Search Count: 99
All
Selected
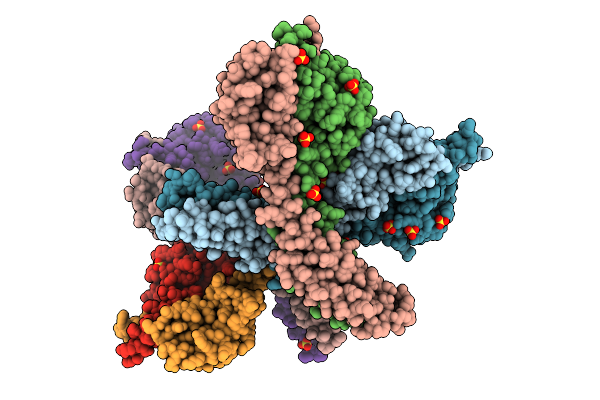 |
Crystal Structure Of The Histidine Kinase Vc2136 From Vibrio Cholerae Serotype O1
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-08-20 Classification: TRANSFERASE Ligands: SO4, CL |
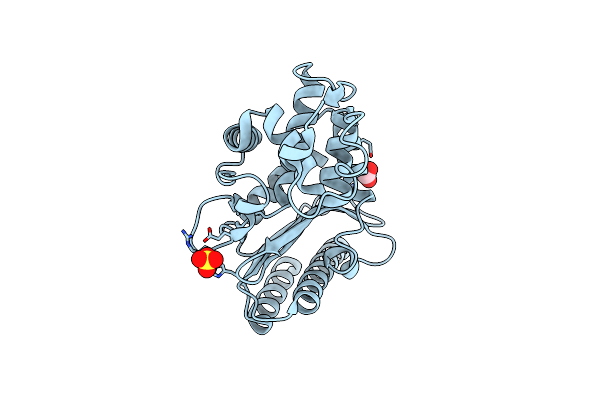 |
High Resolution Structure Of Class A Beta-Lactamase From Bordetella Bronchiseptica Rb50
Organism: Bordetella bronchiseptica rb50
Method: X-RAY DIFFRACTION Resolution:1.05 Å Release Date: 2024-06-12 Classification: HYDROLASE Ligands: FMT, SO4 |
 |
Structure Of Class A Beta-Lactamase From Bordetella Bronchiseptica Rb50 In A Complex With Avibactam
Organism: Bordetella bronchiseptica rb50
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2024-06-12 Classification: HYDROLASE Ligands: NXL, SIN, FMT, EDO |
 |
Structure Of Class A Beta-Lactamase From Bordetella Bronchiseptica Rb50 In A Complex With Clavulonate
Organism: Bordetella bronchiseptica rb50
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2024-06-12 Classification: HYDROLASE Ligands: MLA, TEM, CL |
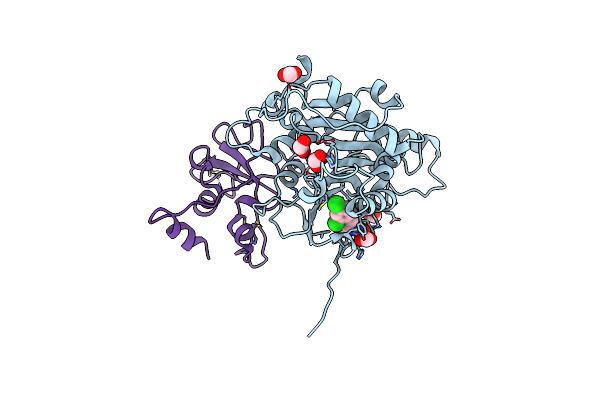 |
Crystal Structure Of The Sars-Cov-2 2'-O-Methyltransferase With Compound 5A Bound To The Cryptic Pocket Of Nsp16
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2023-10-18 Classification: TRANSFERASE |
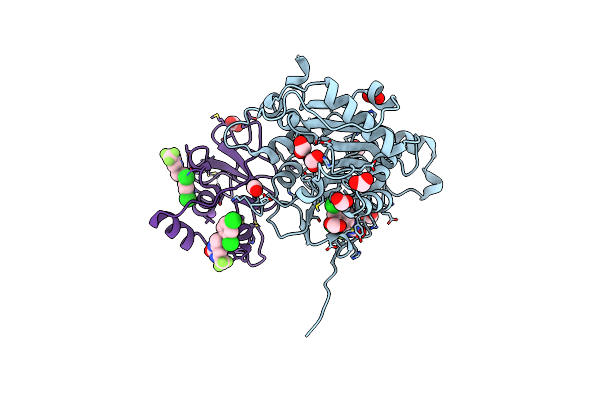 |
Crystal Structure Of Sars-Cov-2 2'-O-Methyltransferase In Complex With Compound 5A Covalently Bound To Nsp16 And Nsp10
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2023-10-18 Classification: TRANSFERASE |
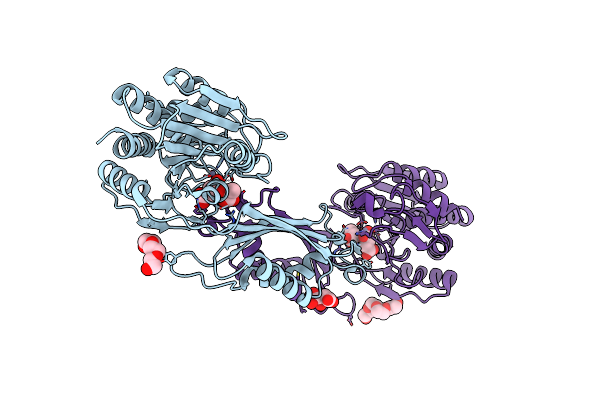 |
Crystal Structure Of The Succinyl-Diaminopimelate Desuccinylase (Dape) From Acinetobacter Baumannii In Complex With Succinic And L-Lactic Acids
Organism: Acinetobacter baumannii atcc 17978
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-11-30 Classification: HYDROLASE Ligands: ZN, SIN, 2OP, PGE, PG4, CIT |
 |
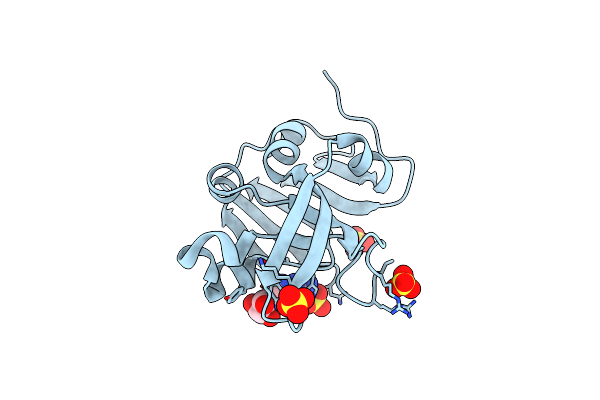
Crystal Structure Of The Putative Bacteriophage Protein From Stenotrophomonas Maltophilia
Organism: Stenotrophomonas maltophilia (strain k279a)
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-11-09 Classification: UNKNOWN FUNCTION Ligands: SO4 |
 |
Crystal Structure Of Putative Pterin Binding Protein (Prur) From Klebsiella Pneumoniae In Complex With Neopterin
Organism: Klebsiella pneumoniae subsp. pneumoniae sa1
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2022-08-10 Classification: PTERIN BINDING PROTEIN Ligands: NEU, CL, SO4 |
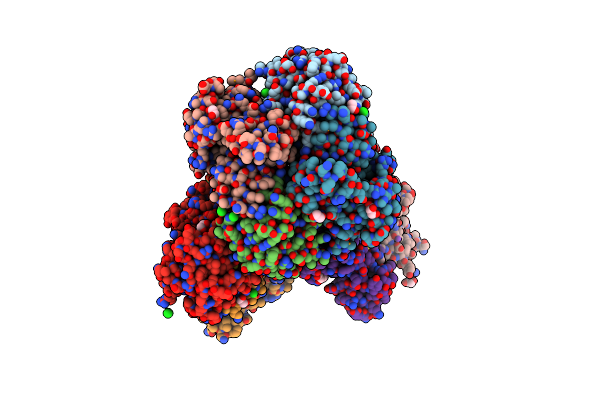 |
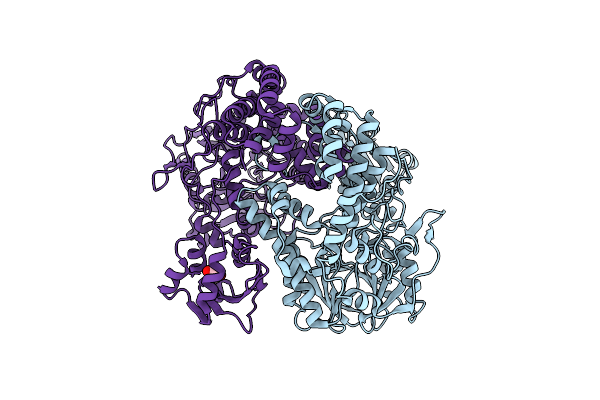
Crystal Structure Of Putataive Short-Chain Dehydrogenase/Reductase (Fabg) From Klebsiella Pneumoniae Subsp. Pneumoniae Ntuh-K2044 In Complex With Nadh
Organism: Klebsiella pneumoniae subsp. pneumoniae ntuh-k2044
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2022-03-02 Classification: BIOSYNTHETIC PROTEIN Ligands: K, CL, NAI, GOL, EDO |
 |
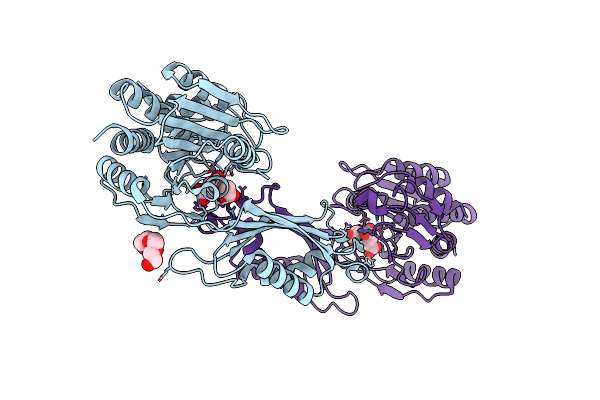
Organism: Klebsiella pneumoniae subsp. pneumoniae ntuh-k2044
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: EDO |
 |
Crystal Structure Of The Succinyl-Diaminopimelate Desuccinylase (Dape) From Acinetobacter Baumannii In Complex With Succinic Acid
Organism: Acinetobacter baumannii atcc 17978
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2021-12-15 Classification: HYDROLASE Ligands: SIN, ACT, PGE, ZN |
 |
Crystal Structure Of The Putative Hydrolase From Stenotrophomonas Maltophilia
Organism: Stenotrophomonas maltophilia (strain k279a)
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2021-12-01 Classification: HYDROLASE Ligands: CL, NA, FMT |
 |
Crystal Structure Of The Peptidyl-Prolyl Cis-Trans Isomerase (Ppib) From Streptococcus Pyogenes.
Organism: Streptococcus pyogenes mgas5005
Method: X-RAY DIFFRACTION Resolution:1.28 Å Release Date: 2021-12-01 Classification: ISOMERASE |
 |
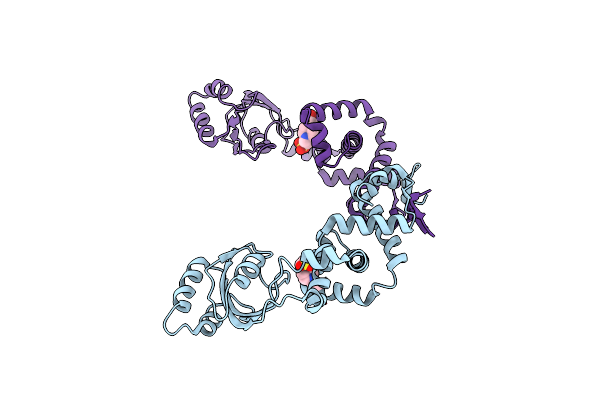
Crystal Structure Of Peptidyl-Prolyl Cis-Trans Isomerasefrom (Ppib) Streptococcus Pneumoniae R6
Organism: Streptococcus pneumoniae (strain atcc baa-255 / r6)
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2021-12-01 Classification: ISOMERASE Ligands: CL, EDO, MES |
 |
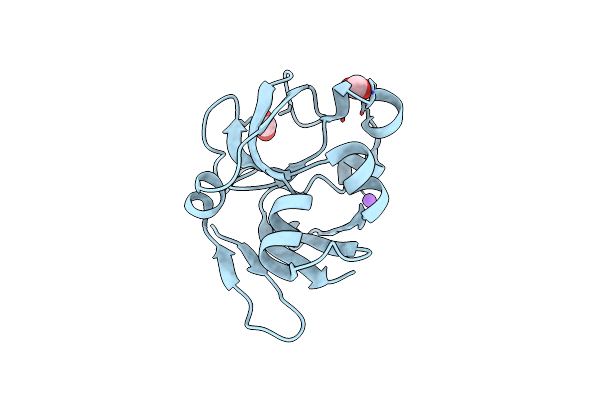
Crystal Structure Of Peptidylprolyl Isomerase Prsa From Streptococcus Mutans.
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2021-12-01 Classification: ISOMERASE Ligands: EPE, CL |
 |
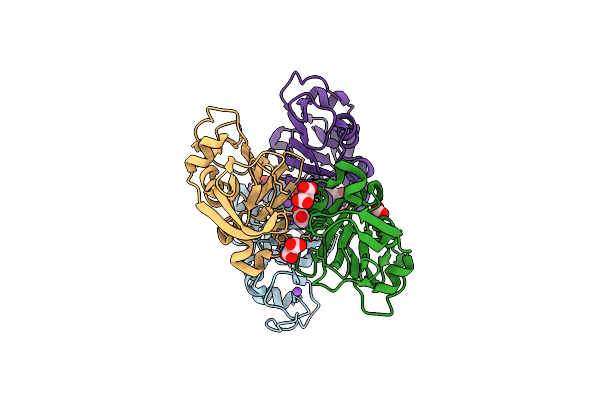
High Resolution Crystal Structure Of Putative Pterin Binding Protein Prur (Vv2_1280) From Vibrio Vulnificus Cmcp6
Organism: Vibrio vulnificus (strain cmcp6)
Method: X-RAY DIFFRACTION Resolution:0.99 Å Release Date: 2021-11-17 Classification: Pterin binding protein Ligands: FMT, NA |
 |
1.50 Angstroms Resolution Crystal Structure Of Putative Pterin Binding Protein Prur (Atu3496) From Agrobacterium Fabrum Str. C58
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2021-11-17 Classification: Pterin binding protein |
 |
1.83 Angstroms Resolution Crystal Structure Of Putative Pterin Binding Protein Prur (Atu3496) From Agrobacterium Fabrum Str. C58
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2021-11-17 Classification: Pterin binding protein Ligands: CL, NA, PG4, PEG |
 |
High Resolution Crystal Structure Of Putative Pterin Binding Protein (Prur) From Vibrio Cholerae O1 Biovar El Tor Str. N16961 In Complex With Neopterin
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2021-11-17 Classification: PTERIN-BINDING PROTEIN Ligands: NEU |

