Search Count: 3,135
All
Selected
 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: REPLICATION Ligands: MG, ZN, ADP, ATP |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: REPLICATION Ligands: ZN, ADP, MG |
 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: REPLICATION Ligands: ZN, ADP, MG |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: REPLICATION Ligands: ZN, ADP, MG |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-05-13 Classification: REPLICATION Ligands: ZN, ADP, MG |
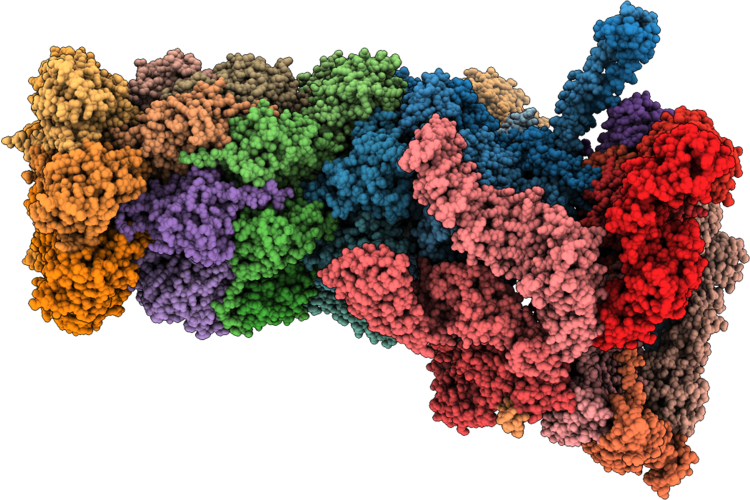 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ea2.1
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
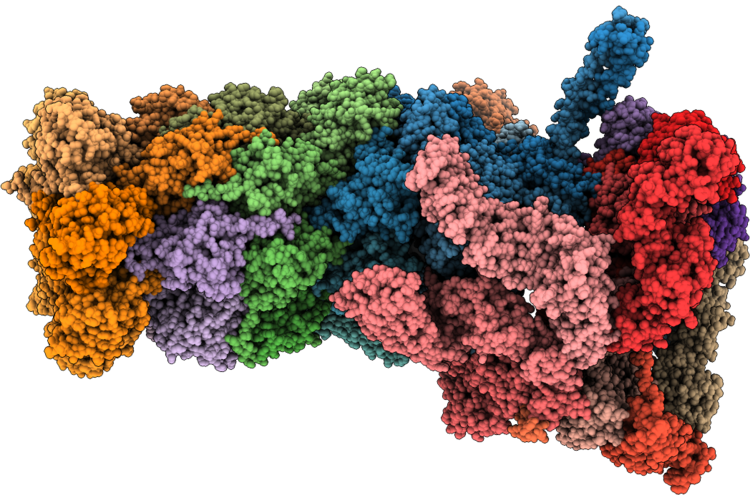 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ea2.2
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
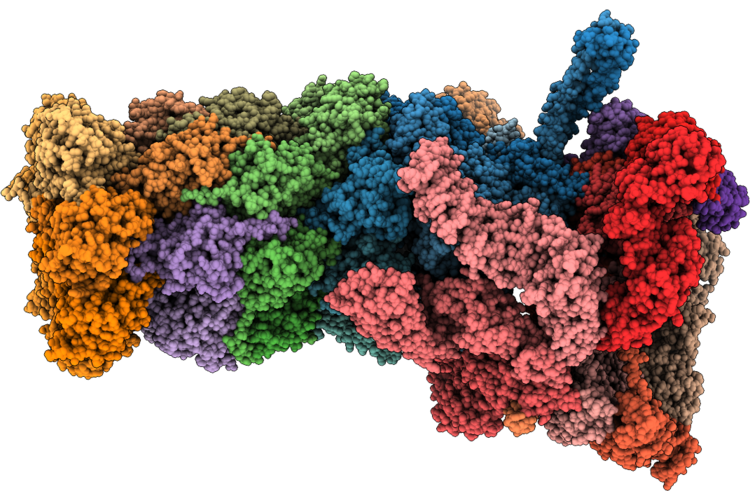 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ea2.3
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Eb.1
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
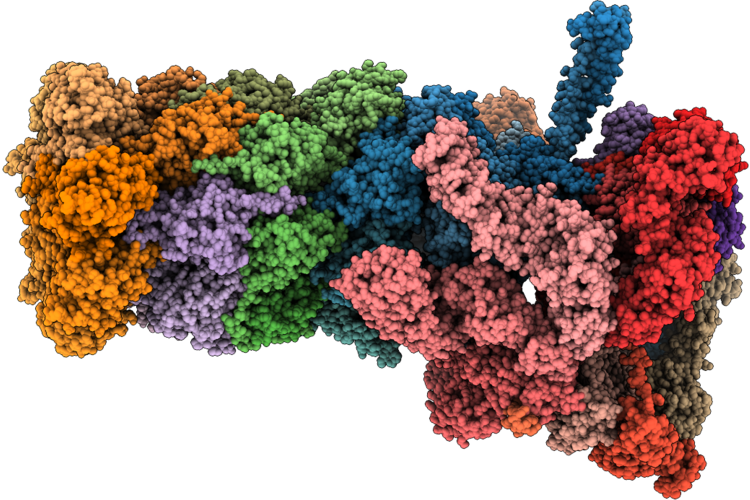 |
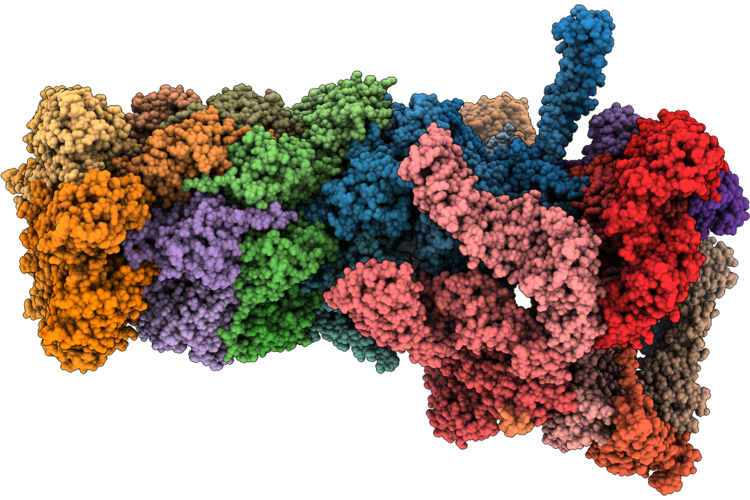
Structure Of Substrate-Engaged 26S Proteasome Rp-Cp Subcomplex In State Eb.2
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
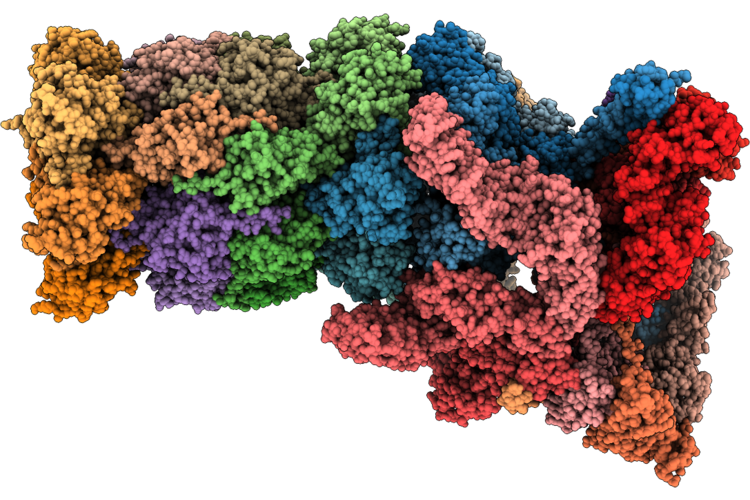
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Eb.3
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
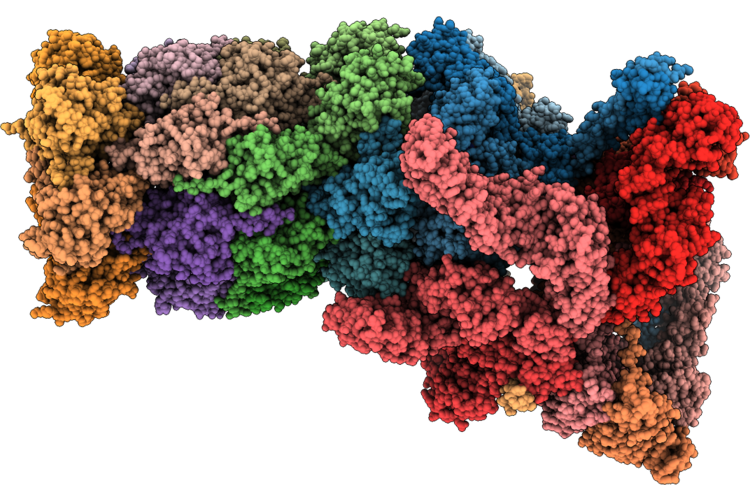
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ec1
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ADP, ATP, MG, ZN |
 |
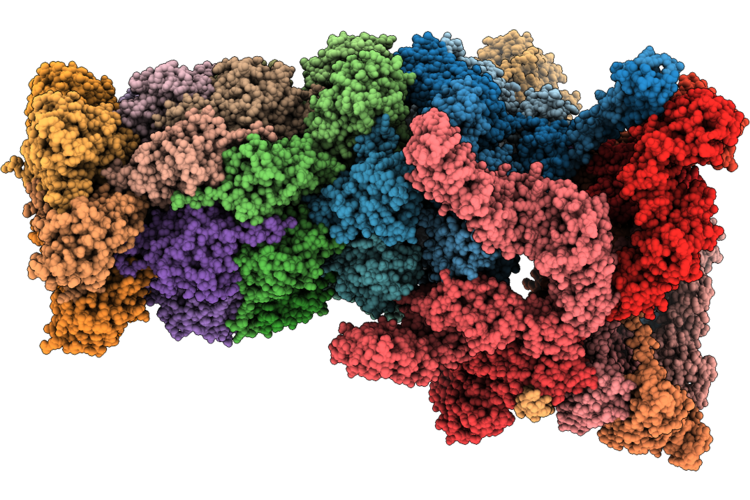
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ec2
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ed0.1
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ed0.2
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ed0.3
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ed1.1
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ed1.2
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ed1.3
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ed2.1
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |

