Search Count: 1,293
All
Selected
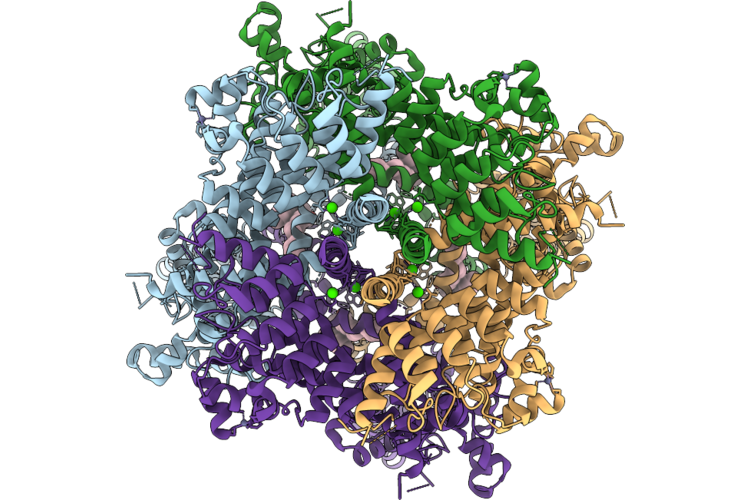 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: METAL TRANSPORT Ligands: CA, ZN, A1L5I |
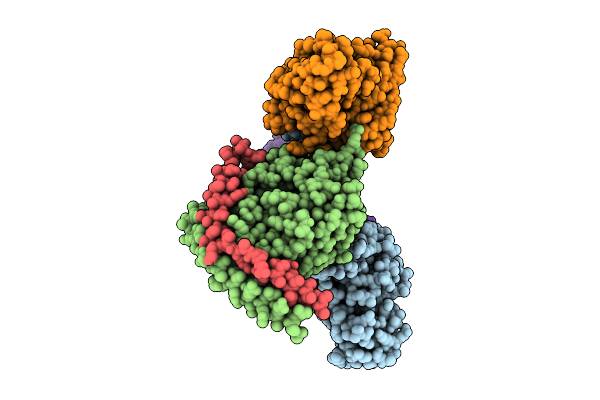 |
Cryoem Structure Of Mu-Opioid Receptor - Gi Protein Complex Bound To Fluornitrazene (Fnz)
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: A1B79 |
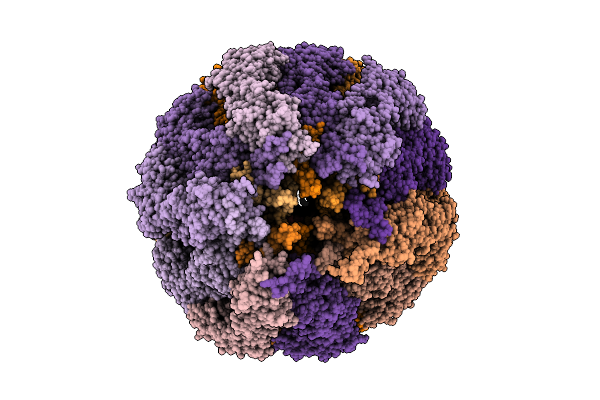 |
Structure Of The Tip Region Of The Intial Complex In Bacterial Flagellar Filament Assembly At 3.68 Angstroms Resolution, Conformation 3.
Organism: Salmonella enterica subsp. enterica serovar typhimurium str. lt2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PROTEIN FIBRIL |
 |
Structure Of The Intial Complex In Filament Assembly At 3.23 Angstroms Resolution, Conformation 2.
Organism: Salmonella enterica subsp. enterica serovar typhimurium str. lt2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: PROTEIN FIBRIL |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Organism: Bos taurus, Mus musculus, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN |
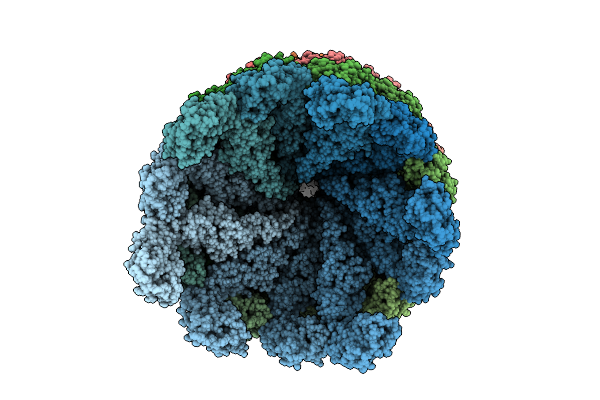 |
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PROTEIN FIBRIL |
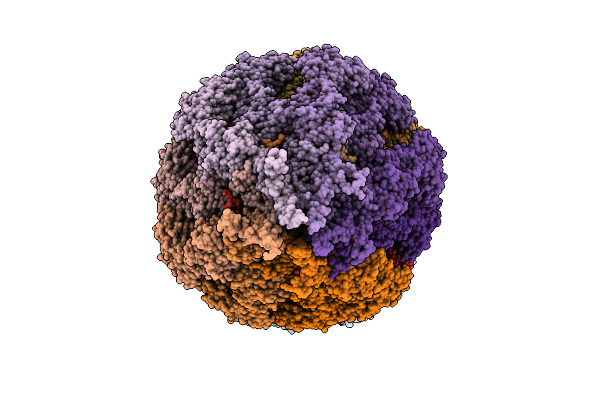 |
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PROTEIN FIBRIL |
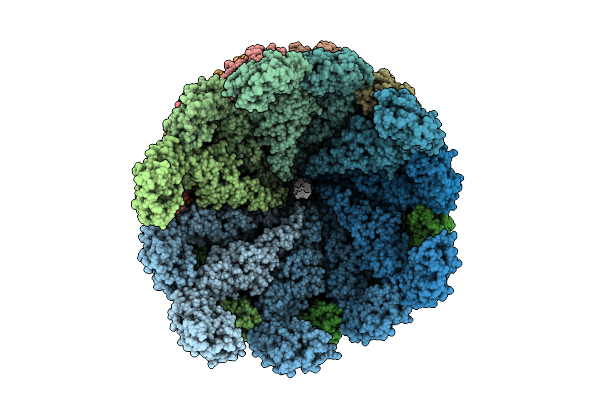 |
Organism: Salmonella enterica subsp. enterica serovar typhimurium str. lt2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PROTEIN FIBRIL |
 |
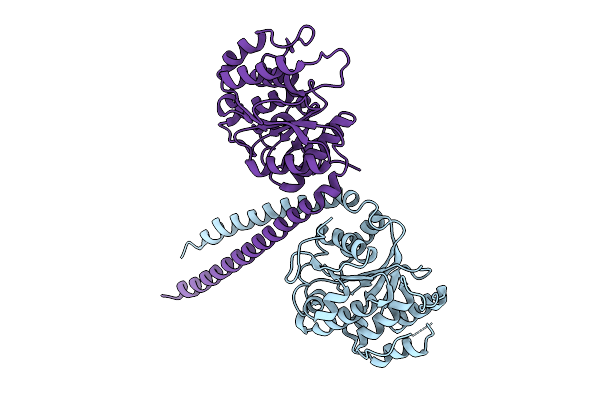
Crystal Structure Of Mctb From Mycobacterium Tuberculosis At 2.3 Angstroms Resolution
Organism: Mycobacterium tuberculosis cdc1551
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: GOL, IMD |
 |
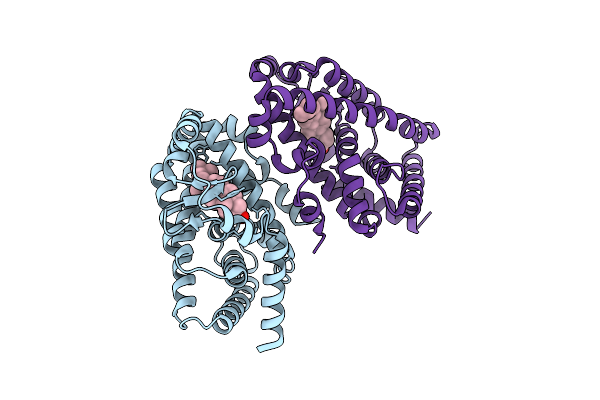
Crystal Structure Of Mctb From Mycobacterium Tuberculosis At 3.25 Angstroms Resolution
Organism: Mycobacterium tuberculosis cdc1551
Method: X-RAY DIFFRACTION Resolution:3.25 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN |
 |
Crystal Structure Of The Mouse Roralpha Ligand Binding Domain In Fusion With An Nrip1 Lxxll Peptide
Organism: Mus musculus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-11 Classification: TRANSCRIPTION Ligands: CLR |
 |
Organism: Mus musculus, Homo sapiens, Bos taurus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN |
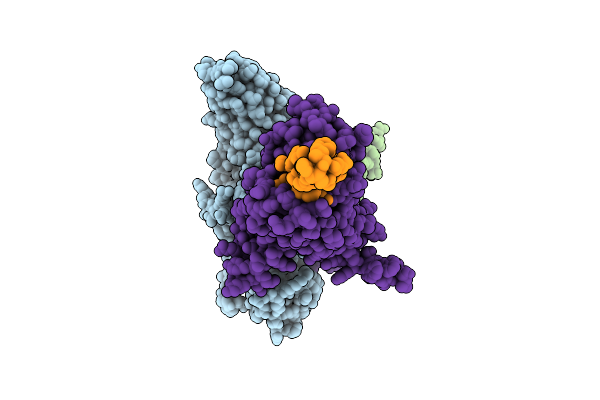 |
Organism: Bos taurus, Mus musculus, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN |
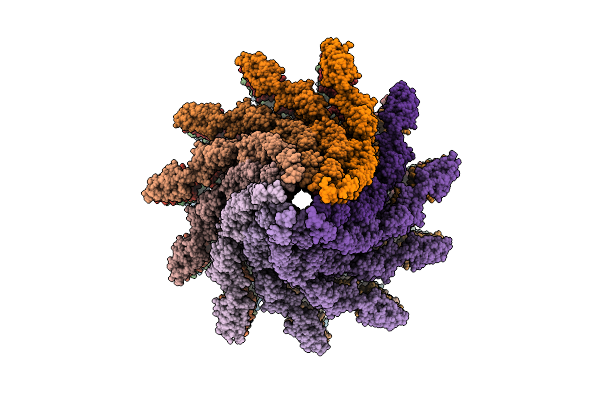 |
Structure Of The Flagellar Filament In Short-Length At 3.02 Angstroms Resolution
Organism: Salmonella enterica subsp. enterica serovar typhimurium str. lt2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: PROTEIN FIBRIL |
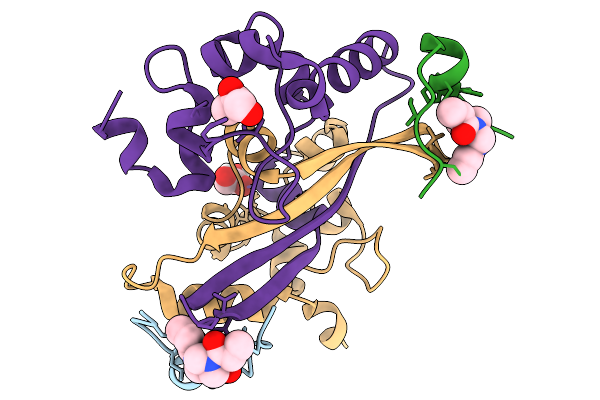 |
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: LFI, GOL |
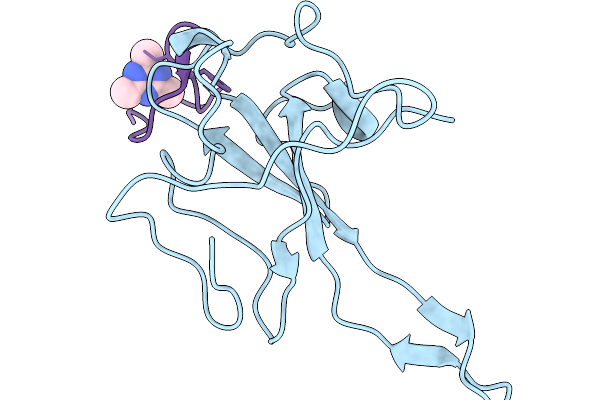 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: KZ0 |
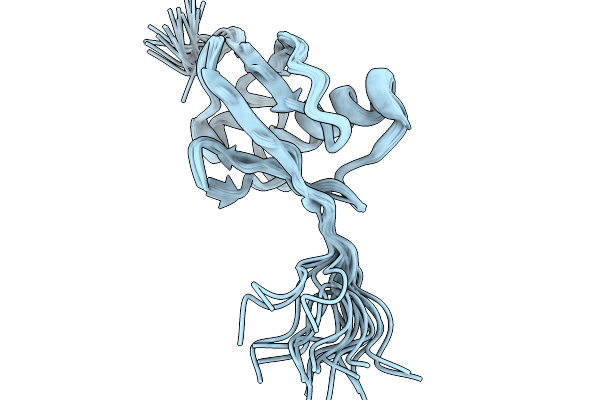 |
Nmr Structure Of Proteinmpnn-Desighed Ubiquitin Variant R4 At Ph 3 With 8 M Urea
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
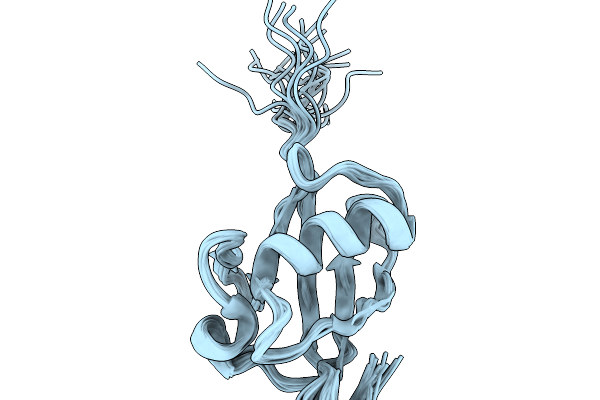 |
Nmr Structure Of Proteinmpnn-Desighed Ubiquitin Variant R4 At Ph 6.3 With 8 M Urea
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |

