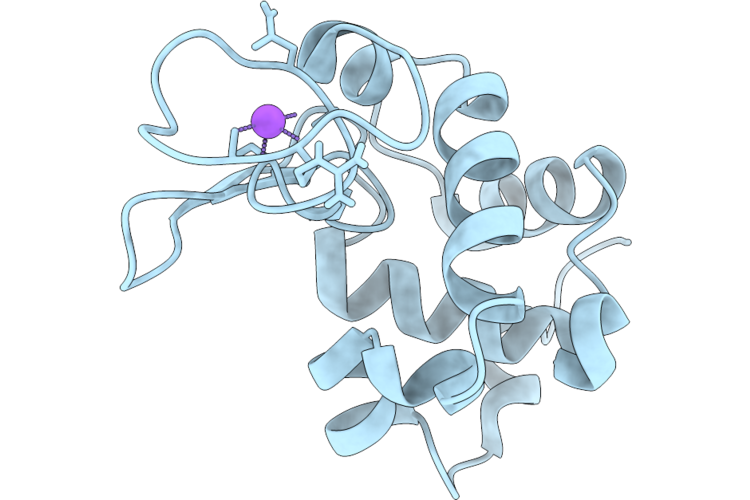 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CL, NA |
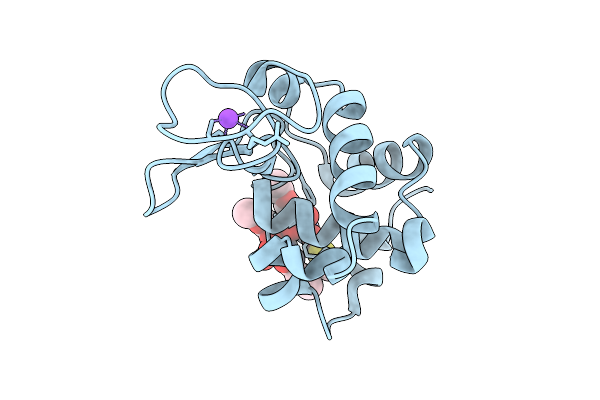 |
X-Ray Structure Of Lysozyme Obtained Upon Reaction With The Dioxidovanadium(V) Complex With Thiophene-2-Carboxylic Acid (3-Ethoxy-2-Hydroxybenzylidene)Hydrazide
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-04-01 Classification: HYDROLASE |
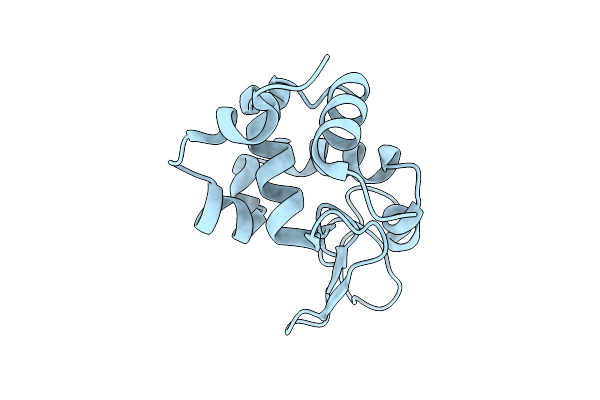 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: CL |
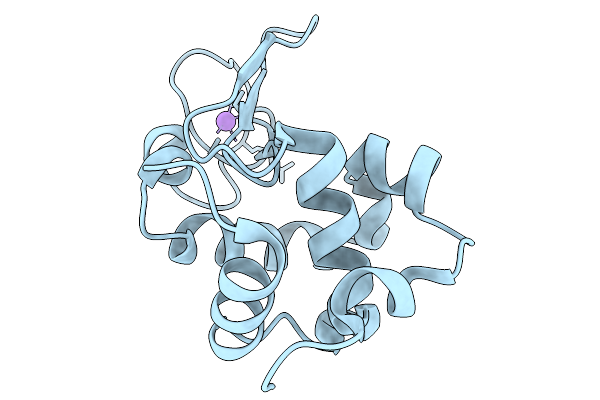 |
In Situ Structure Of Egg-White Lysozyme Using A Goniometer-Compatible Chip-Based Platform
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-03-25 Classification: HYDROLASE |
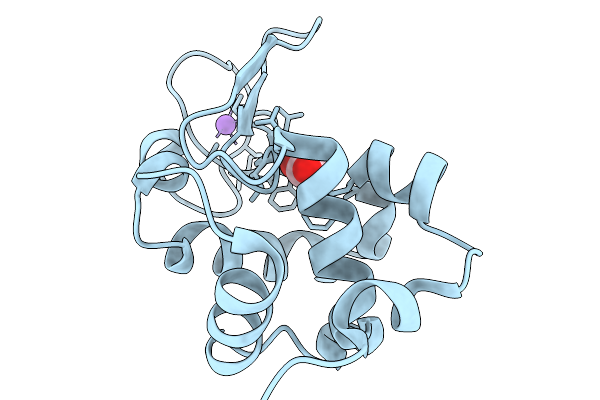 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-25 Classification: HYDROLASE |
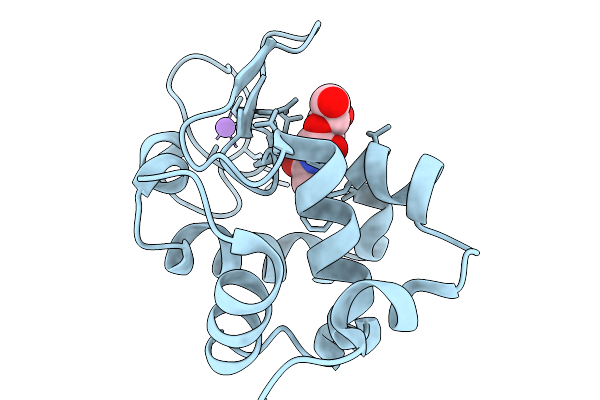 |
Time-Resolved Mixing Crystallography Structure Of 3-5 Micrometers Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 2 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |

