Search Count: 3,892
All
Selected
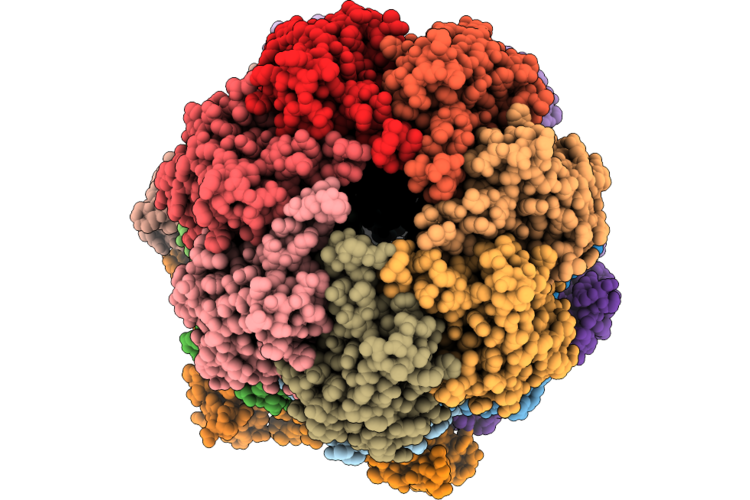 |
Organism: Enterobacter
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-22 Classification: CHAPERONE Ligands: ATP |
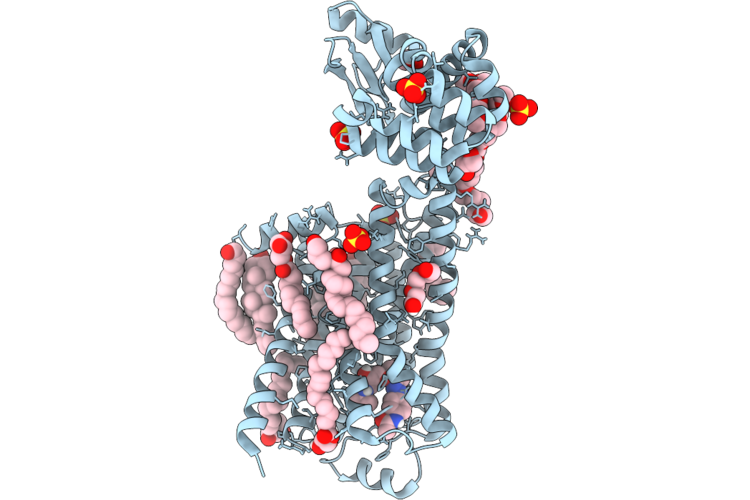 |
Dark Structure Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State Recorded At Swissfel
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, CLR, 12P, OLC, 1PE, EDO, GOL, A1JHU, PLM |
 |
Dark Structure Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State Recorded At Lcls
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, CLR, PLM, 12P, A1JHU, OLC, EDO, GOL |
 |
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-03-18 Classification: METAL BINDING PROTEIN Ligands: MTE, B3P, MOO, EFK, CL |
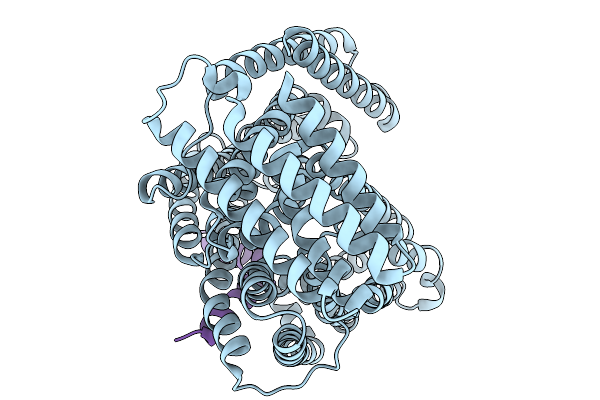 |
Organism: Escherichia coli k-12, Enterobacteria phage m
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN |
 |
Cryo-Em Structure Of The Escherichia Phage Hk446 Rip1 In Complex With The Enterobacteria Phage T6 Small Terminase
Organism: Enterobacteria phage t6, Escherichia phage hk446
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: VIRAL PROTEIN |
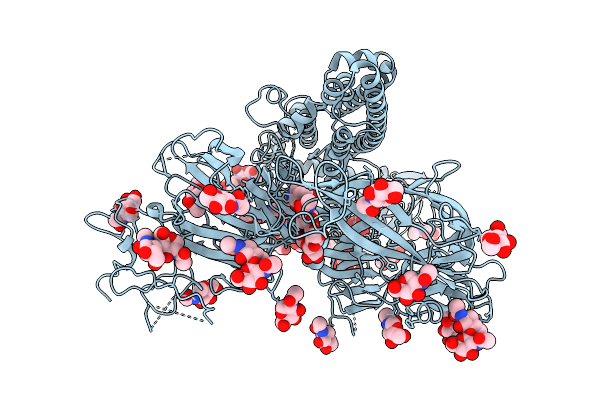 |
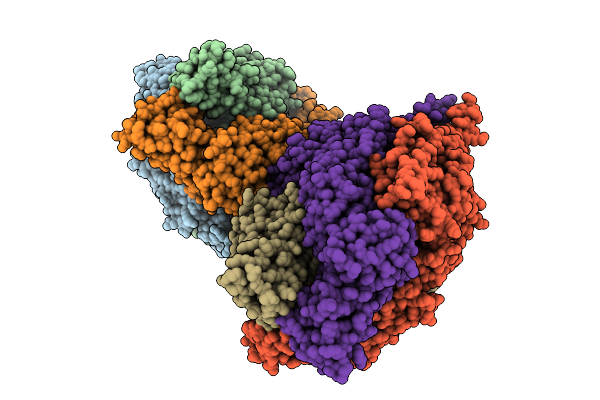
Organism: Feline infectious peritonitis virus (strain 79-1146), Enterobacteria phage t6
Method: ELECTRON MICROSCOPY Resolution:2.78 Å Release Date: 2026-01-28 Classification: PROTEIN BINDING Ligands: NAG |
 |
Organism: Human respiratory syncytial virus, Enterobacteria phage t6
Method: X-RAY DIFFRACTION Resolution:2.92 Å Release Date: 2025-12-17 Classification: VIRAL PROTEIN |
 |
Organism: Respiratory syncytial virus, Enterobacteria phage t6
Method: X-RAY DIFFRACTION Resolution:2.77 Å Release Date: 2025-12-17 Classification: VIRAL PROTEIN |
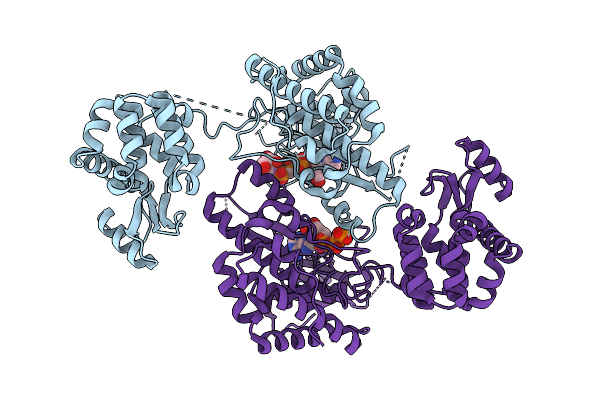 |
Crystal Structure Of Phosphatidyl Inositol 4-Kinase Ii Beta In Complex With Hh5129
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-12-03 Classification: TRANSFERASE Ligands: A1IVA |
 |
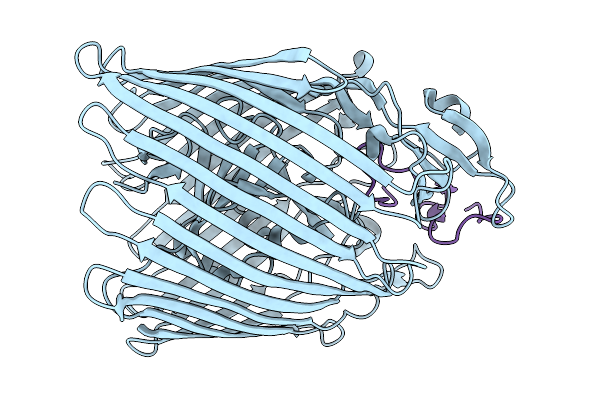
Organism: Enterobacteria phage t4
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: ISOMERASE |
 |
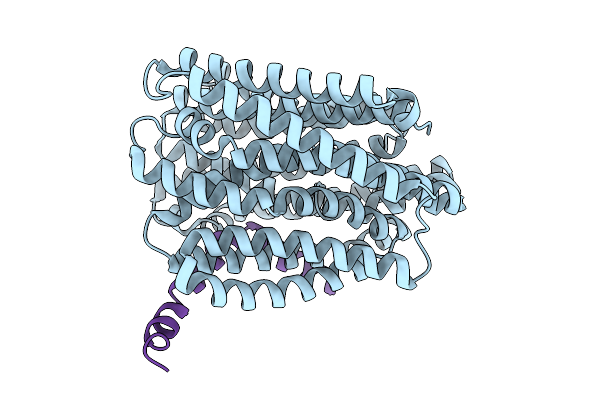
Organism: Escherichia coli, Enterobacteriaceae
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN |
 |
Organism: Escherichia coli, Enterobacteria phage m
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN |
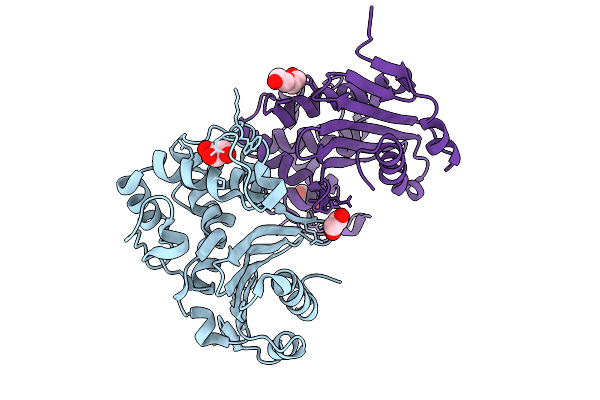 |
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2025-09-03 Classification: ANTIMICROBIAL PROTEIN Ligands: CL, GOL, EDO, PEG |
 |
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2025-09-03 Classification: ANTIMICROBIAL PROTEIN Ligands: NXL, A1IY4, CL, EDO, 2PE |
 |
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2025-09-03 Classification: ANTIMICROBIAL PROTEIN Ligands: OP0, EDO, GOL, PEG, CL, DMS |
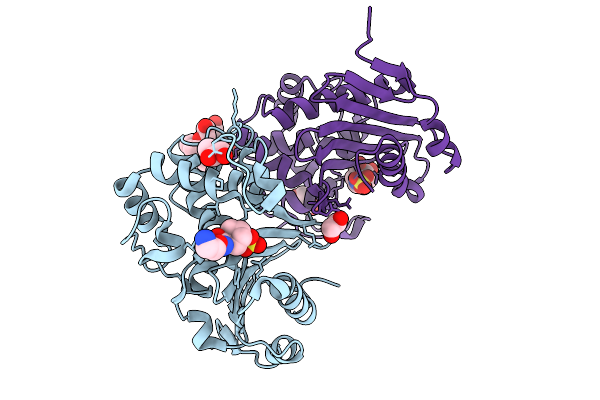 |
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2025-09-03 Classification: ANTIMICROBIAL PROTEIN Ligands: OP0, EDO, GOL, A1IYS, CL, 2PE |
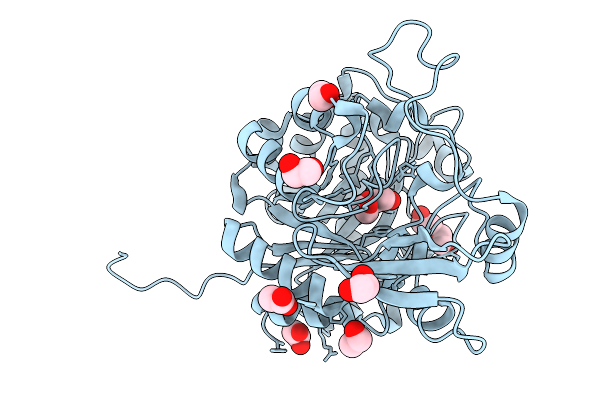 |
Organism: Enterobacter cloacae complex sp. ecnih14
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: EDO, ACT, PEG |
 |
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2025-07-30 Classification: TRANSFERASE |
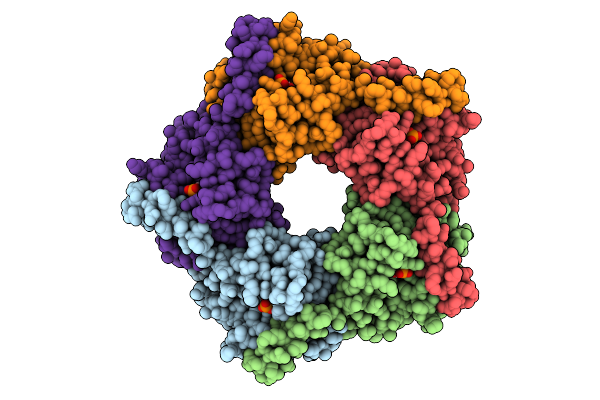 |
X-Ray Structure Of Enterobacter Cloaca Transaldolase In Complex With D-Fructose-6-Phosphate.
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2025-07-30 Classification: TRANSFERASE Ligands: F6R |

