Search Count: 4,497
All
Selected
 |
Organism: Streptomyces fagopyri
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-03-18 Classification: BIOSYNTHETIC PROTEIN |
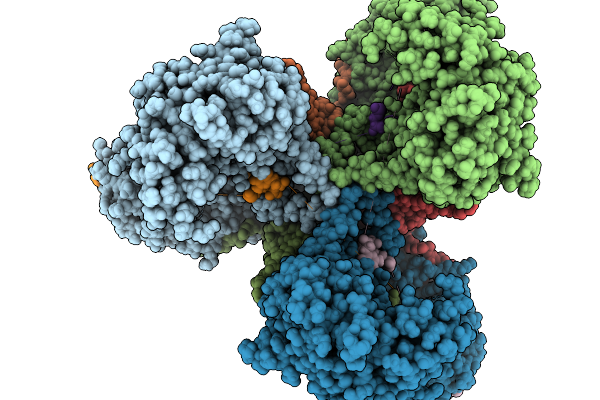 |
Organism: Escherichia coli nctc 86
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-02-18 Classification: ANTIVIRAL PROTEIN Ligands: MG |
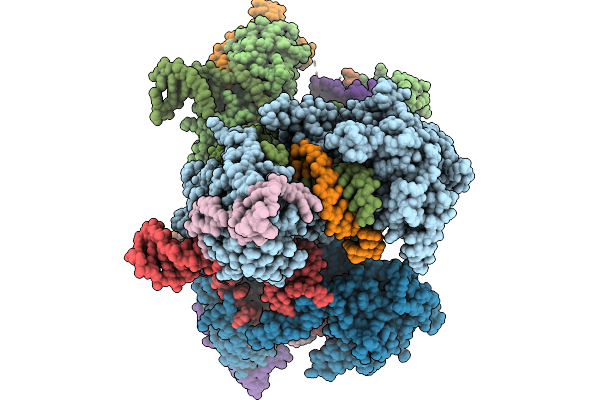 |
Organism: Escherichia coli nctc 86
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-02-04 Classification: ANTIVIRAL PROTEIN |
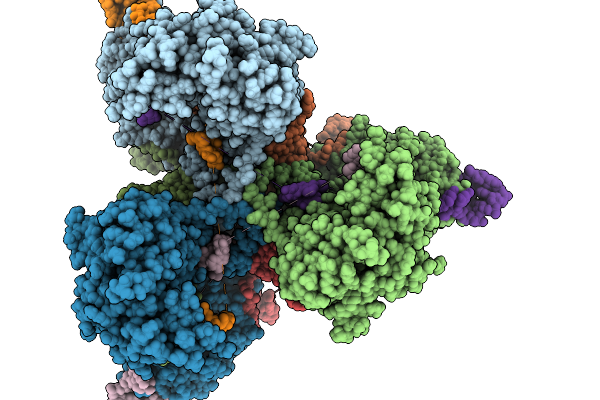 |
Organism: Escherichia coli nctc 86
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-02-04 Classification: ANTIVIRAL PROTEIN Ligands: MG |
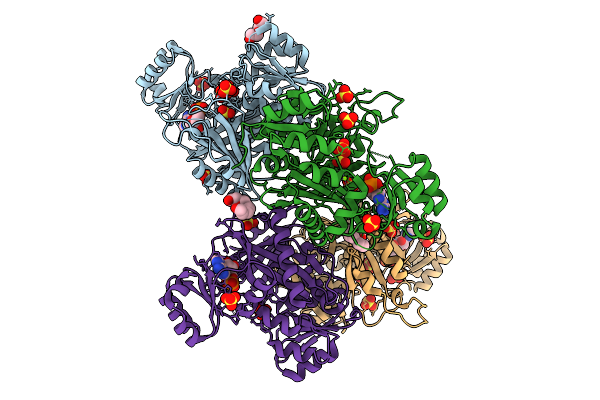 |
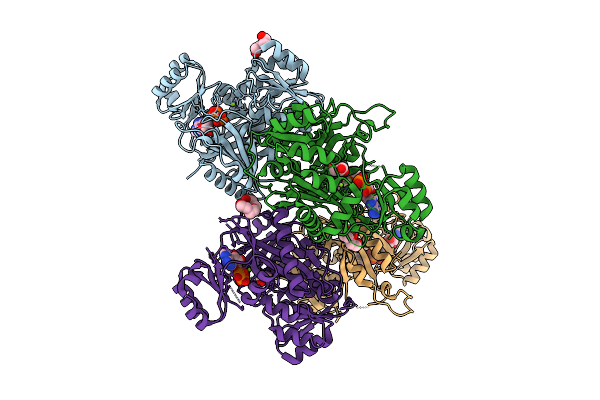
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Amp, Adp And Sulfate Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2026-01-21 Classification: LYASE Ligands: AMP, PEG, MG, SO4, PG4, ADP |
 |
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Atp Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-12-24 Classification: LYASE Ligands: ATP, PEG, MG, PGE, GOL, ADP, PG4 |
 |
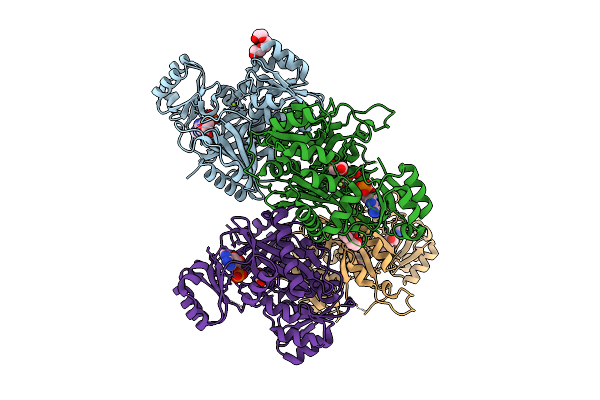
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Adp Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2025-12-24 Classification: LYASE Ligands: ADP, PEG, MG, PGE, GOL, PG4 |
 |
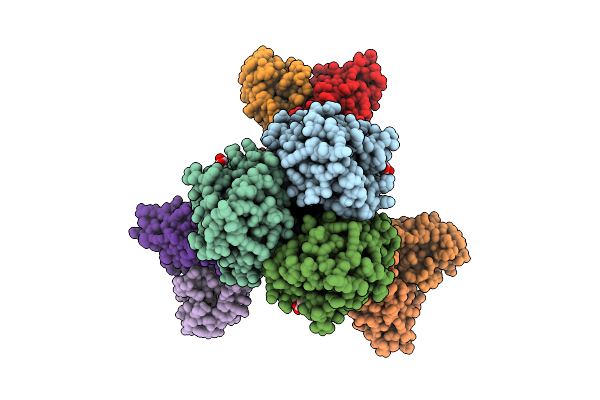
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Amp Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2025-12-24 Classification: LYASE Ligands: AMP, PGE, MG, PEG, PG4, GOL |
 |
Organism: Homo sapiens, Influenza a virus (a/california/01/2009(h1n1))
Method: ELECTRON MICROSCOPY Resolution:4.10 Å Release Date: 2025-11-12 Classification: IMMUNE SYSTEM/VIRAL PROTEIN Ligands: NAG |
 |
Organism: Homo sapiens, Influenza a virus (a/california/01/2009(h1n1))
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-11-12 Classification: IMMUNE SYSTEM/VIRAL PROTEIN Ligands: NAG |
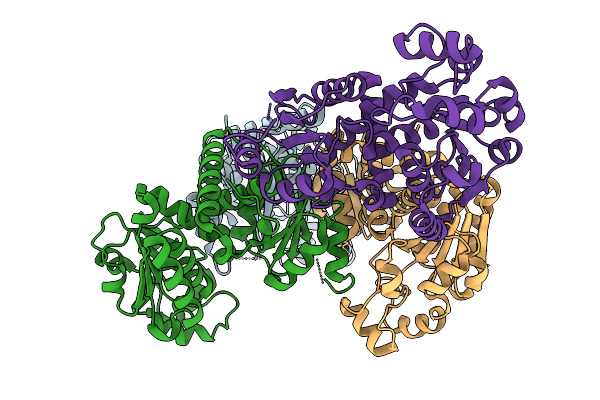 |
Crystal Structure Of Ornithine Carbamoyltransferase From Burkholderia Xenovorans
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2025-09-24 Classification: TRANSFERASE Ligands: CL |
 |
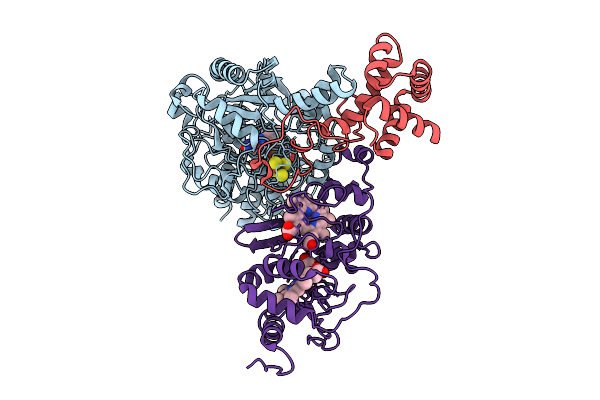
Crystal Structure Of Ornithine Carbamoyltransferase From Burkholderia Xenovorans In Complex With Phosphono Carbamate
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.67 Å Release Date: 2025-09-24 Classification: TRANSFERASE Ligands: CP, PO4, CL |
 |
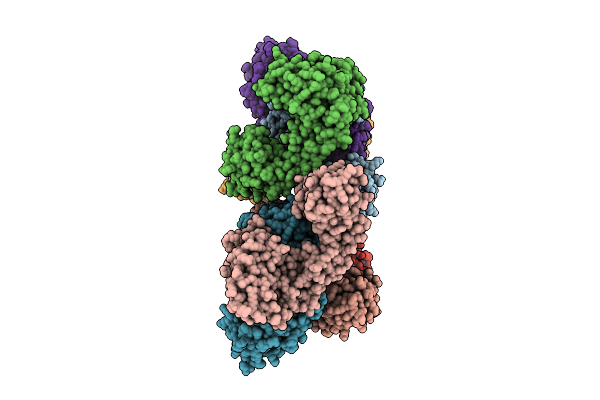
Cryo-Em Structure Of Fructose Dehydrogenase Variant From Gluconobacter Japonicus Truncating Heme 1C And C-Terminal Hydrophobic Regions
Organism: Gluconobacter japonicus
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: OXIDOREDUCTASE Ligands: FAD, F3S, HEC |
 |
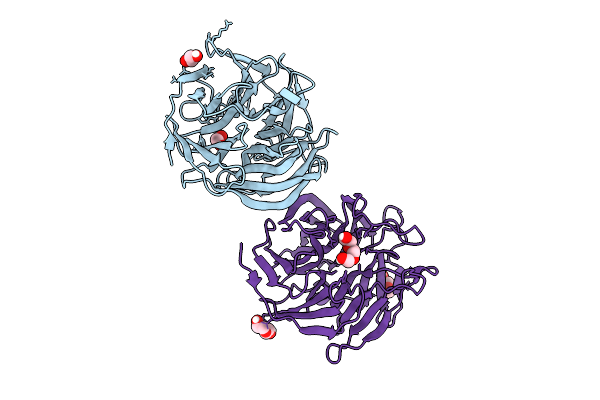
Organism: Corynebacterium glutamicum atcc 13032
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-07-30 Classification: CELL CYCLE |
 |
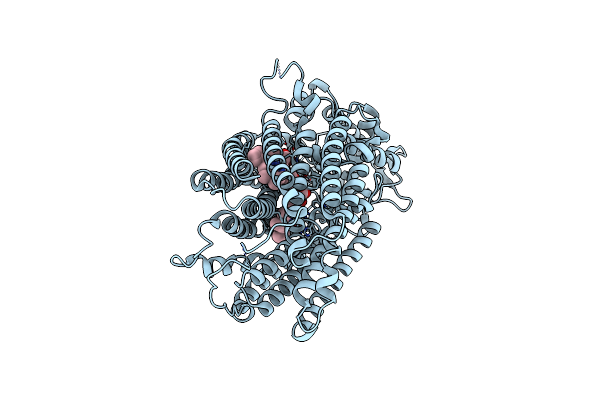
Crystal Structure Of Complement1.4 (Cent1.4), An Engineered Photoenzyme For Selective [2+2]-Cycloadditions
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, PEG, PGE |
 |
Cryoem Structure Of Monomeric Quinol Dependent Nitric Oxide Reductase From Neisseria Meningitidis
Organism: Neisseria meningitidis alpha14
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: OXIDOREDUCTASE Ligands: HEM, FE, CA |
 |
Cryoem Structure Of Dimeric Quinol Dependent Nitric Oxide Reductase From Neisseria Meningitidis
Organism: Neisseria meningitidis alpha14
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: OXIDOREDUCTASE Ligands: HEM, FE, CA |
 |
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2025-05-14 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO |
 |
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-05-14 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, EDO |
 |
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-05-14 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, PGE, MG |

