Search Count: 62
All
Selected
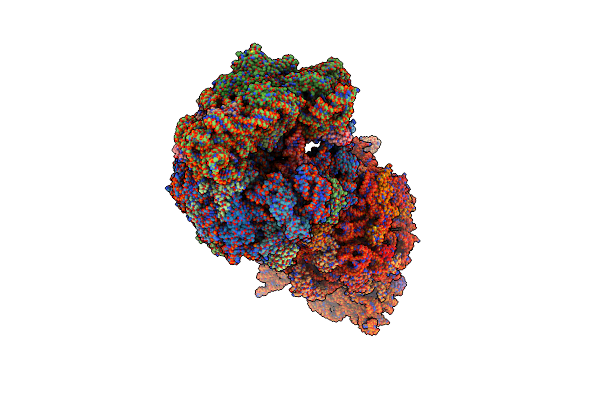 |
Crystal Structure Of The Wild-Type Thermus Thermophilus 70S Ribosome In Complex With Spectinomycin, Mrna, Deacylated A- And E-Site Trnaphe, And Deacylated P-Site Trnamet At 2.60A Resolution
Organism: Escherichia coli, Escherichia phage t4, Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2024-08-07 Classification: RIBOSOME/INHIBITOR,ANTIBIOTIC Ligands: MG, K, ZN, SCM, SF4 |
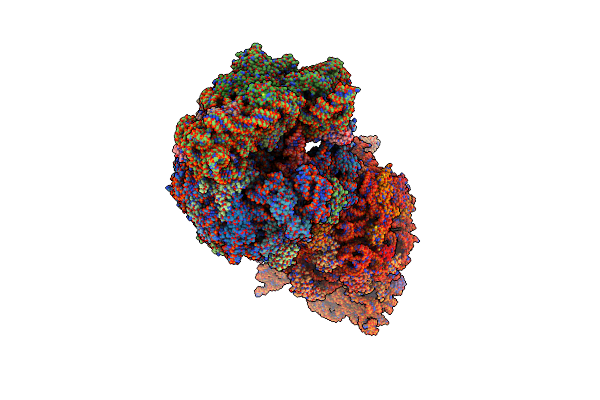 |
Crystal Structure Of The Wild-Type Thermus Thermophilus 70S Ribosome In Complex With Spectinomycin Derivative 2694, Mrna, Deacylated A- And E-Site Trnaphe, And Deacylated P-Site Trnamet At 2.75A Resolution
Organism: Escherichia coli, Escherichia phage t4, Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2024-08-07 Classification: RIBOSOME/INHIBITOR Ligands: MG, ZN, Y7K, SF4, K |
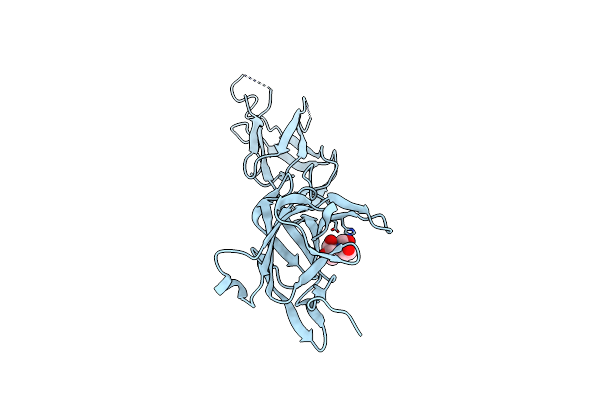 |
Crystal Structure Of Srtb-Anchored Collagen-Binding Adhesin Fragment (Residues 206-565) From Clostridioides Difficile Strain 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2021-10-20 Classification: CELL ADHESION Ligands: BTB |
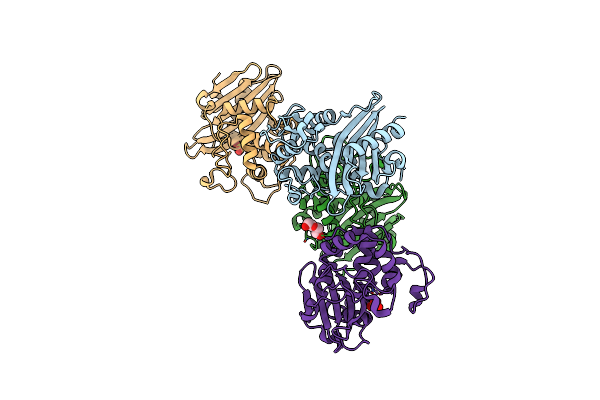 |
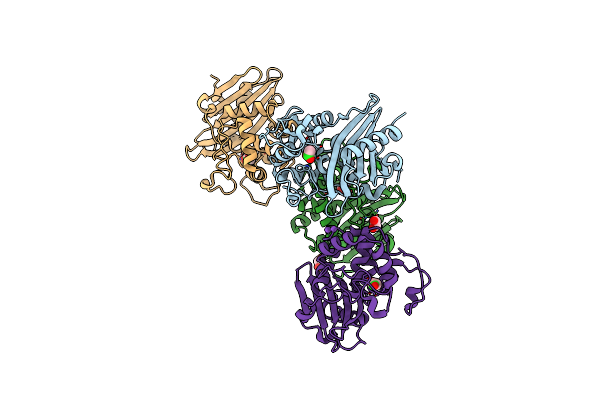
Crystal Structure Of C79A Mutant Of Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2021-08-11 Classification: HYDROLASE Ligands: PEG, SO4 |
 |
Crystal Structure Of K83A Mutant Of Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2021-08-11 Classification: HYDROLASE Ligands: CL, ACT, EDO, NA |
 |
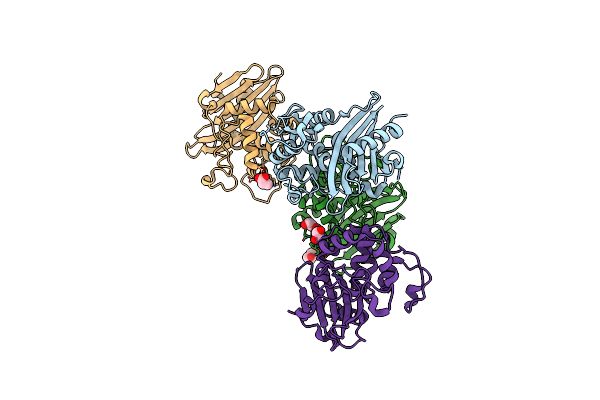
Crystal Structure Of Wild Type Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2020-05-27 Classification: HYDROLASE Ligands: PEG, NA |
 |
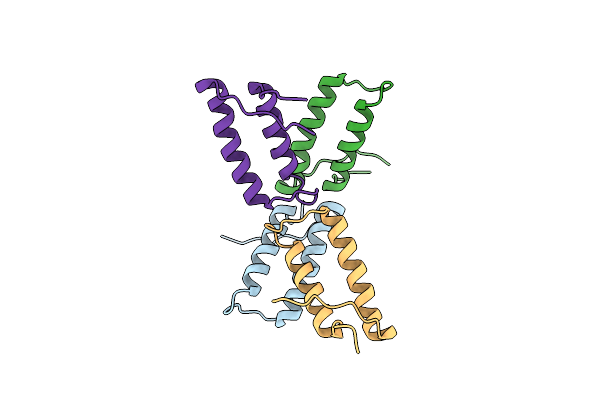
1.85 Angstrom Resolution Crystal Structure Of Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Peptoclostridium difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2019-12-25 Classification: HYDROLASE Ligands: PGE, PEG, PPI, GOL |
 |
1.78 Angstrom Resolution Crystal Structure Of Hypothetical Protein Cd630_05490 From Clostridioides Difficile 630.
Organism: Peptoclostridium difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2018-12-12 Classification: UNKNOWN FUNCTION |
 |
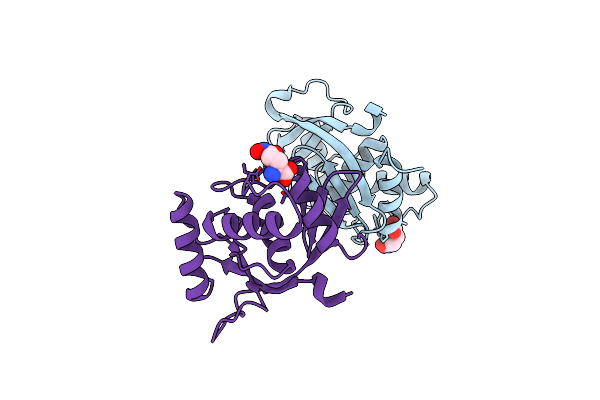
A 1.85A X-Ray Structure From Peptoclostridium Difficile 630 Of A Hypothetical Protein
Organism: Peptoclostridium difficile
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2017-02-01 Classification: STRUCTURAL GENOMICS, UNKNOWN FUNCTION |
 |
2.05 Angstrom Resolution Crystal Structure Of Peptidoglycan-Binding Protein From Clostridioides Difficile In Complex With Glutamine Hydroxamate.
Organism: Peptoclostridium difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2016-12-14 Classification: HYDROLASE Ligands: HGA |
 |
1.95 Angstrom Resolution Crystal Structure Of Stage Ii Sporulation Protein D (Spoiid) From Clostridium Difficile In Apo Conformation
Organism: Peptoclostridium difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2016-12-14 Classification: HYDROLASE Ligands: ZN, NA, CL, FMT, DMS |
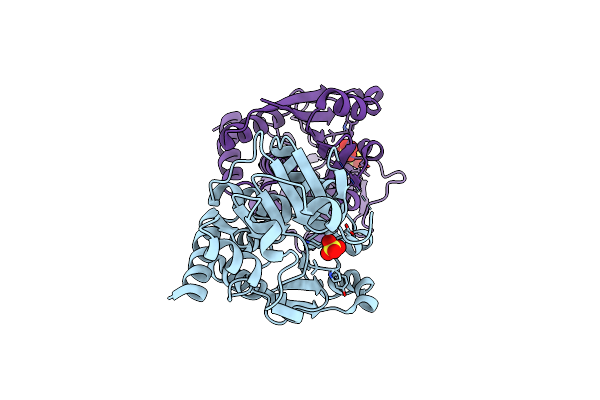 |
1.5 Angstrom Crystal Structure Of Shikimate Dehydrogenase 1 From Peptoclostridium Difficile.
Organism: Peptoclostridium difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2015-10-07 Classification: OXIDOREDUCTASE Ligands: SO4 |
 |
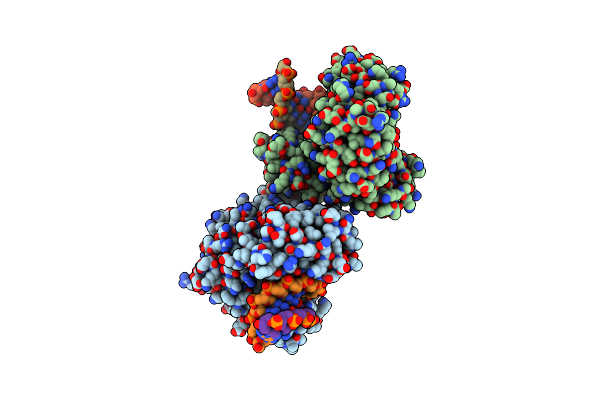
Ternary Complex Of Y-Family Dna Polymerase Dpo4 With (5'S)-8,5'-Cyclo-2'-Deoxyguanosine And Dttp
Organism: Sulfolobus solfataricus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2015-01-14 Classification: TRANSFERASE/DNA Ligands: MG, DOC, CA, TTP |
 |
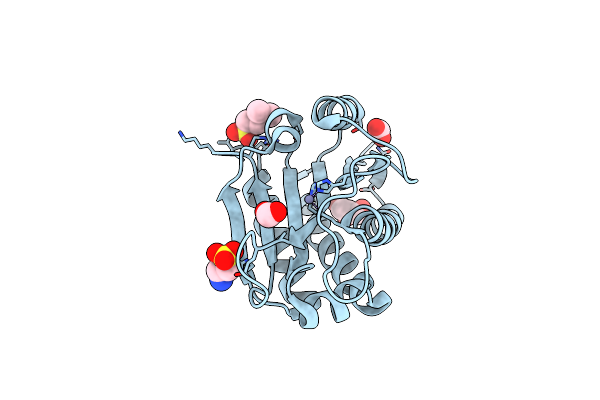
Ternary Complex Of Y-Family Dna Polymerase Dpo4 With (5'S)-8,5'-Cyclo-2'-Deoxyguanosine And Dctp
Organism: Sulfolobus solfataricus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2015-01-14 Classification: TRANSFERASE/DNA Ligands: DCP, MG, DOC |
 |
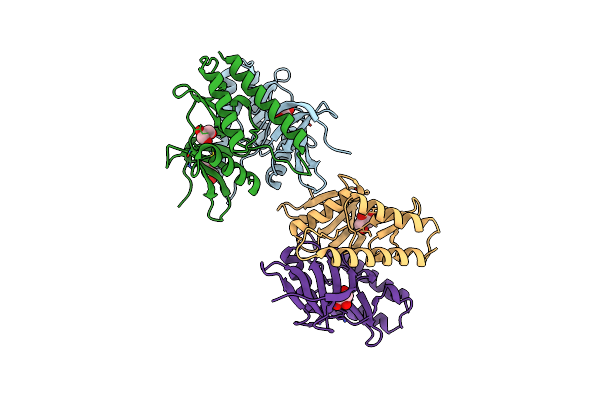
The Crystal Structure Of N-Acetylmuramoyl-L-Alanine Amidase From Clostridium Difficile 630
Organism: Peptoclostridium difficile
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2014-11-05 Classification: HYDROLASE Ligands: ZN, GOL, EPE, FMT |
 |
2.60 Angstrom Resolution Crystal Structure Of A Protein Kinase Domain Of Type Iii Effector Nleh2 (Ecs1814) From Escherichia Coli O157:H7 Str. Sakai
Organism: Escherichia coli o157:h7
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2014-01-15 Classification: HYDROLASE Ligands: GOL, PEG |
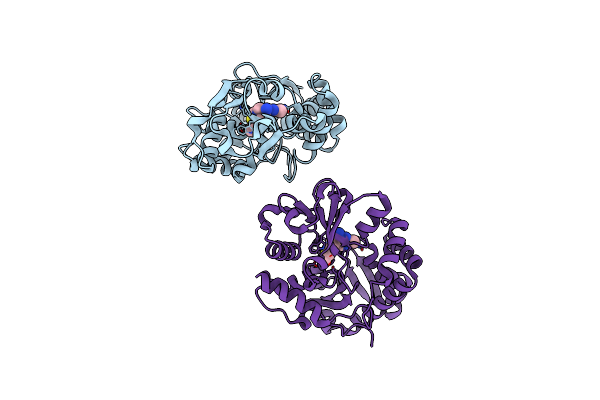 |
Crystal Structure Of The Thiamin-Bound Form Of Substrate-Binding Protein Of Abc Transporter From Clostridium Difficile
Organism: Clostridium difficile
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2013-12-04 Classification: TRANSPORT PROTEIN Ligands: VIB |
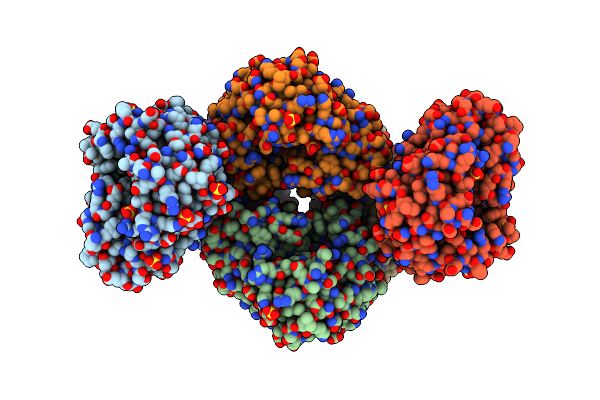 |
Organism: Yersinia pestis
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2013-10-16 Classification: OXIDOREDUCTASE Ligands: SO4 |
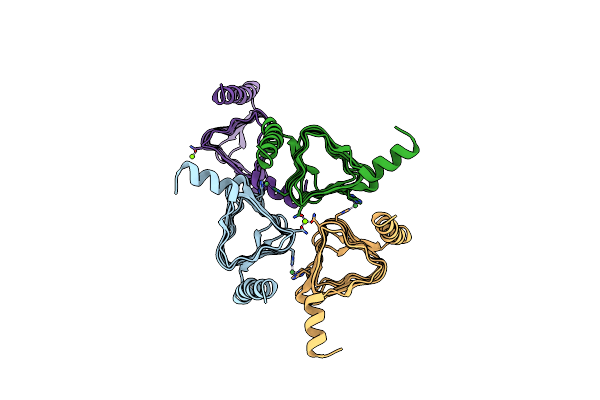 |
2.0 Angstrom Resolution Crystal Structure Of Putative Carbonic Anhydrase From Clostridium Difficile.
Organism: Clostridium difficile
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2013-09-04 Classification: TRANSFERASE Ligands: MG, NI |
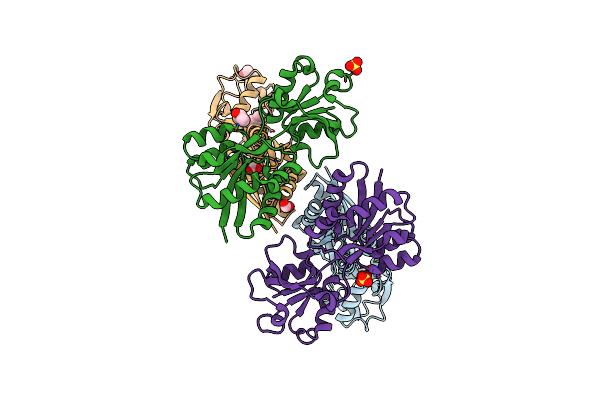 |
Substrate Binding Domain Of Putative Molybdenum Abc Transporter From Clostridium Difficile
Organism: Clostridium difficile
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2013-05-08 Classification: TRANSPORT PROTEIN Ligands: FMT, SBT, SO4 |

