Search Count: 35
All
Selected
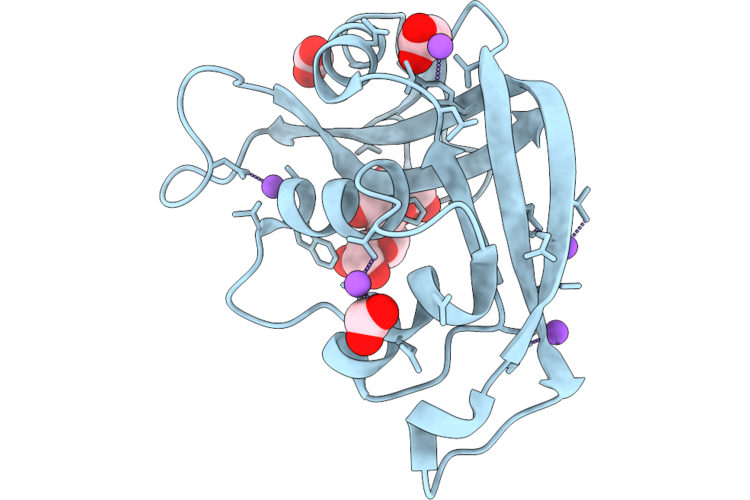 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:0.94 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, FMT, NA |
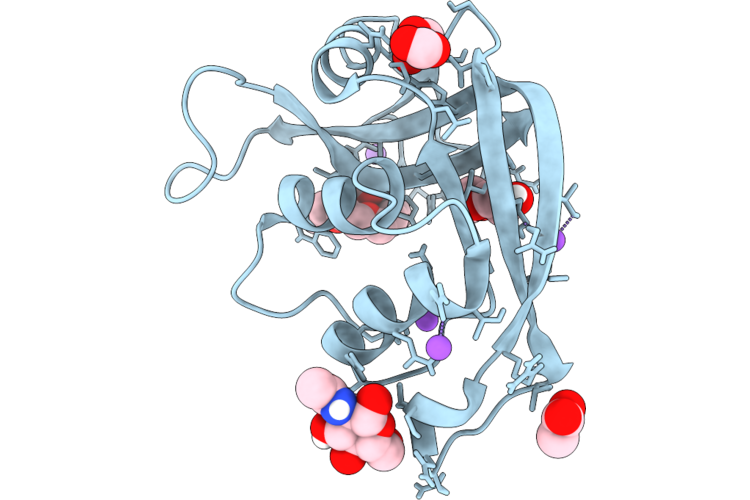 |
Structural Characterization Of A N-Acetyl-D-Glucosamine-Binding Site In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, NAG, PG4, FMT, PEG, NA |
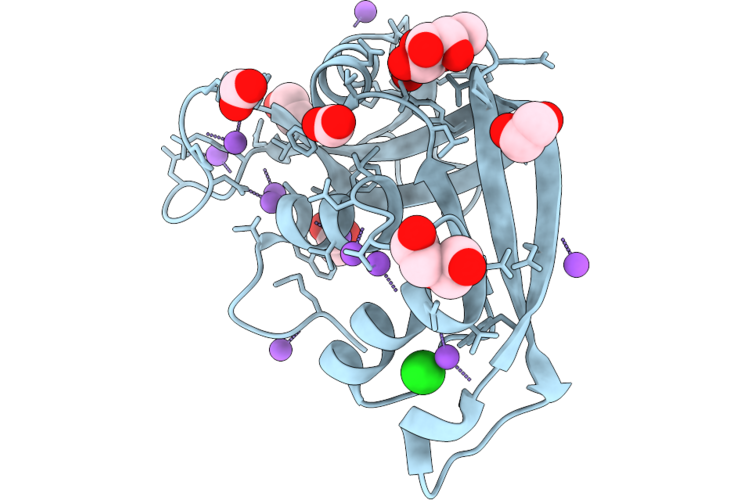 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: GLA, ACT, FMT, PEG, EDO, NA, CL |
 |
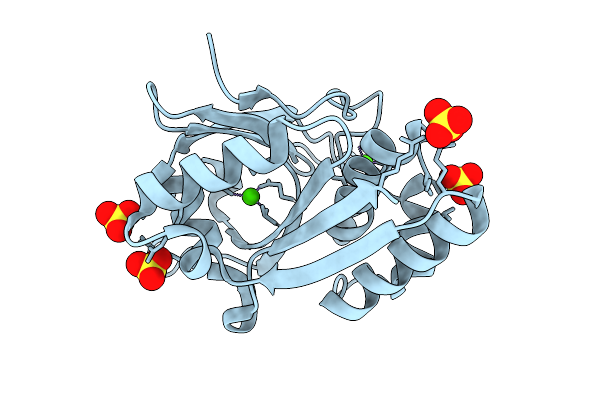
Structural Characterization Of A N-Acetylmuramic Acid-Binding Site In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.01 Å Release Date: 2026-04-15 Classification: PROTEIN BINDING Ligands: MUB, AMU, ACT, FMT, NA, CL |
 |
Organism: Metagenomes
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: LYASE Ligands: CA, SO4 |
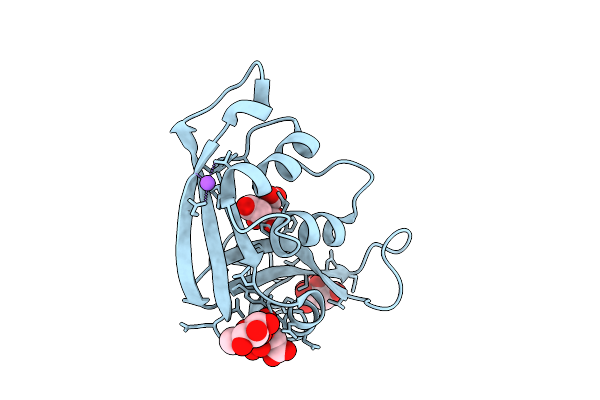 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: CIT, GLC, NA, CL |
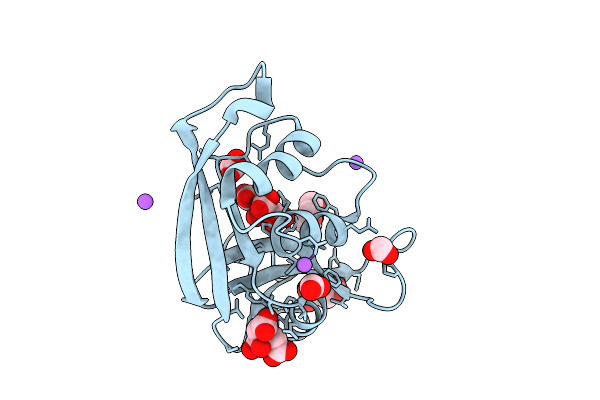 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.17 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: CIT, FMT, PEG, ACT, NA |
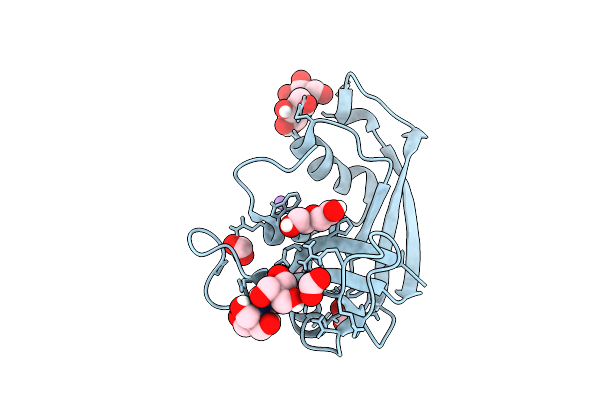 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: B3P, FMT, PEG, NA |
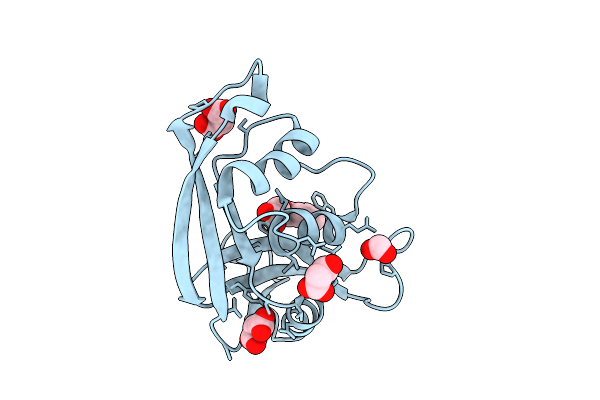 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: MLI, PEG, ACT, FMT |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: PEG, ACT, FMT, FUL, FUC |
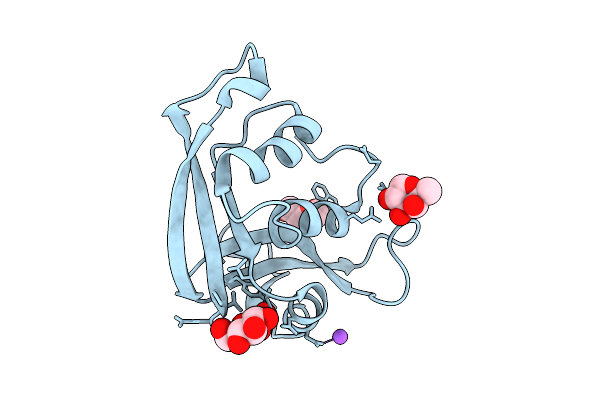 |
Structural Characterization Of Glucose- And Fucose-Binding Sites In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.01 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: PEG, FUC, ACT, GLC, NA |
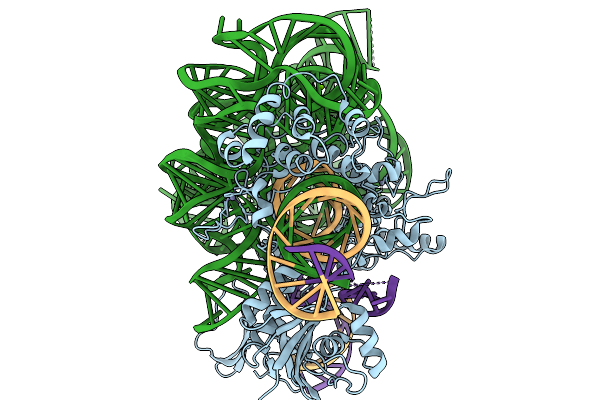 |
Organism: Metagenome, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN Ligands: MG, ZN |
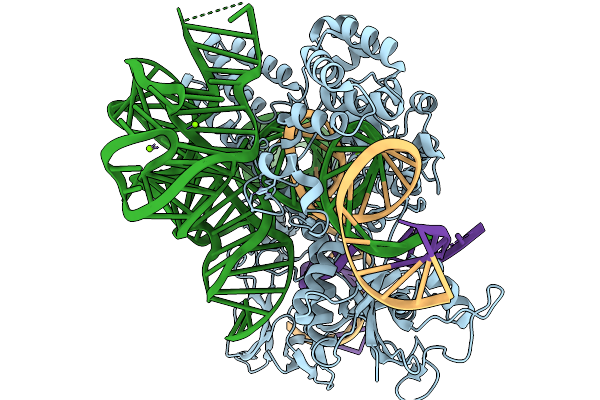 |
Organism: Nitrospirota bacterium, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN Ligands: MG, ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2025-06-11 Classification: TRANSFERASE Ligands: AMP, PO4, MG |
 |
Crystal Structure Of The Light-Driven Proton Pump Heimdallarchaeial Rhodopsin Heimdallr1
Organism: Candidatus heimdallarchaeota archaeon
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-04-02 Classification: PROTON TRANSPORT Ligands: RET, OLC, NO3 |
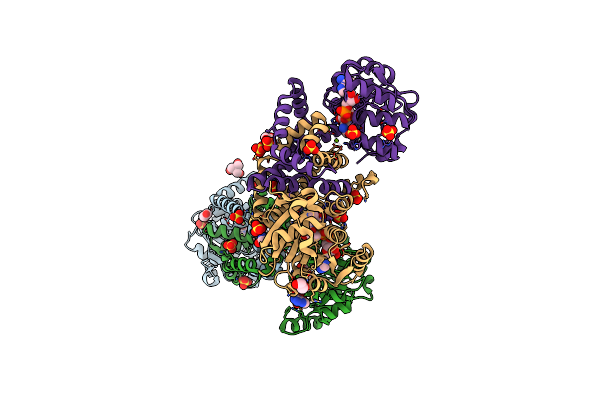 |
Organism: Syntrophothermus lipocalidus (strain dsm 12680 / tgb-c1)
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2024-07-17 Classification: OXIDOREDUCTASE Ligands: NAI, SO4, MG, EDO, TRS |
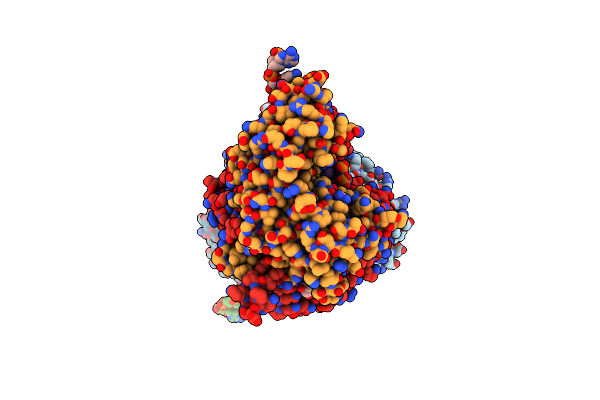 |
Organism: Geobacter sulfurreducens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-07-03 Classification: DNA/RNA/CELL INVASION Ligands: MG |
 |
Organism: Geobacter sulfurreducens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-07-03 Classification: DNA/RNA/CELL INVASION Ligands: MG |
 |
Organism: Thermoflavifilum thermophilum, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-11-08 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Thermoflavifilum thermophilum
Method: ELECTRON MICROSCOPY Release Date: 2023-10-18 Classification: DNA BINDING PROTEIN |

