Search Count: 540
All
Selected
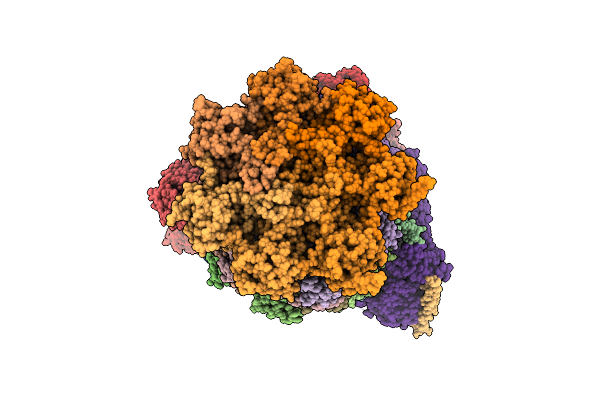 |
Organism: Homo sapiens, Escherichia coli str. k-12 substr. mg1655, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.42 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: ZN, ATP, MG, ADP, LDZ |
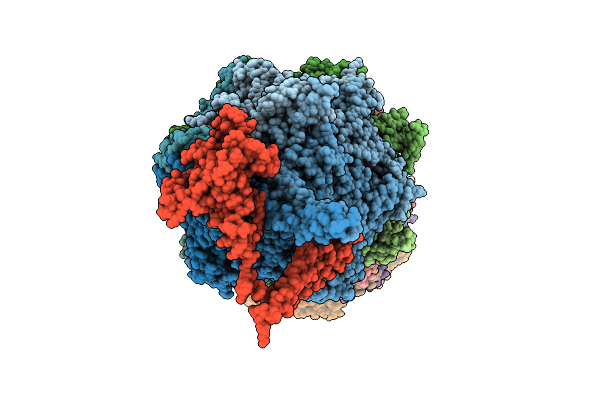 |
Substrate-Free Human 26S Proteasome Purified By Midnolin, 20S Proteasome, Rpts And Rpn11 Part
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: HYDROLASE Ligands: ZN, ATP, MG, ADP, LDZ |
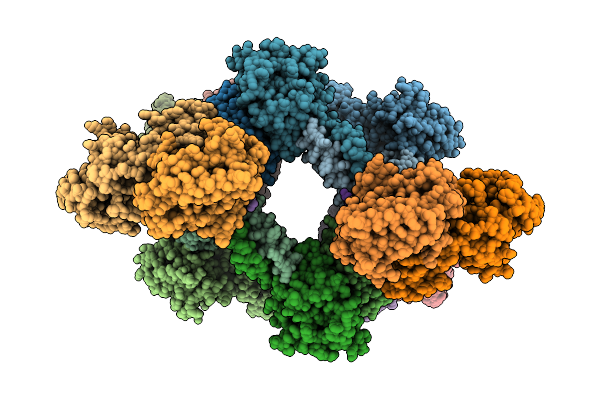 |
Organism: Enterococcus faecalis
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN Ligands: CA |
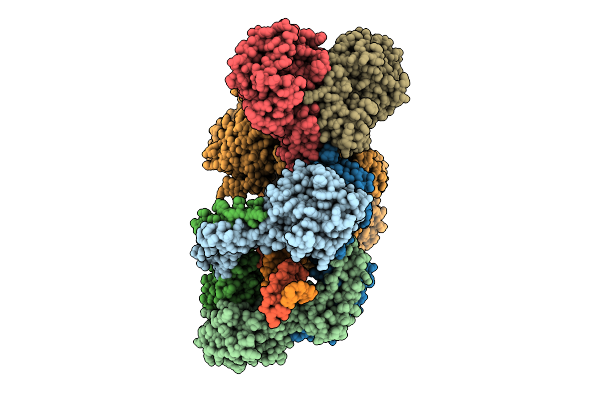 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.76 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA Ligands: CA |
 |
The Crystal Structure Of Paib From Bacillus Stearothermophilus Bound To Hem
Organism: Geobacillus kaustophilus hta426
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2026-04-01 Classification: TRANSCRIPTION Ligands: HEM |
 |
Structure Of Human 26S Proteasome Complexed With Midnolin, 19S Proteasome With Ubl Bound
Organism: Escherichia coli k-12, Homo sapiens, Purpureocillium lilacinum
Method: ELECTRON MICROSCOPY Resolution:3.65 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: ADP, ATP, ZN, MG |
 |
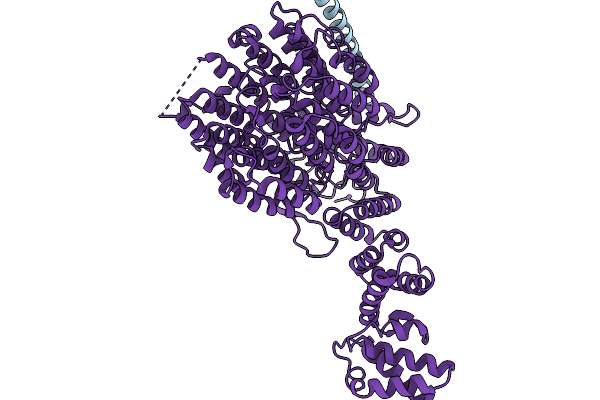
Structure Of Human 26S Proteasome Complexed With Midnolin, 19S Proteasome With Ubl And Catch Domain Resolved
Organism: Escherichia coli k-12, Homo sapiens, Pseudotamlana agarivorans
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: HYDROLASE Ligands: ADP, ATP, ZN, MG |
 |
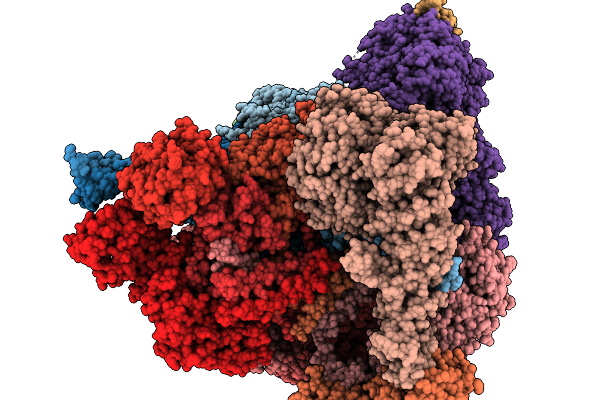
Focused Refinement Of Rpn1 And The C-Terminal Helix Of Midnolin In The Substrate-Engaged Human 26S Proteasome
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: HYDROLASE |
 |
Substrate-Engaged Human 26S Proteasome Bound To Midnolin With Rpt1 At Top Of Spiral Staircase
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: HYDROLASE Ligands: ZN, ATP, MG, ADP, LDZ |
 |
Substrate-Engaged Human 26S Proteasome Bound To Midnolin With Rpt5 At Top Of Spiral Staircase
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: HYDROLASE Ligands: ZN, ATP, MG, ADP, LDZ |
 |
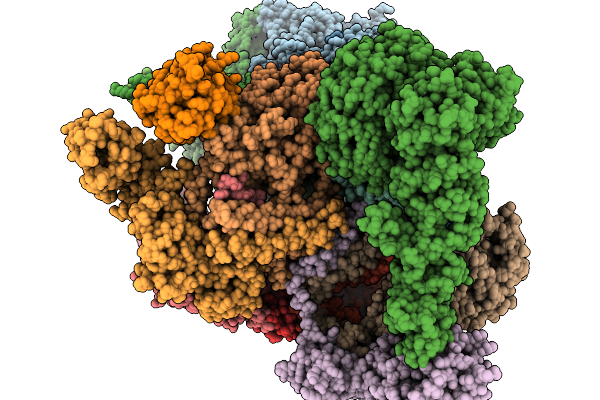
Substrate-Engaged Human 26S Proteasome Bound To Midnolin With Rpt2 At Top Of Spiral Staircase
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: HYDROLASE Ligands: ADP, ATP, MG, LDZ, ZN |
 |
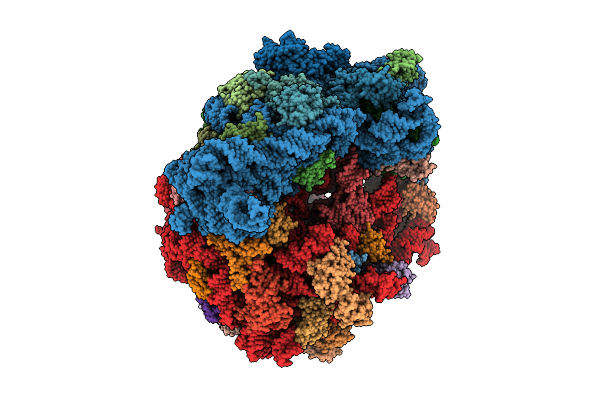
Focused Refinement Of 19S In The Substrate-Engaged Human 26S Proteasome Bound To Midnolin With Rpt6 At Top Of Spiral Staircase
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: HYDROLASE Ligands: ADP, ZN, ATP, MG |
 |
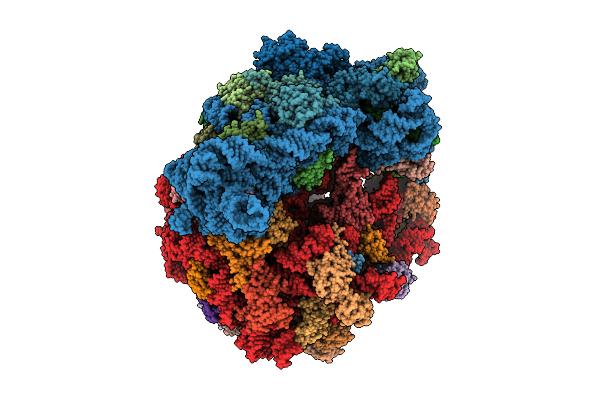
Structure Of Wt E.Coli Ribosome 70S Subunit With Complexed With Mrna, P-Site Fmet-Nh-Trnafmet And A-Site (S)-Betahydroxybock Charged Nh-Trnapyl
Organism: Escherichia coli, Methanomethylophilus alvi
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: RIBOSOME Ligands: ZN, MG, FME, K, PAR, SPD, SPM, A1B71 |
 |
Structure Of Wt E.Coli Ribosome 70S Subunit With Complexed With Mrna, P-Site Fmet-Nh-Trnafmet And A-Site (R) Beta-2-Hydroxy-Boclysine Acid Charged Nh-Trnapyl
Organism: Escherichia coli, Methanomethylophilus alvi
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: RIBOSOME Ligands: PAR, MG, SPD, FME, SPM, K, ZN, A1B70 |
 |
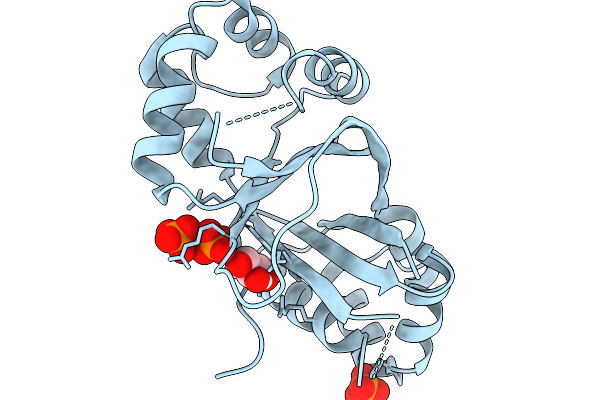
Crystal Structure Of Human Nfix In Complex With Tggca(N3)Tgcca Palindromic Dna
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
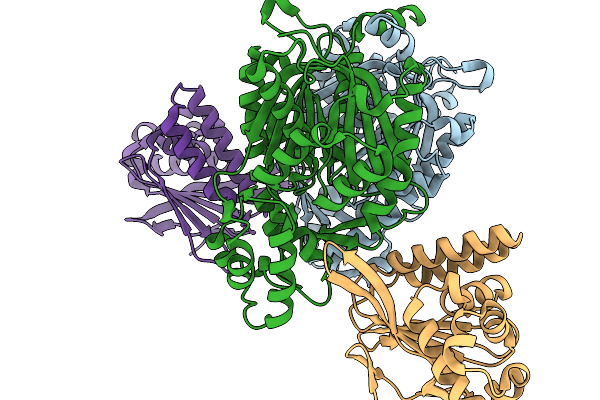
Crystal Structure Of A Polyketide Decarboxylase Abx(+)O From Actinomycetes Sp. Ma7150
Organism: Actinomycetota bacterium
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-03-04 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL, PO4 |
 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-03-04 Classification: TRANSFERASE |
 |
Organism: Psychrobacter lutiphocae dsm 21542, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA Ligands: CA, GOL, PEG |

