Search Count: 2,241
All
Selected
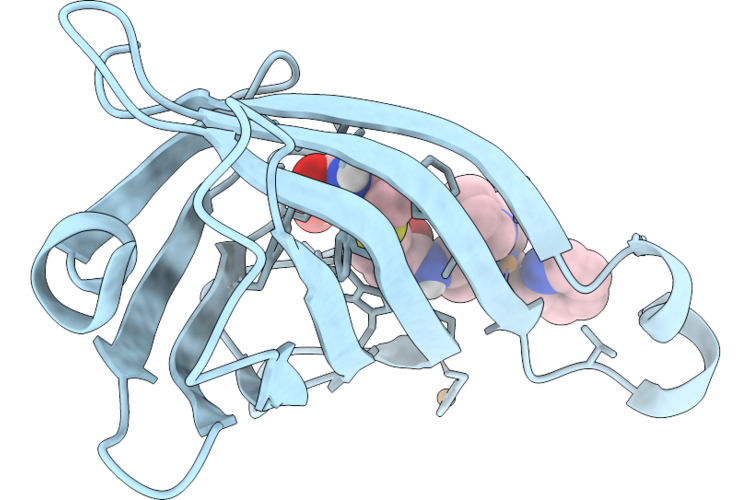 |
Organism: Streptomyces avidinii
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2026-05-20 Classification: METAL BINDING PROTEIN Ligands: A1BIA, CU, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-05-20 Classification: METAL BINDING PROTEIN Ligands: GOL, CU |
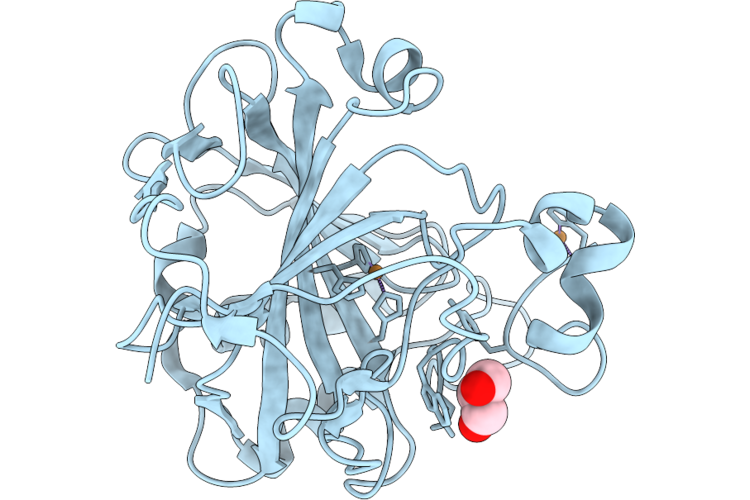 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-05-20 Classification: METAL BINDING PROTEIN Ligands: GOL, CU |
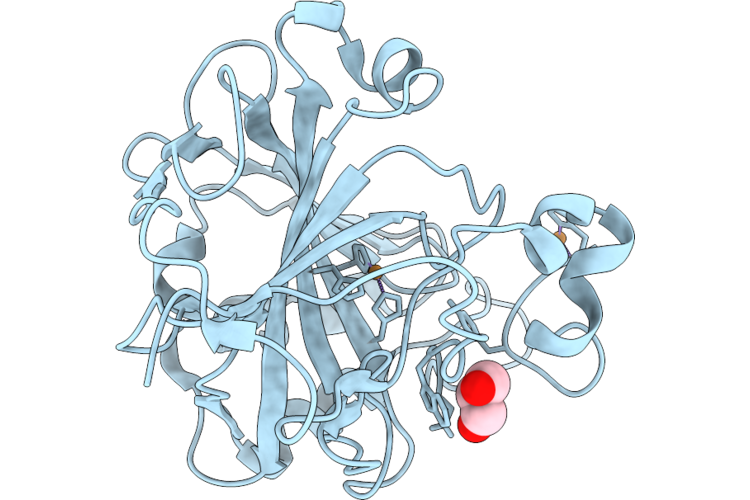 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-05-20 Classification: METAL BINDING PROTEIN Ligands: GOL, CU |
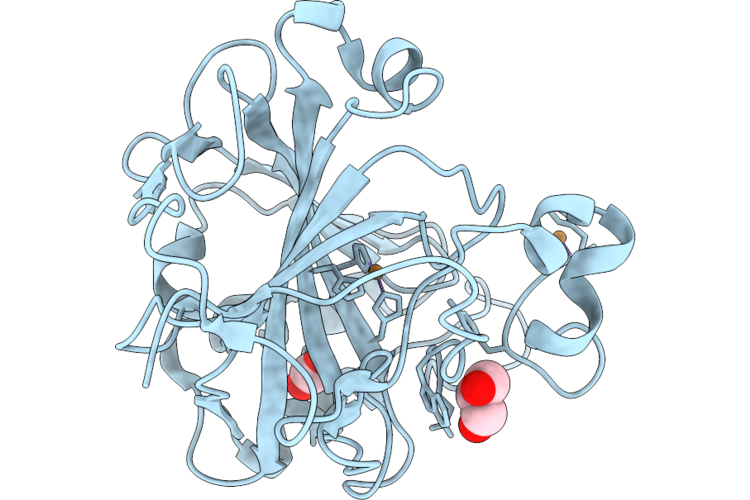 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-05-20 Classification: METAL BINDING PROTEIN Ligands: CO2, GOL, CU |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-05-20 Classification: METAL BINDING PROTEIN Ligands: CO2, GOL, CU |
 |
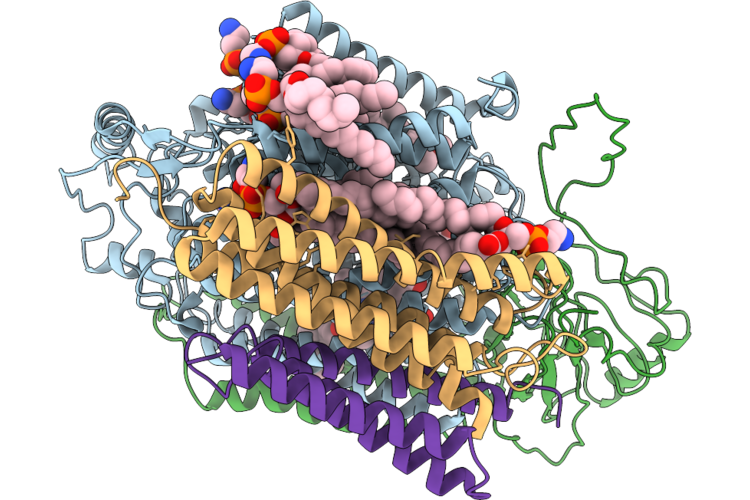
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 8.0.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MEMBRANE PROTEIN Ligands: HEM, FE, CA, LMT, CU, UQ5 |
 |
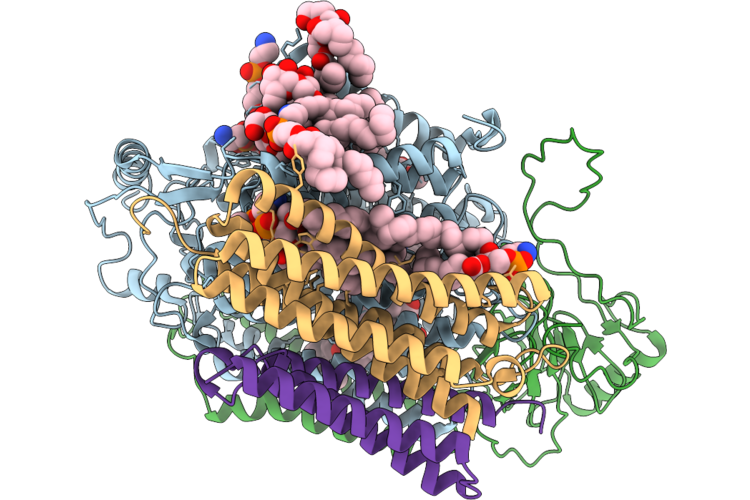
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.05 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: 3PE, HEM, HEO, CU, UQ8 |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.56 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, LMG, UQ8 |
 |
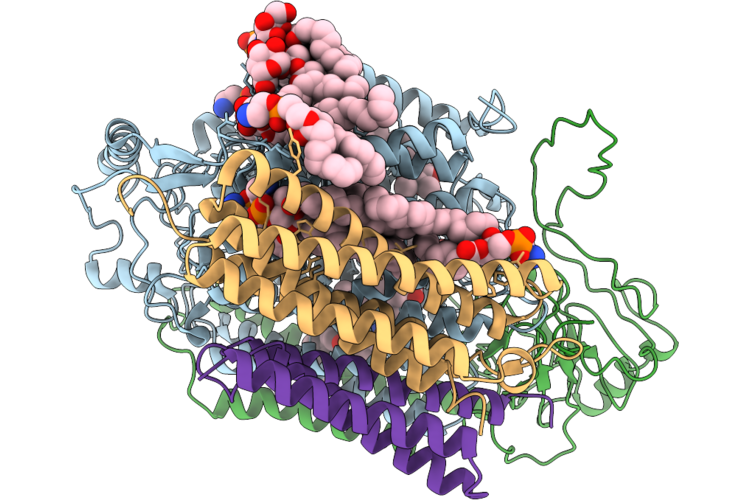
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.12 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, LMG, UQ8 |
 |
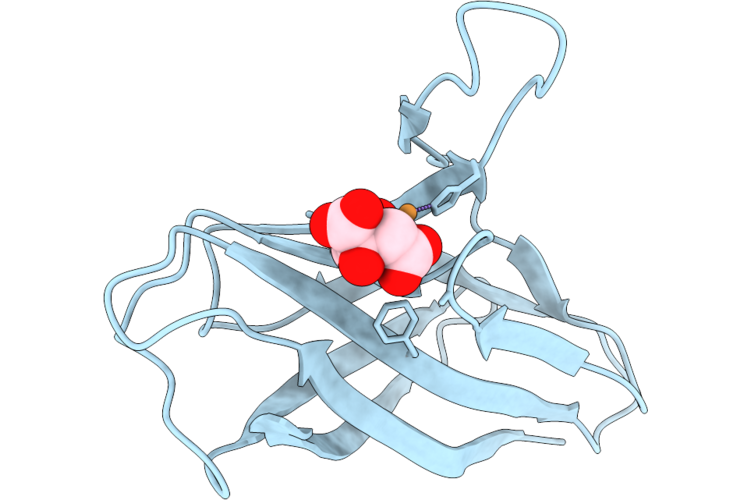
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.28 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, LMG, UQ8 |
 |
Organism: Cereibacter sphaeroides
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-05-06 Classification: CHAPERONE Ligands: CU, FLC |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-04-22 Classification: DE NOVO PROTEIN Ligands: CU, FMT, ACA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: TXP, FAD, CU |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, LMG, UQ8 |
 |
Organism: Arabidopsis thaliana, Escherichia coli k12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: SIGNALING PROTEIN Ligands: NAG, CU |
 |
Cryo-Em Structure Of Complex Iv On The Bovine Heart Submitochondrial Particles, Iv-A
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEA, CU, MG, PER, PGV, CUA, NA, ZN, PEK |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: CU, ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.37 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: CU, ZN |
 |
Atomic Resolution (1.02 A) Xfel Structure Of Nitrite-Bound Copper Nitrite Reductase From Bradyrhizobium Sp. Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium diazoefficiens usda 110
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: CU, NO2, GLC, FRU, SO4 |

