Search Count: 77,680
All
Selected
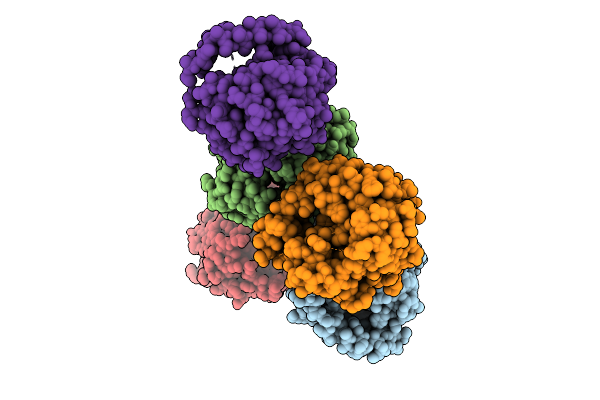 |
Crystal Structure Of Apo Alpha/Beta-Hydrolase Macrolide Esterase Estt From Sphingobacterium Faecium (S126A Mutant)
Organism: Sphingobacterium faecium
Method: X-RAY DIFFRACTION Resolution:3.19 Å Release Date: 2026-04-08 Classification: HYDROLASE |
 |
Crystal Structure Of Apo Alpha/Beta-Hydrolase Macrolide Esterase Estt From Sphingobacterium Thalpophilum (S102A Mutant)
Organism: Sphingobacterium thalpophilum
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2026-04-08 Classification: HYDROLASE |
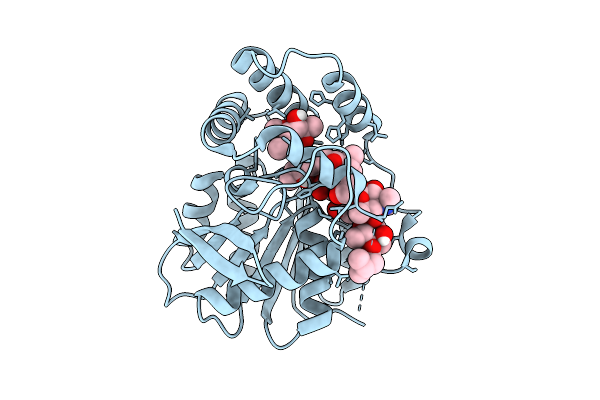 |
Crystal Structure Of Alpha/Beta-Hydrolase Macrolide Esterase Estt From Bacillus Cereus (S102A Mutant) In Complex With Linearized Tylvalosin
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: A1DAR |
 |
Crystal Structure Of Alpha/Beta-Hydrolase Macrolide Esterase Estx From Escherichia Coli (S102A Mutant) In Complex With Linearized Tylosin
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: A1E2O |
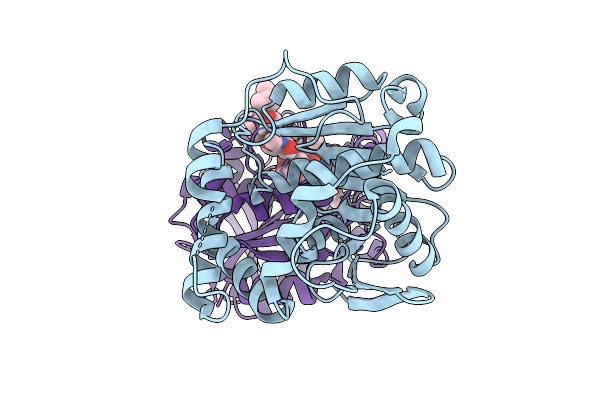 |
Crystal Structure Of Alpha/Beta-Hydrolase Macrolide Esterase Estx From Escherichia Coli (S102A Mutant) In Complex With Linearized Tylvalosin
Organism: Escherichia coli #1/h766
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: A1DAR |
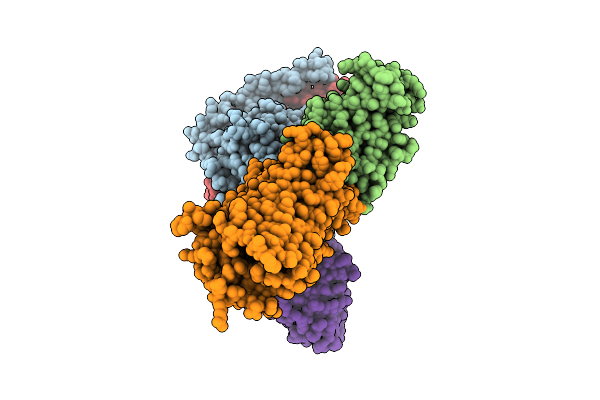 |
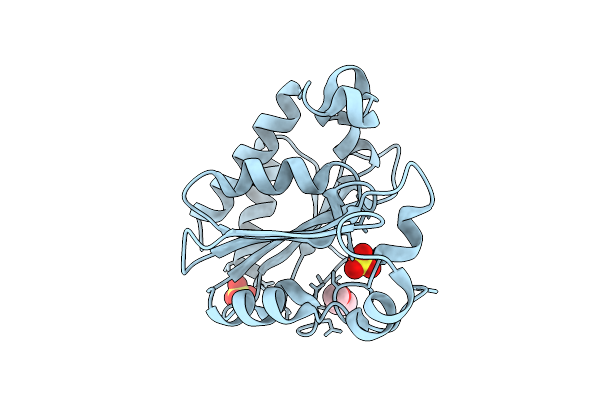
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: A1E2G |
 |
Crystal Structure Of Poly-Gamma-Glutamate Hydrolase From Bacillus Phage Pm1
Organism: Bacillus phage pm1
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: PEG, SO4 |
 |
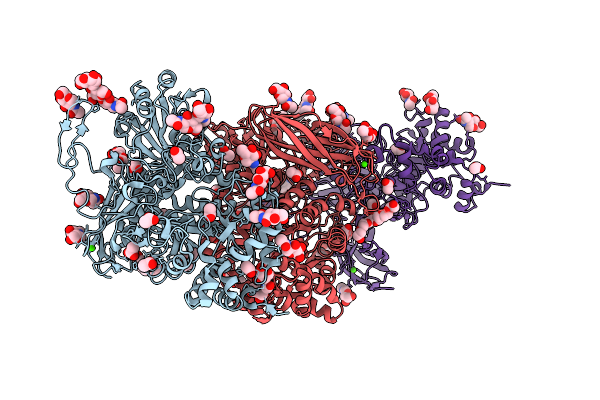
Crystal Structure Of Poly-Gamma-Glutamate Hydrolase From Bacillus Phage Pm1 In Complex With A Zinc Ion
Organism: Bacillus phage pm1
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: PEG, ZN |
 |
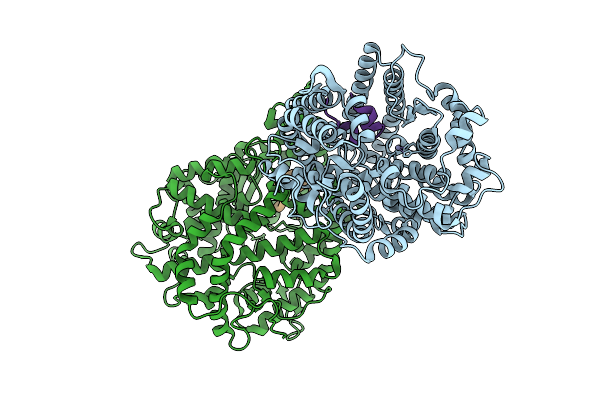
Crystal Structure Of Isoprimeverose-Producing Enzyme From Phaeoacremonium Minimum
Organism: Rattus rattus, Phaeoacremonium minimum
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAG, ACT, PG4, CA, PGE, P6G |
 |
Organism: Homo sapiens, Synthetic plasmid
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A5Z, CL |
 |
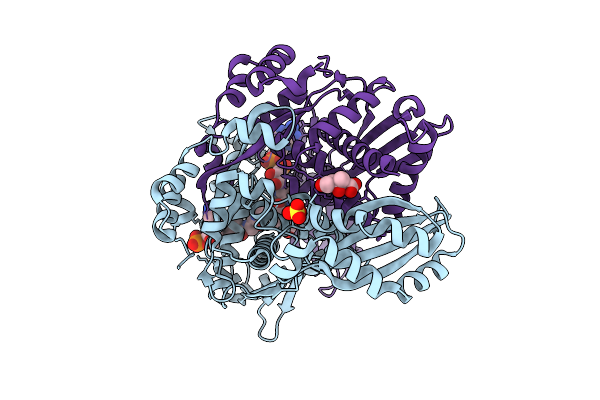
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A54, CL |
 |
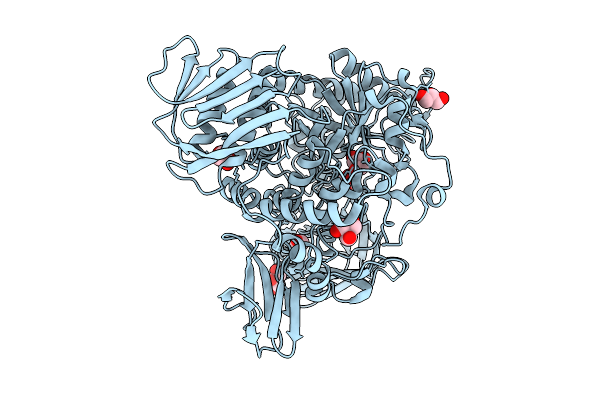
Structure Of Pmhmgr Bound To Mevalonate, Coa And Nad 2 Minutes 20 Seconds After Reaction Initiation At Ph 9
Organism: Pseudomonas sp. 'mevalonii'
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE/SUBSTRATE/INTERMEDIATE Ligands: COA, SO4, A1AFY, ZKE, NAI, MEV |
 |
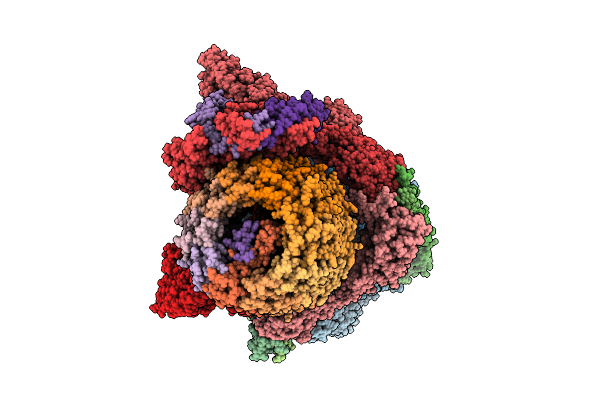
Sdacs: A Synergistic Strategy Using Both Crystallographic And Alphafold Structures To Improve Thermostability And Activity Of N-Acetylhexosaminidase
Organism: Cellvibrio mixtus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: TRS, GOL |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: HYDROLASE Ligands: ADP |
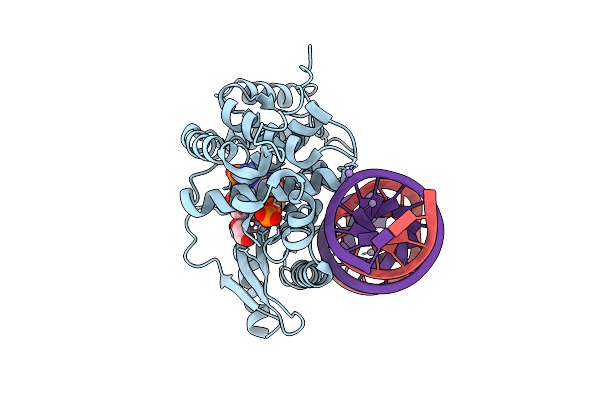 |
Crystal Structure Of The Pre-Reactive State Of Porcine Oas1 In Complex With Dsrna, Two Apcpp Substrate Analogs, Three Catalytic Mn2+ Ions.
Organism: Sus scrofa, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE/RNA Ligands: APC, MN, EDO |
 |
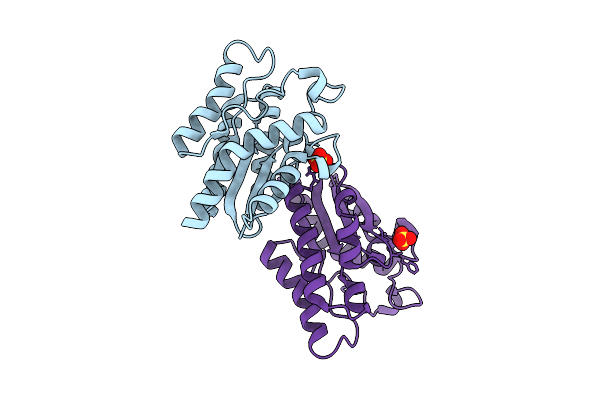
Organism: Rattus norvegicus, Sus scrofa
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-04-08 Classification: CELL CYCLE Ligands: GTP, GDP, A1B97, SO4 |
 |
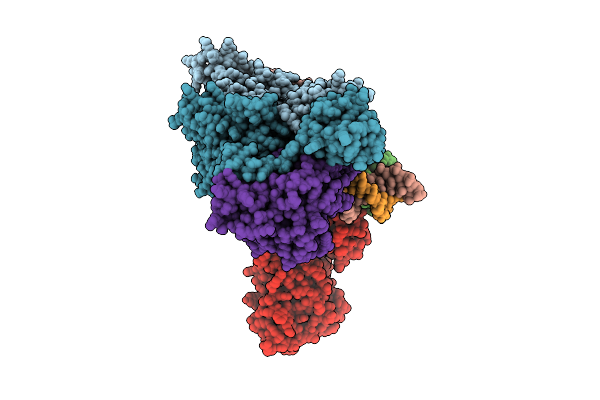
Organism: Chloracidobacterium thermophilum
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: SO4 |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, MG, ATP |
 |
Crystal Structure Of The S-Adenosyl-L-Homocysteine Hydrolase From P. Aeruginosa, Q65N Mutant Probed With Rubidium To Confirm Disruption Of A Potassium Binding Site.
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAD, GOL, PO4 |

