Search Count: 20,863
All
Selected
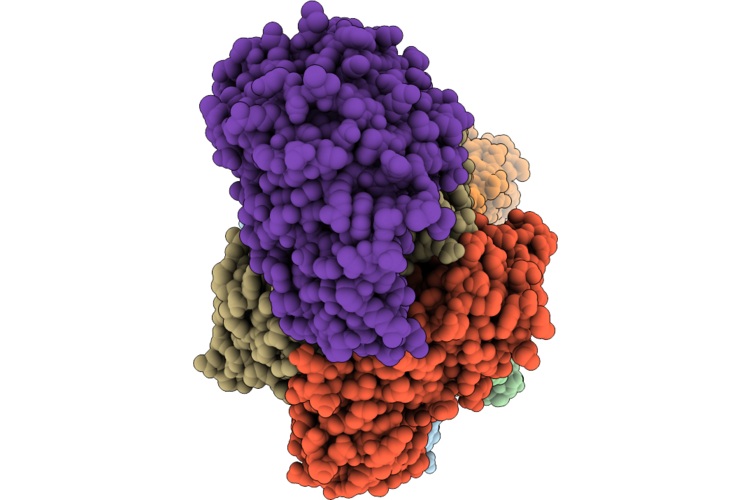 |
Cryoem Structure Of Human Mata2 In Complex With Mat2B Isoform V1 At 2.6 A Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-13 Classification: TRANSFERASE |
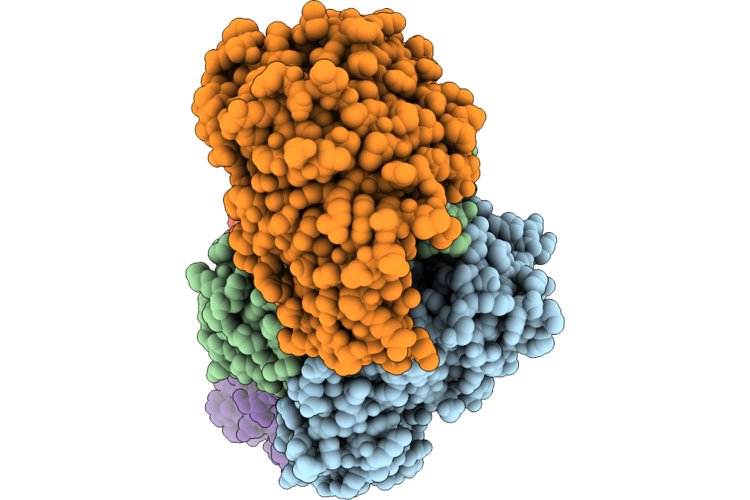 |
Cryoem Structure Of Human Mata2 In Complex With Mat2B Isoform V1 At 2.6 A Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-05-13 Classification: TRANSFERASE |
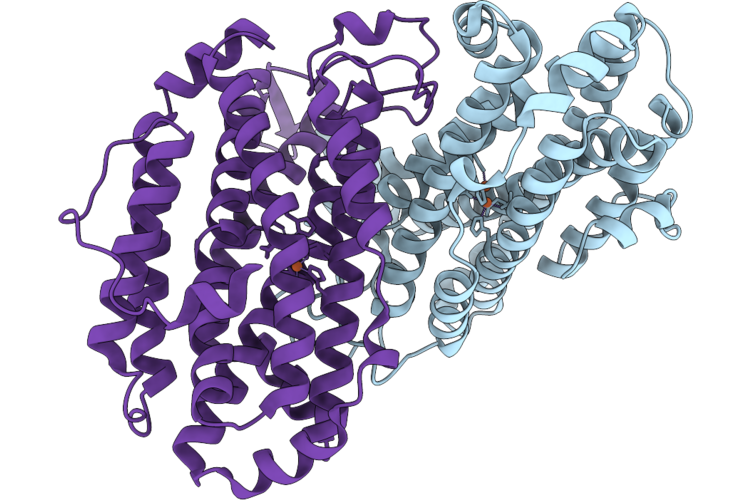 |
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
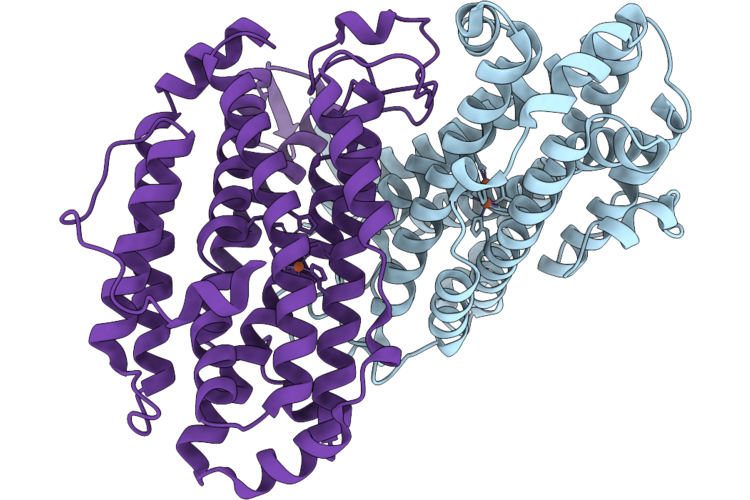 |
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |
 |
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli, Hexagonal P6122 Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
 |
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form, Hexagonal P6122
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |
 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 8.0 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Arg720Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
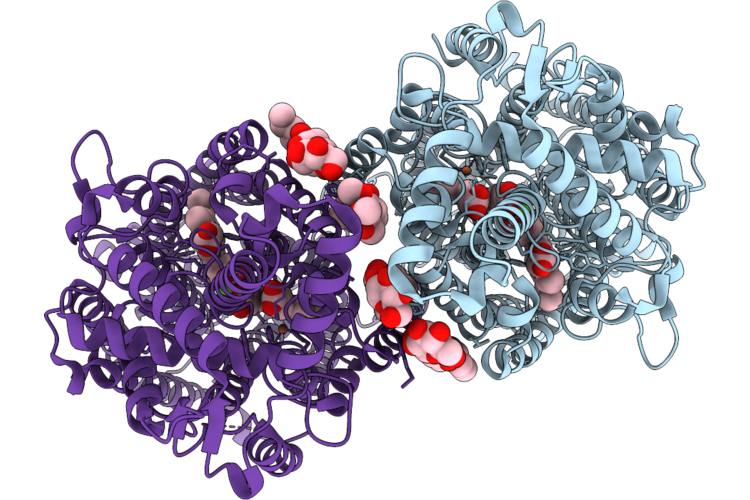 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, UQ5 |
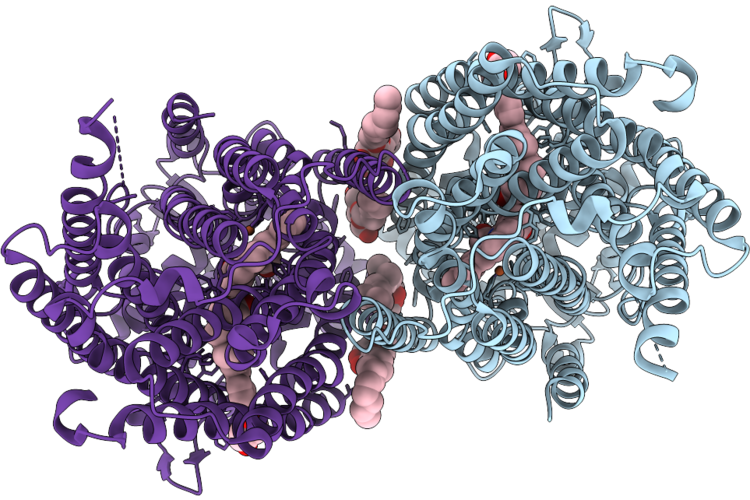 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant With Quino At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, UQ5, HQE, LMT |
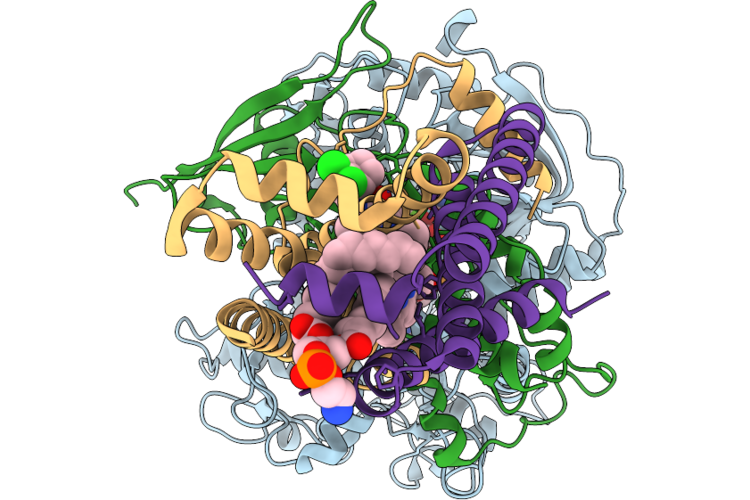 |
The Cryo-Em Structure Of Human Succinate Dehydrogenase In Complex With Benzovindiflupyr
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FAD, FES, SF4, F3S, A1EGM, HEM, PEV |
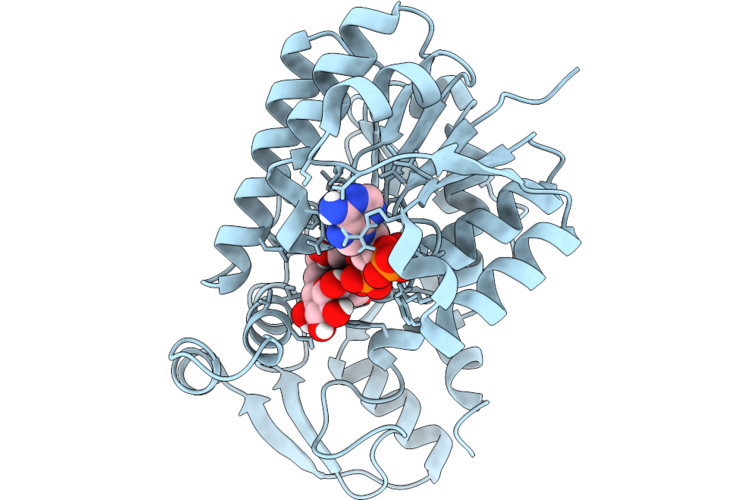 |
Crystal Structure Of An Nta622L Variant In Complex With Nadph And Nicotinic Acid N-Glucoside
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-06 Classification: PLANT PROTEIN Ligands: A1JBN, NDP |
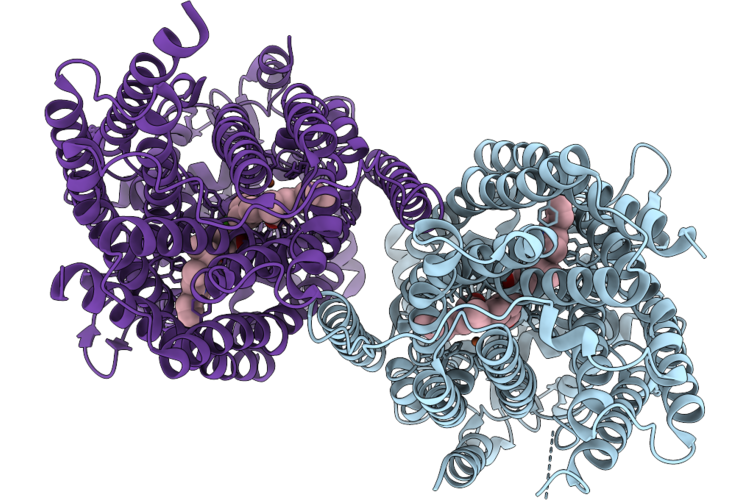 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 6.5
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
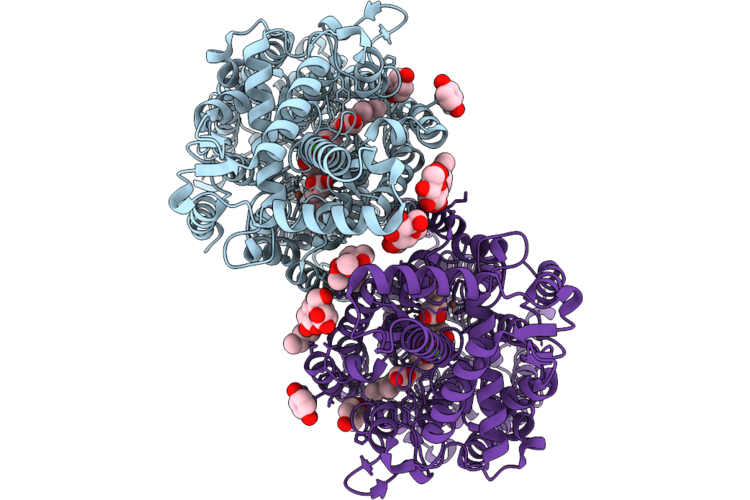 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase With Hqe At Ph 6.5
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, HQE, UQ5 |
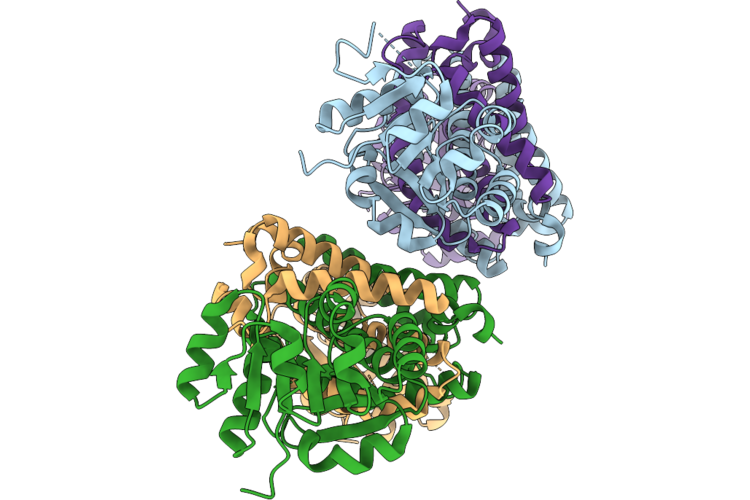 |
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE |
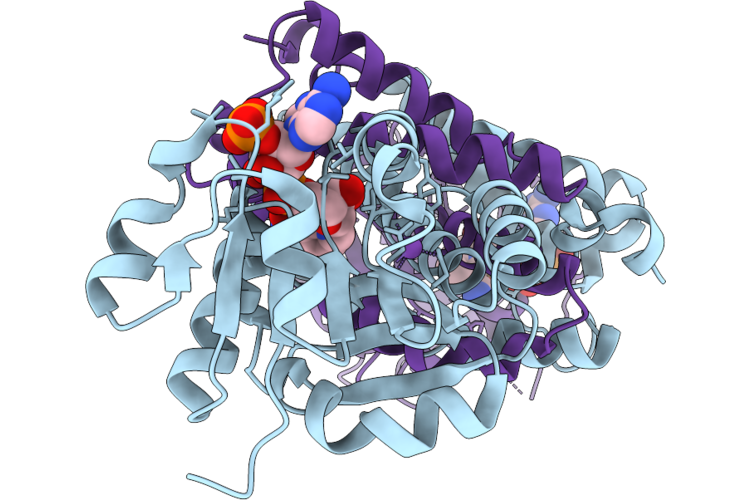 |
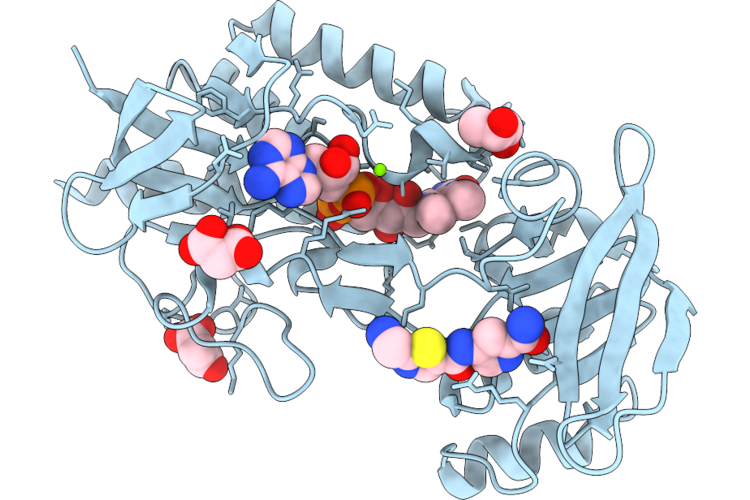
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: NAP, NA |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z4666192264
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1JAA |
 |
Crystal Structure Of An Nta622L Variant In Complex With Nadph And Trigonelline
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-29 Classification: PLANT PROTEIN Ligands: NDP, A1JBP |
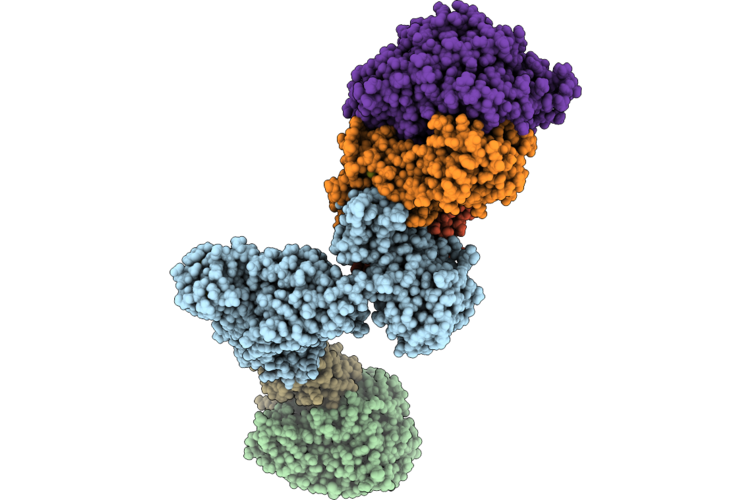 |
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 2 Substate A (Composite Structure)
Organism: Methanothermobacter marburgensis str. marburg
Method: ELECTRON MICROSCOPY Resolution:2.19 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, NFU |
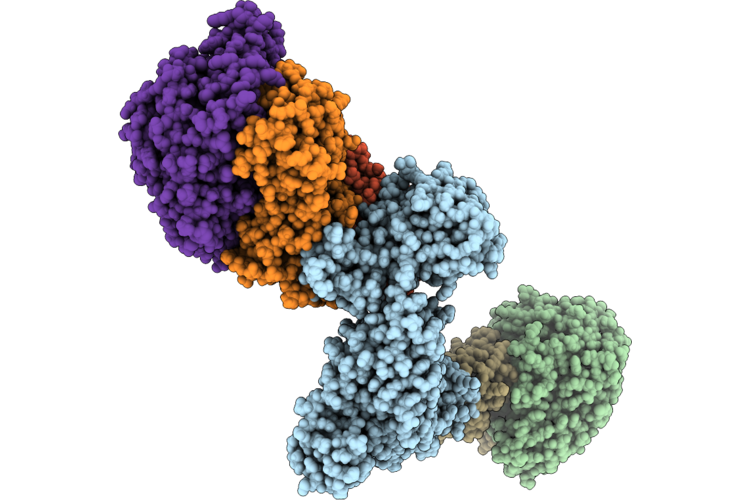 |
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 2 Substate B (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:3.25 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, NFU |

