Search Count: 97,310
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: CYTOSOLIC PROTEIN Ligands: A1C4S, ZN |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: XDC, ZN, MG, CA, GOL, NA, NAG, MAN, PO4, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN, MG, CA, NA, NAG, GOL, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
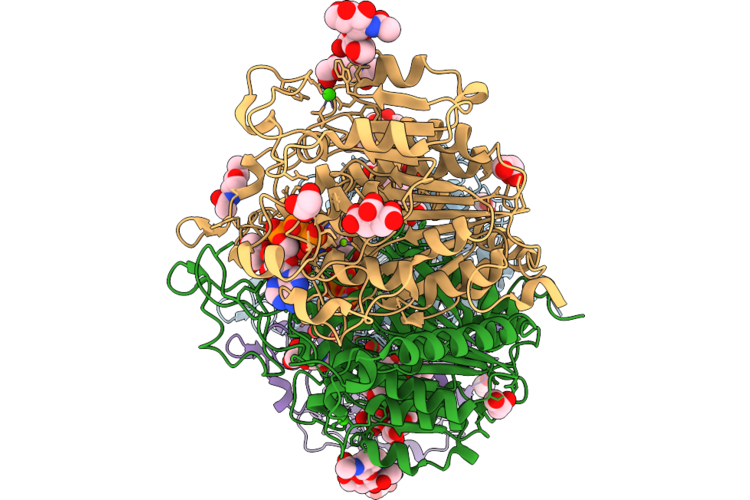
Tissue Non-Specific Alkaline Phosphatase -S110A Bound To Ppi With Ethylene Glycol
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: POP, ZN, MG, CA, NA, NAG, MAN, PO4, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ATP, ZN, MG, CA, GOL, NAG, FLC, PO4 |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.28 Å Release Date: 2026-04-22 Classification: DNA BINDING PROTEIN |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.81 Å Release Date: 2026-04-22 Classification: DNA BINDING PROTEIN |
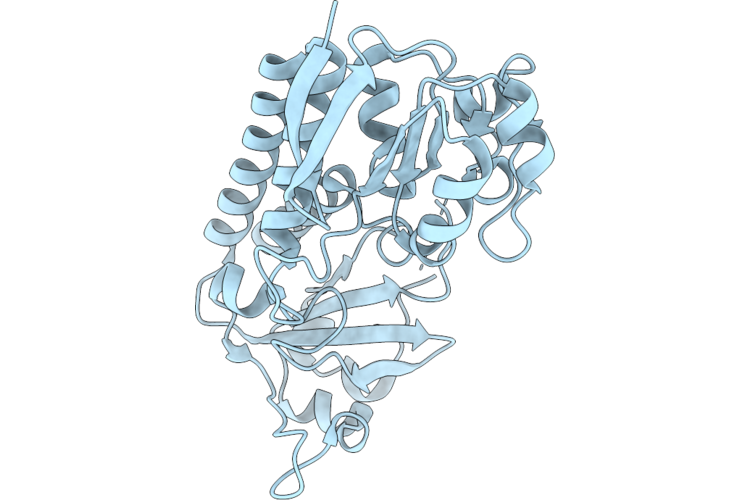 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus In The Apo-Form
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN |
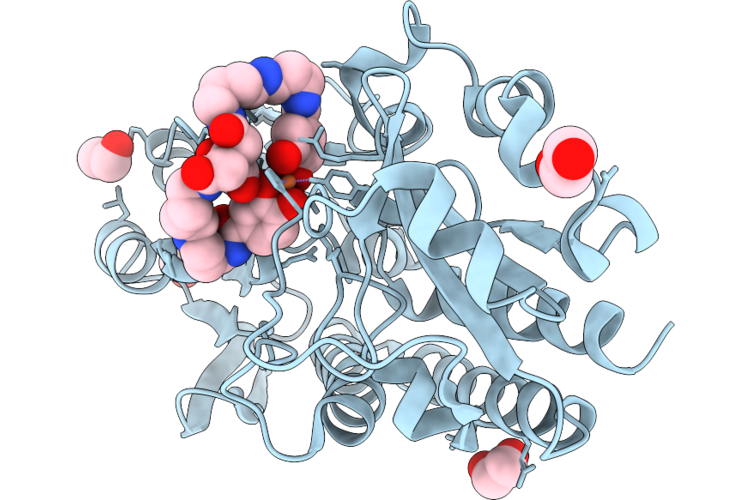 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Petrobactin
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: F8W, FE, EDO |
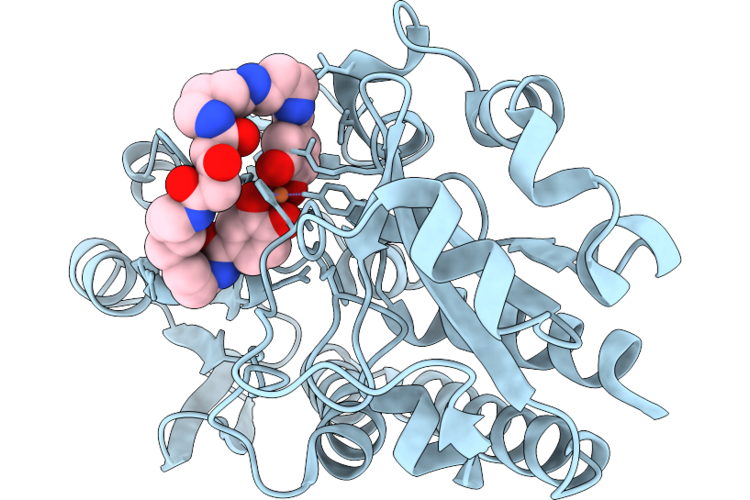 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Petrobactin Photoproduct, Fepbv
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: A1MDH, FE |
 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Siderophore Mimic, Fe(3,4-Dhb)2
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: DHB, FE, EDO, IPA |
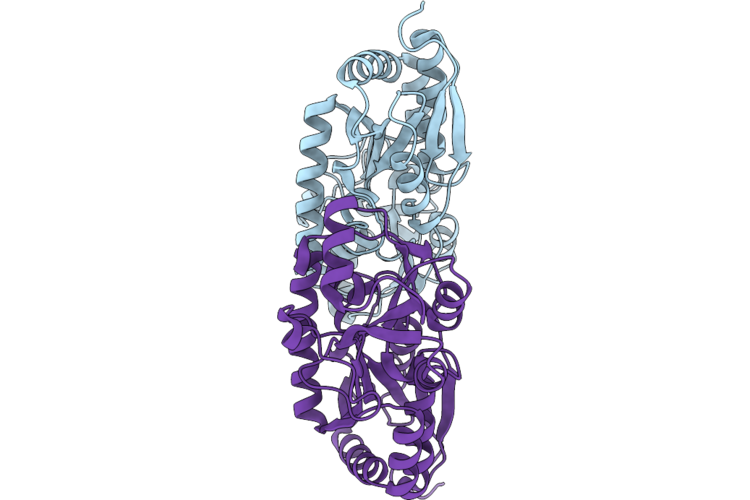 |
Crystal Structure Of A Tctc Solute Binding Protein From Vibrio Sp. C42, No Ligand
Organism: Vibrio atlanticus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING |
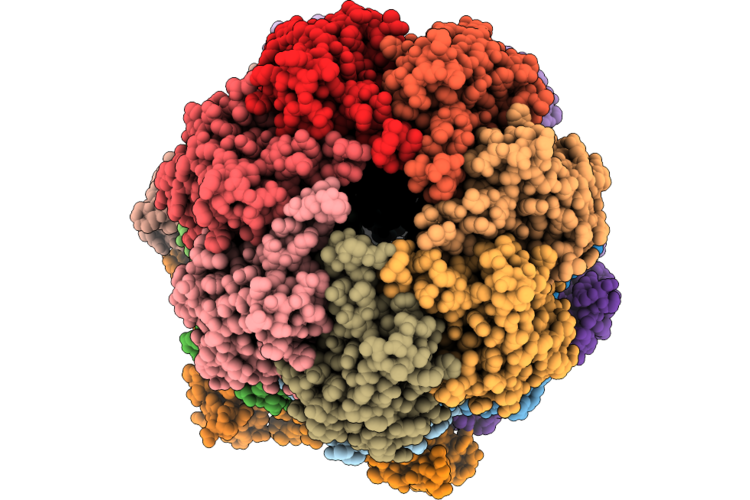 |
Organism: Enterobacter
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-22 Classification: CHAPERONE Ligands: ATP |
 |
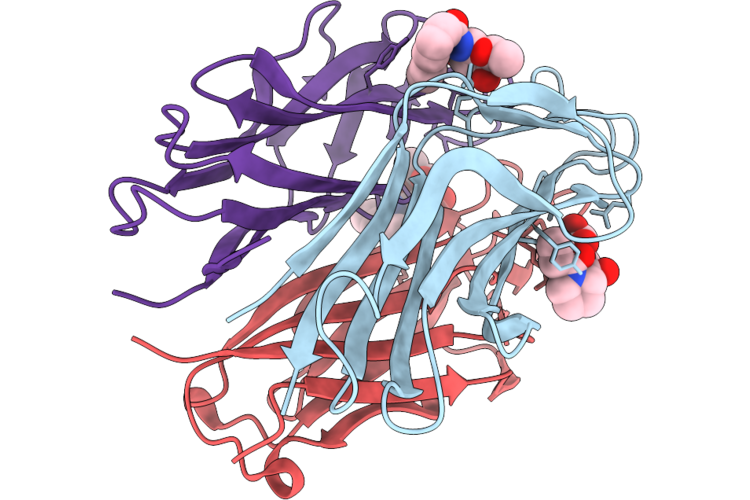
N Terminal Domain Of Bc2L-C Lectin (1-131) In Complex With A Beta-Fucosylamide Side-Product
Organism: Burkholderia cenocepacia j2315
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2026-04-22 Classification: SUGAR BINDING PROTEIN Ligands: A1IRO |
 |
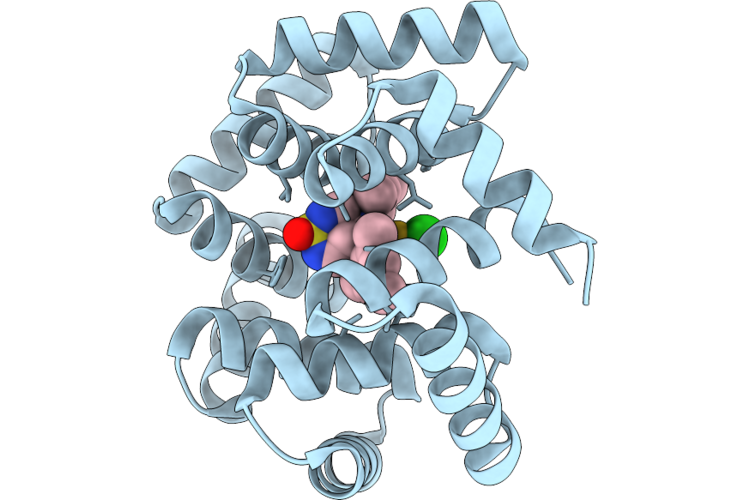
N Terminal Domain Of Bc2L-C Lectin (1-131) With Covalent Beta-Fucosylamide Ligand
Organism: Burkholderia cenocepacia j2315
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-22 Classification: SUGAR BINDING PROTEIN Ligands: A1IRP |
 |
Crystal Structure Of A Computationally Designed Protein Bound To A Au-Containing Cofactor ([(Sulfanhc)Au.Trp])
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-22 Classification: DE NOVO PROTEIN Ligands: A1IRM |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PEG, DSN, CA |
 |
Crystal Structure Of Gh19 E228Q Domain Of D29-Lysa In Complex With (Glcnac)4
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, MPD |

