Search Count: 1,138
All
Selected
 |
Androgen Receptor Ligand-Binding Domain In Complex With 11-Ketodihydrotestosterone
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: A1IQ9, SPD, SO4, GOL, IMD |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And Imidazole
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: IMD, MN |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 2,5,6-Triaminopyrimidin-4(3H)-One.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: A1L8V, IMD, MN |
 |
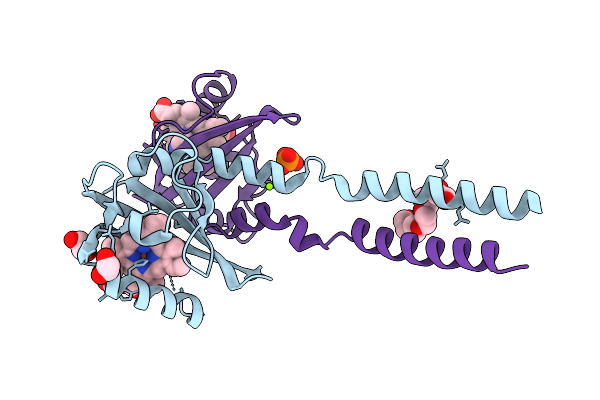
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: GOL, A1L8W, MN, IMD |
 |
Crystal Structure Of Heme Binding Pas Domain From One Component Transcription Factor, Fg214
Organism: Fimbriimonas ginsengisoli gsoil 348
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-25 Classification: TRANSCRIPTION Ligands: HEM, IMD, PO4, CL, PG4, EDO, MG, FMT, POL |
 |
Organism: Rattus norvegicus, Gallus gallus, Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-03-25 Classification: CELL CYCLE Ligands: GTP, MG, GOL, CA, CL, GDP, MES, A1I6L, IMD, ACP |
 |
Scia(H42Q) Endonuclease From Escherichia Phage T5 In Complex With 10 Bp Dna (Gcgc Sequence 2)
Organism: Escherichia phage t5
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN Ligands: IMD, ZN |
 |
Scia(H42A) Endonuclease From Escherichia Phage T5 In Complex With 10 Bp Dna (Gcgc Sequence 1)
Organism: Escherichia phage t5
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN Ligands: IMD, ACT, ZN |
 |
Crystal Structure Of Mctb From Mycobacterium Tuberculosis At 2.3 Angstroms Resolution
Organism: Mycobacterium tuberculosis cdc1551
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: GOL, IMD |
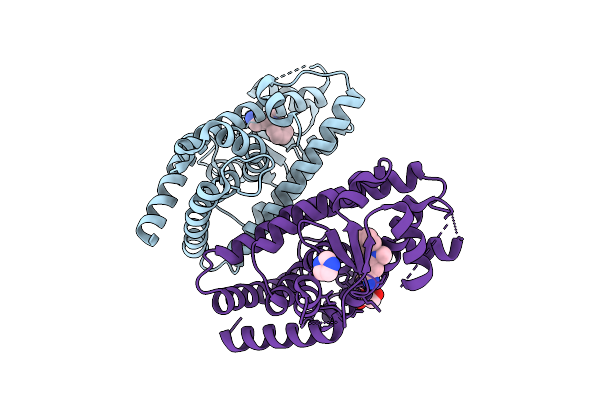 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-18 Classification: TRANSCRIPTION Ligands: A1ENO, IMD, GOL |
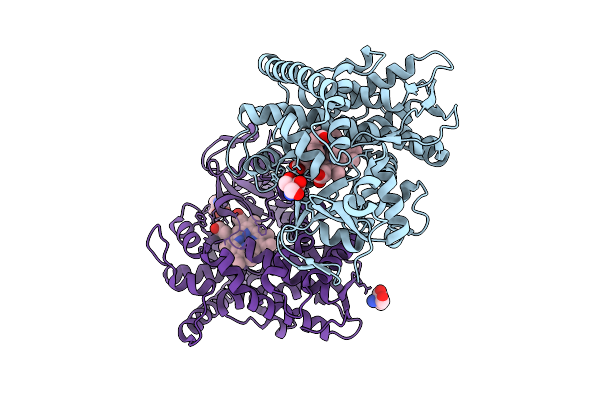 |
Organism: Priestia megaterium (strain nbrc 15308)
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: HEM, IMD, PEG, TRS, CL |
 |
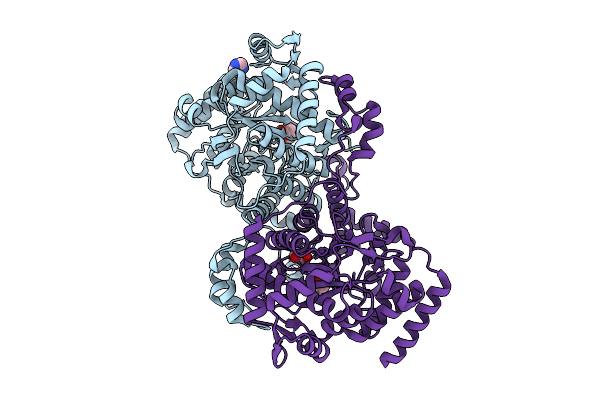
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN Ligands: GOL, EDO, IMD |
 |
Active Site Of Mtgb, A Glycine Betaine Methyltransferase From The Mttb Superfamily
Organism: Desulfitobacterium hafniense y51
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: IMD, BET, EDO |
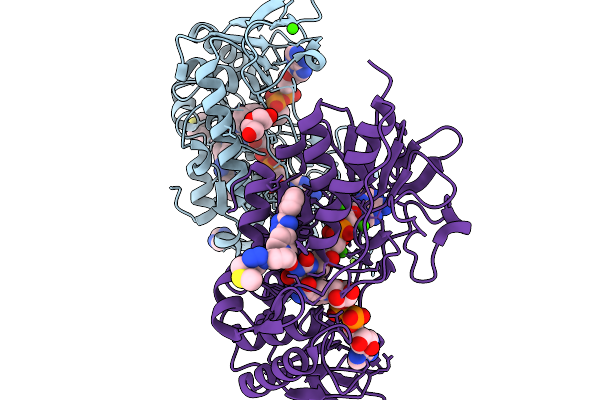 |
Organism: Mammalia
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: FAD, NAD, A1I3Z, IMD, PEG, CA, CL, MG |
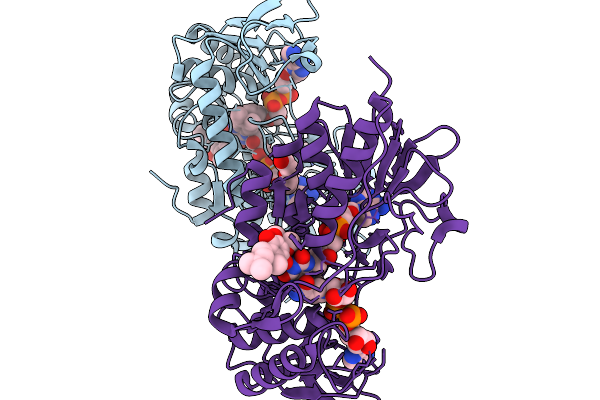 |
Organism: Mammalia
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: FAD, NAD, UQ1, IMD, PEG |
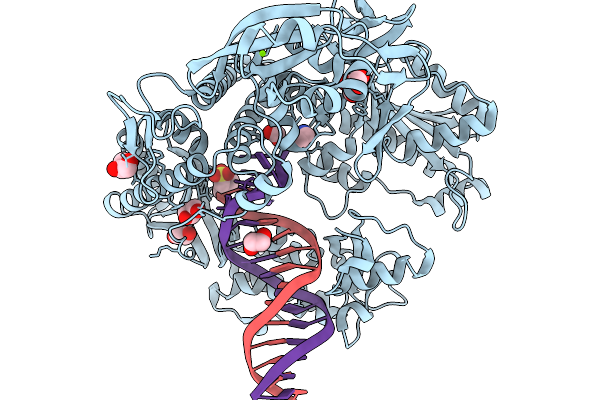 |
Kod-H4 Dna Polymerase Mutant In A Binary Complex With Dna:Dna Containing Two Atna Nucleotides
Organism: Thermococcus kodakarensis kod1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: IMD, GOL, FMT, MES, EDO, MG, MN |
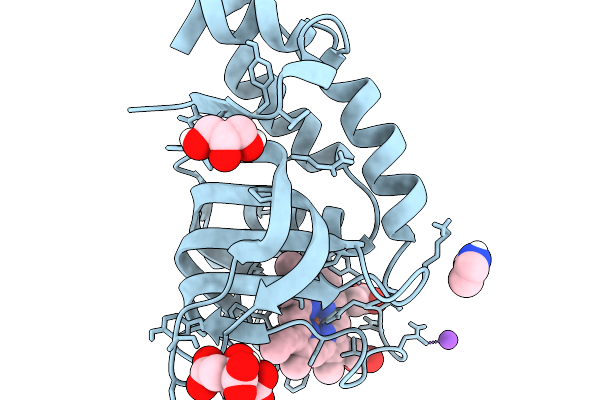 |
Organism: Serratia marcescens
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN Ligands: IMD, CIT, GOL, HEM, NA |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-25 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1C20, IMD |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2026-02-25 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1C2Z, IMD |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: NAG, ZN, A1JPY, MLI, BO3, IMD, EDO, CL |

