Search Count: 55,415
All
Selected
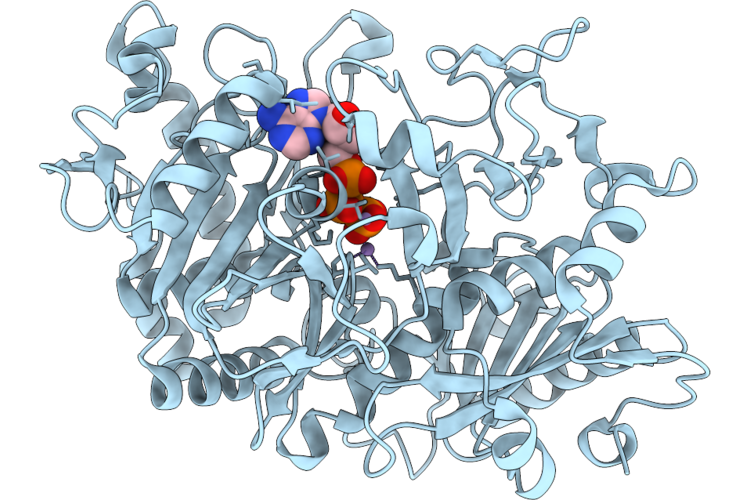 |
Structure Of Escherichia Coli Phosphoenolpyruvate Carboxykinase With Bound Atp, Mg2+, And Mn2+
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-04-29 Classification: LYASE Ligands: ATP, MN, MG |
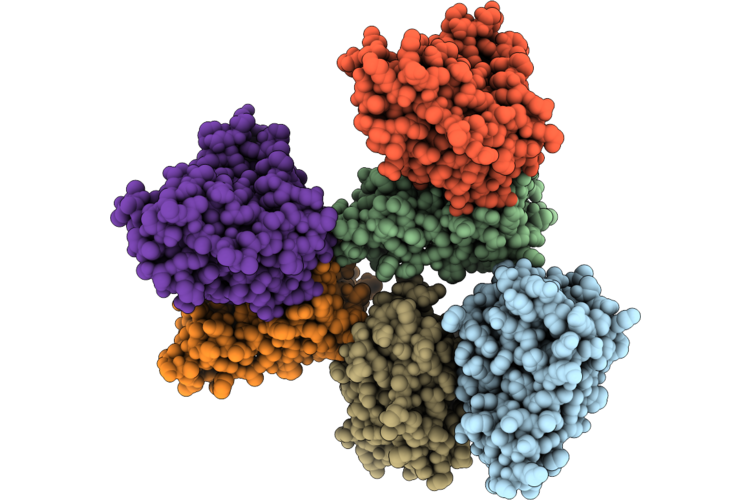 |
Organism: Cryptosporidium parvum iowa ii
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: MN, EDO, PEG |
 |
Organism: Ogataea methanolica
Method: ELECTRON MICROSCOPY Resolution:1.93 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: FAS |
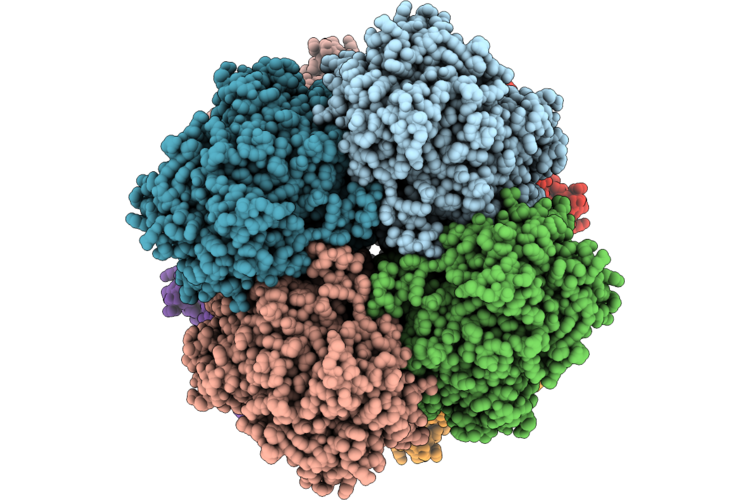 |
Organism: Ogataea methanolica
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: FAD |
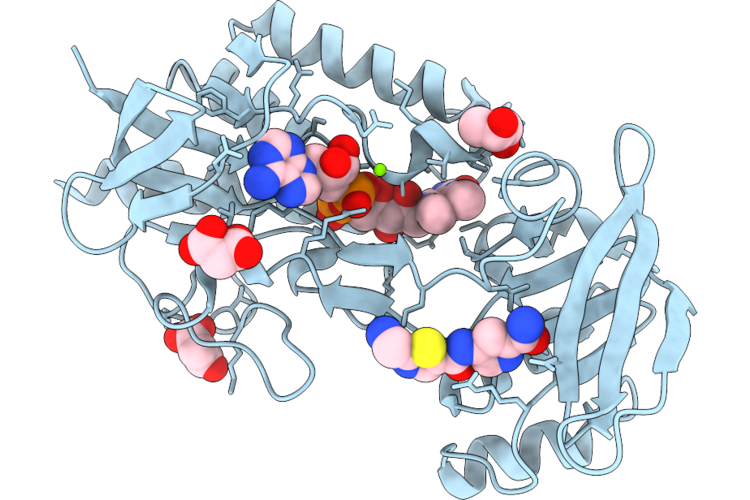 |
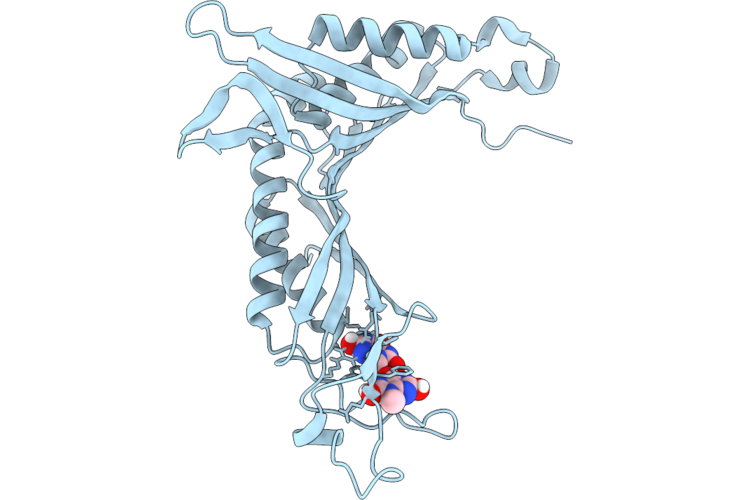
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z4666192264
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1JAA |
 |
Urate Oxidase From Aspergillus Flavus With Its Inhibitor 9-Methyl Uric Acid By Continuous Serial Electron Diffraction (Serialed)
Organism: Aspergillus flavus
Method: ELECTRON CRYSTALLOGRAPHY Resolution:1.25 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: MUA |
 |
Urate Oxidase From Aspergillus Flavus With Its Substrate Uric Acid By Continuous Serial Electron Diffraction (Serialed)
Organism: Aspergillus flavus
Method: ELECTRON CRYSTALLOGRAPHY Resolution:1.75 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: OXY, URC |
 |
Crystal Structure Of Catalytic Domain Of Rv1625C Bound To Inhibitory Darpin G11
Organism: Mycobacterium tuberculosis, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-04-29 Classification: SIGNALING PROTEIN Ligands: GOL |
 |
Crystal Structure Of An Nta622L Variant In Complex With Nadph And Trigonelline
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-29 Classification: PLANT PROTEIN Ligands: NDP, A1JBP |
 |
Crystal Structure Of Human L-Lactate Dehydrogenase B Protein In Complex With Nadh, Oxamate And Sertraline
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: OXM, NAI, SRE |
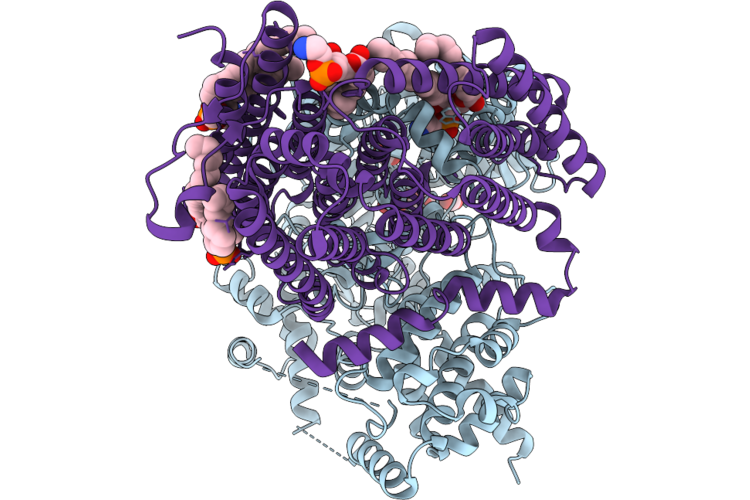 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
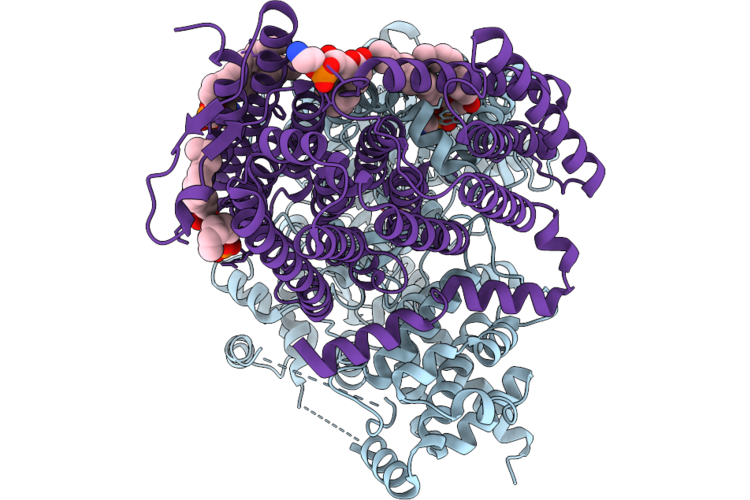 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, BCT, CO2, 6OU |
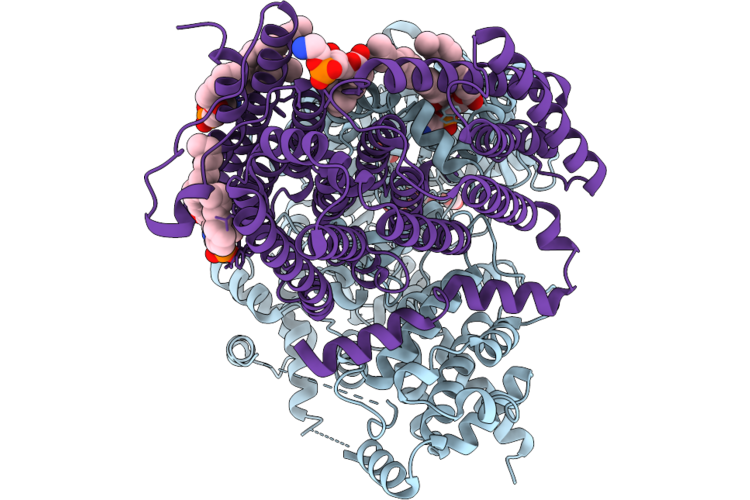 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
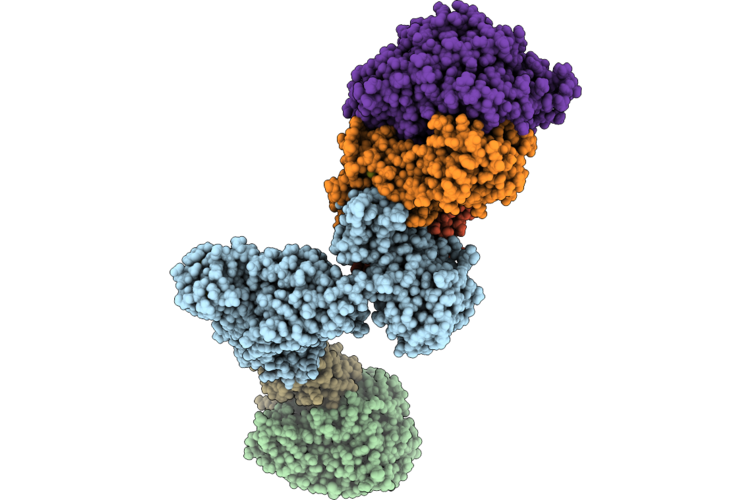 |
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 2 Substate A (Composite Structure)
Organism: Methanothermobacter marburgensis str. marburg
Method: ELECTRON MICROSCOPY Resolution:2.19 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, NFU |
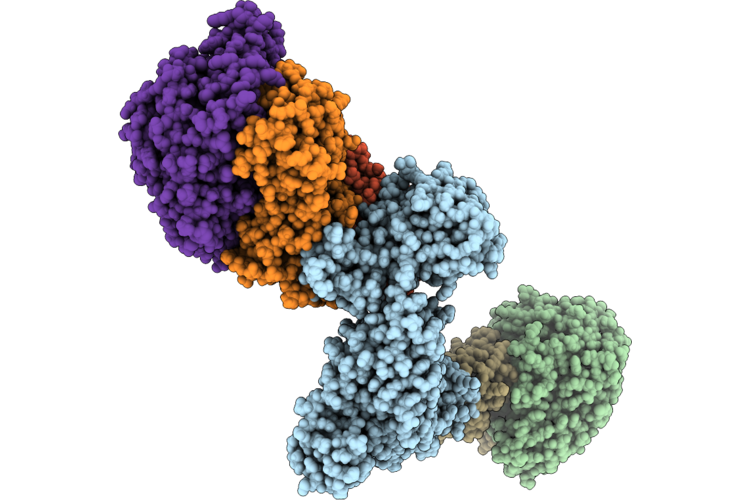 |
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 2 Substate B (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:3.25 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, NFU |
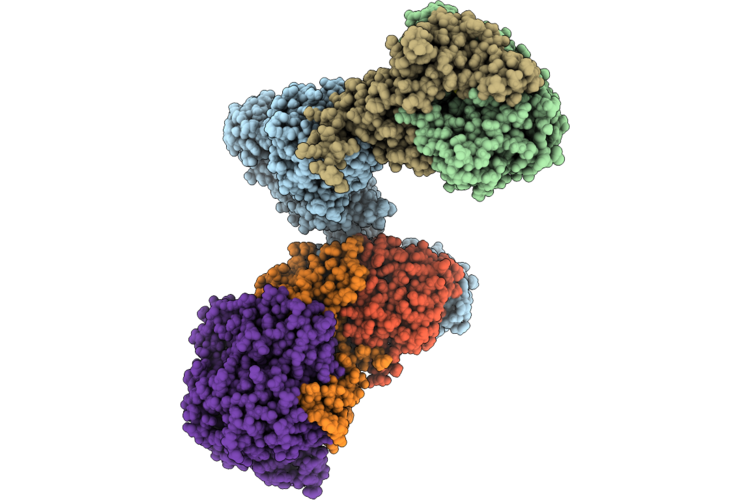 |
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 1 Substate A (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, FES, 9S8, NFU |
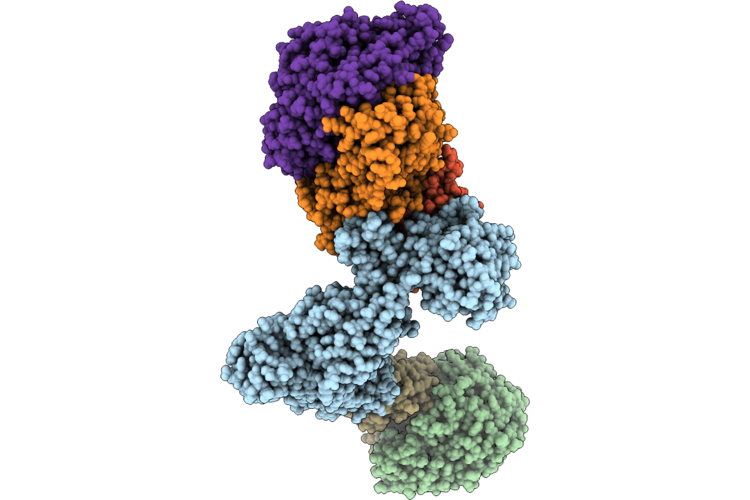 |
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 1 Substate B (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, NFU, FES |
 |
Structure Of The Mvh-Hdr-Fmd Complex Of Methanothermobacter Marburgensis (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, ZN, MGD, MO, H2S, F3S, NFU |
 |
Organism: Glycyrrhiza uralensis
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: NAP, A1EP1 |
 |
Crystal Structure Of The V165A/S219E/A225P Mutant Of Alanine Dehyrogenase From Geobacillus Stearothermophilus In Complex With Nicotinamide Cytosine Dinucleotide
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: A1EP0 |

