Search Count: 9,160
All
Selected
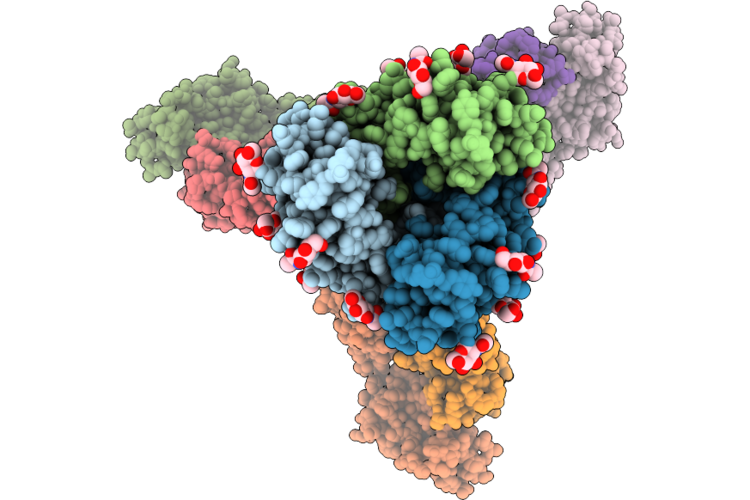 |
Cryo-Em Structure Of Sars-Cov-2 Wide-Type S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab (Focused Refinement Of The S2-76E1)
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
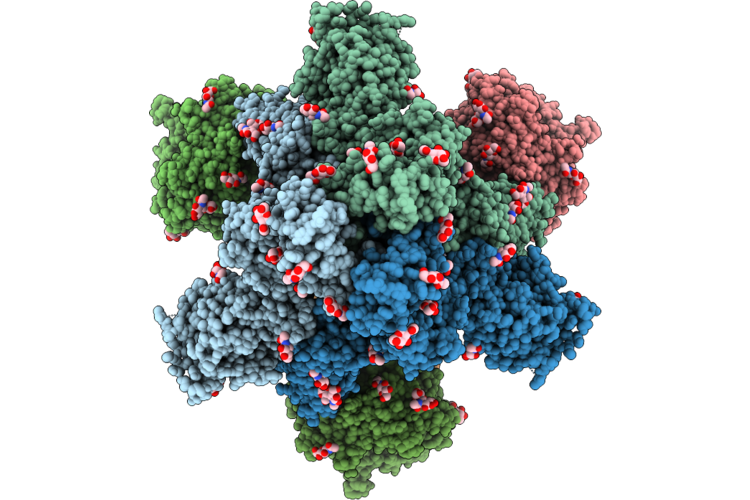 |
Cryo-Em Structure Of Sars-Cov-2 Wide-Type S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
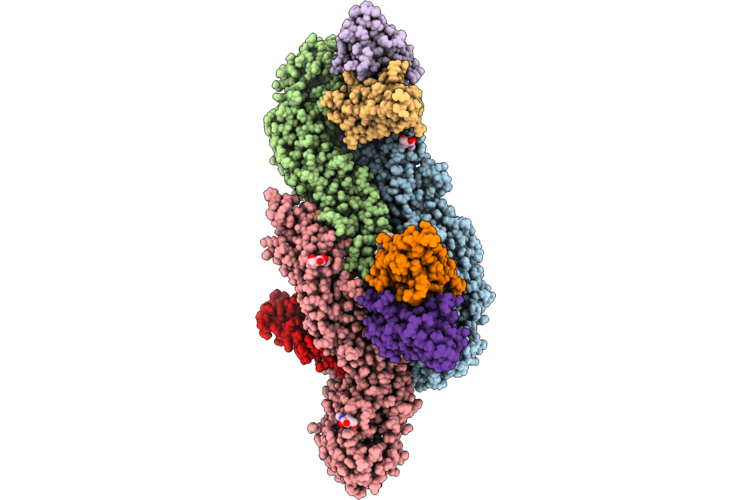 |
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRUS Ligands: NAG |
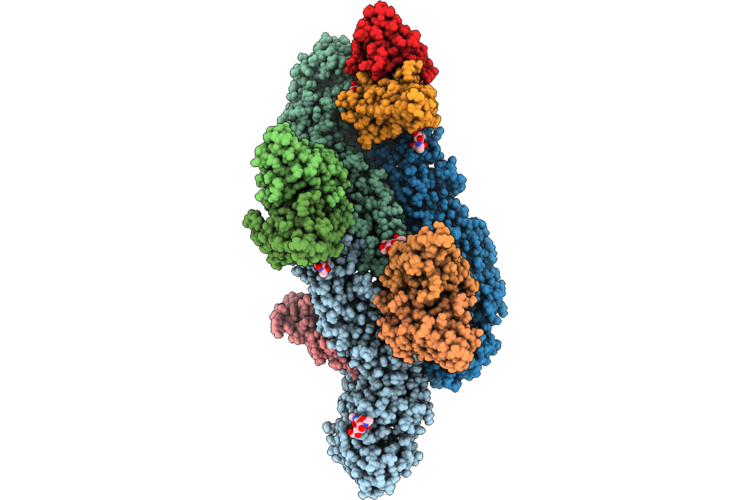 |
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRUS |
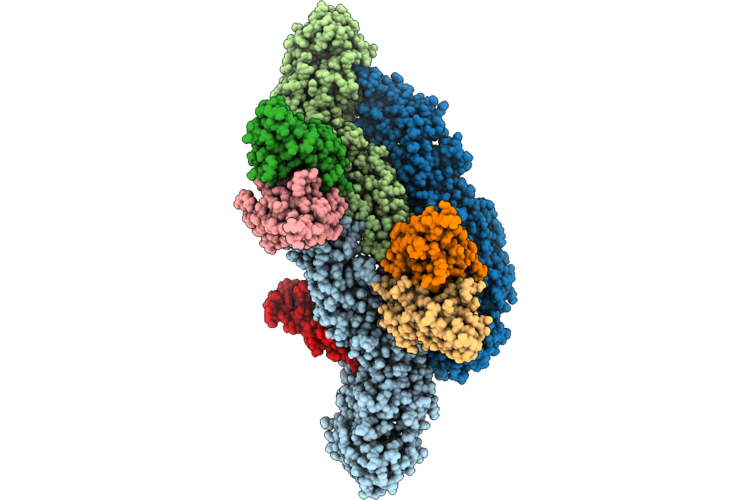 |
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRUS |
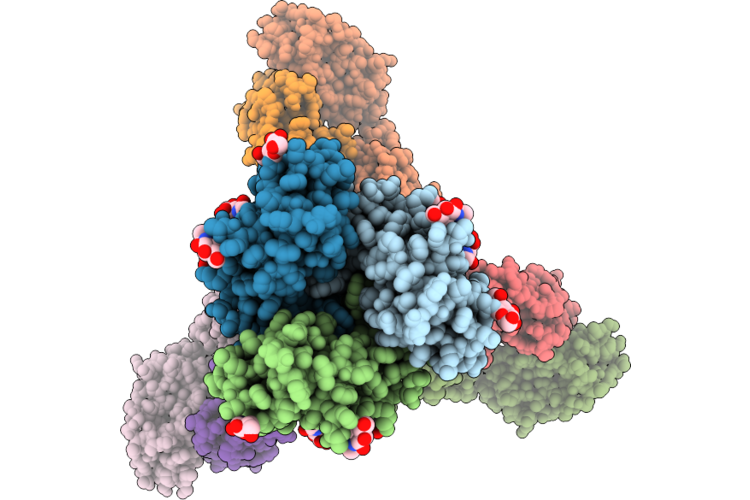 |
Cryo-Em Structure Of Sars-Cov-2 Xbb.1.5 S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab (Focused Refinement Of The S2-76E1)
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
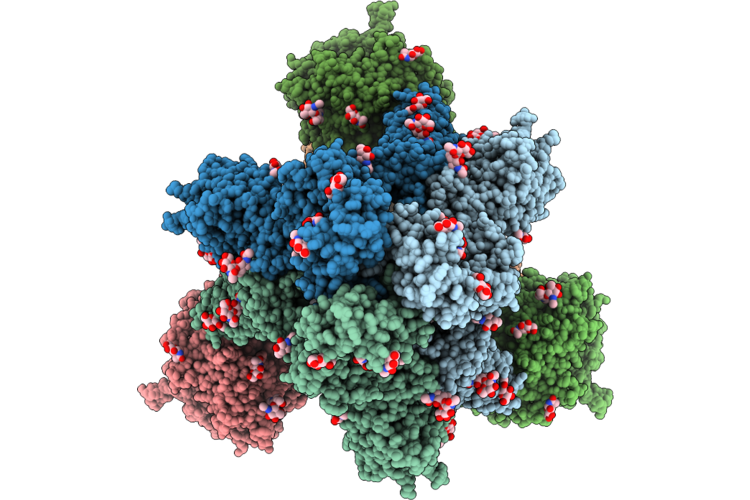 |
Cryo-Em Structure Of Sars-Cov-2 Xbb.1.5 S Trimer In The Early Fusion Intermediate Conformation (E-Fic) Complexed With Ace2 And 76E1-Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
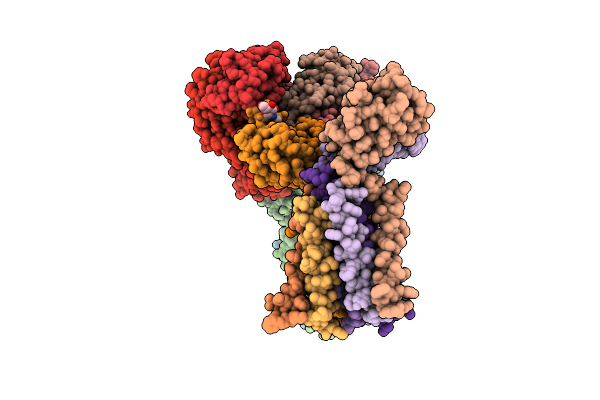 |
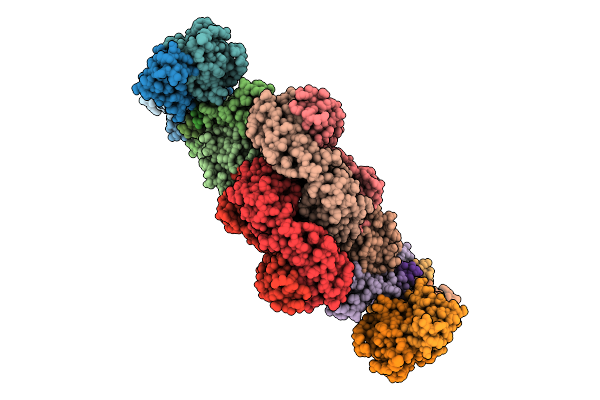
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.35 Å Release Date: 2026-04-15 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
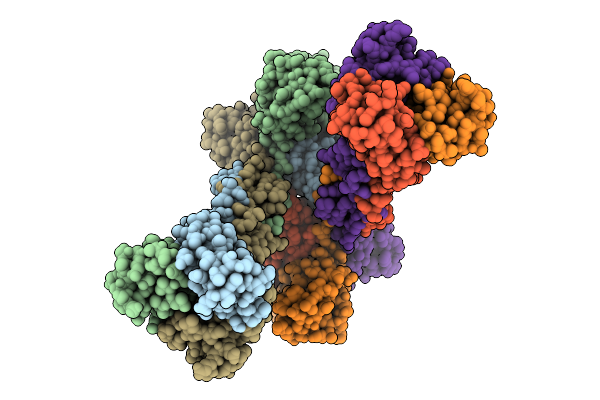
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: IMMUNE SYSTEM |
 |
Crystal Structure Of Gp232 Tail Fiber Recognition Domain From Mycobacteriophage Bxz-1
Organism: Bixzunavirus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN |
 |
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Complex With Neutralizing Antibody Y10F
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
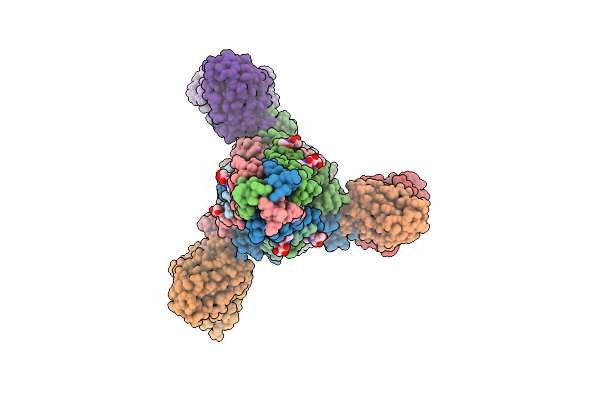
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Complex With Two Neutralizing Antibodies Mab77 And Y10F
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
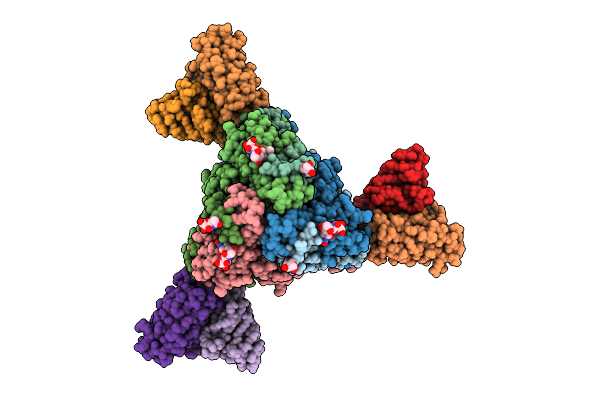
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Postfusion Conformation Bound To Neutralizing Antibody H8
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Prefusion Conformation Bound To Neutralizing Antibody H8
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Crystal Structure Of A Zikv E Glycoprotein Di-Diii Vaccine Candidate In Complex With Human Neutralizing Antibody Mz4
Organism: Homo sapiens, Zika virus zikv/h. sapiens/frenchpolynesia/10087pf/2013
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: NAG |
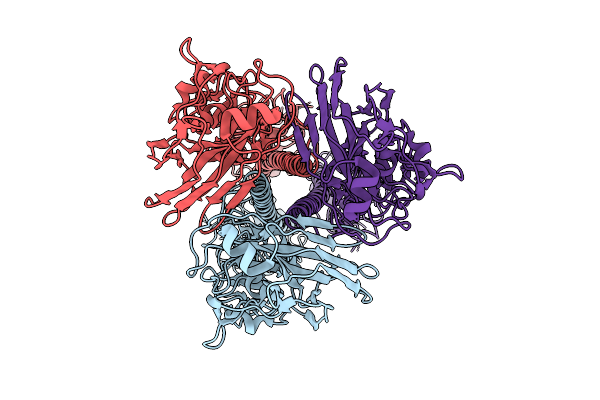 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.39 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
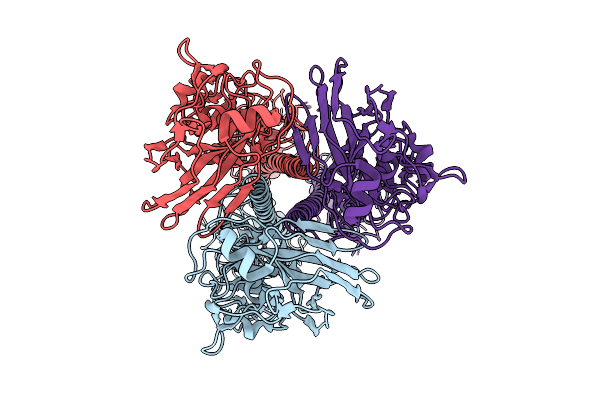 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.16 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
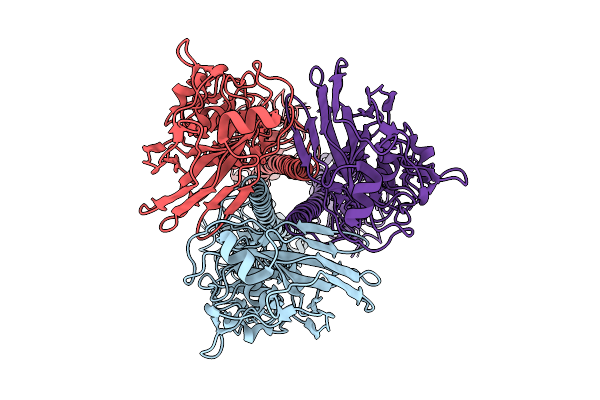
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.67 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
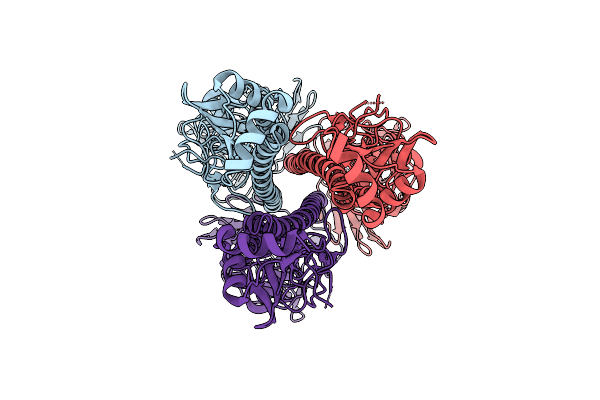
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.47 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
Influenza A Virus Hemagglutinin (A/California/04/2009 H1N1), E47K Ha2 Stabilizing Mutation (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |

