Search Count: 1,736
All
Selected
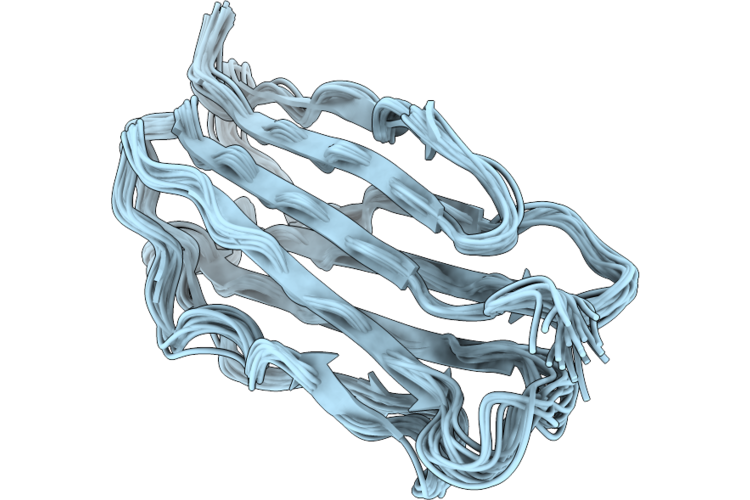 |
Organism: Mattirolomyces terfezioides
Method: SOLUTION NMR Release Date: 2026-04-22 Classification: UNKNOWN FUNCTION |
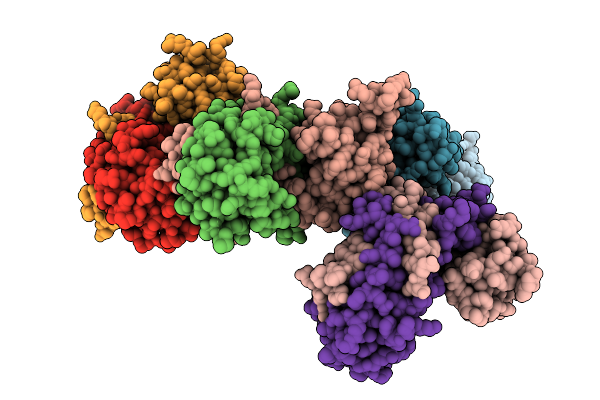 |
Structural Insights Into Hpar-Mediated Recognition Of Hrpx And Hrpg In Xanthomonas Campestris Pv. Campestris
Organism: Xanthomonas campestris pv. campestris (strain atcc 33913 / dsm 3586 / ncppb 528 / lmg 568 / p 25)
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-03-11 Classification: TRANSCRIPTION |
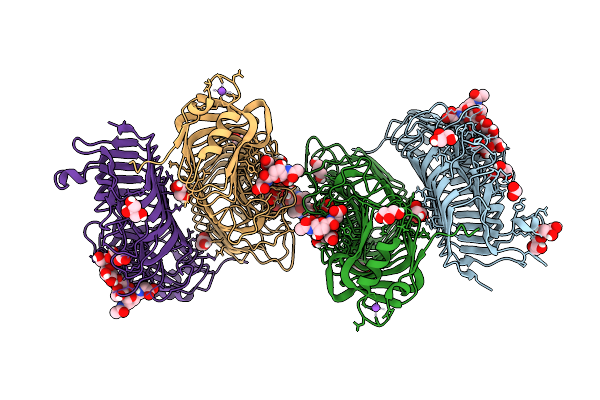 |
Crystal Structure Of The Polysaccharide Lyase Rbmb From Vibrio Cholerae Bound To Vibrio Polysaccharide
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-03-11 Classification: LYASE |
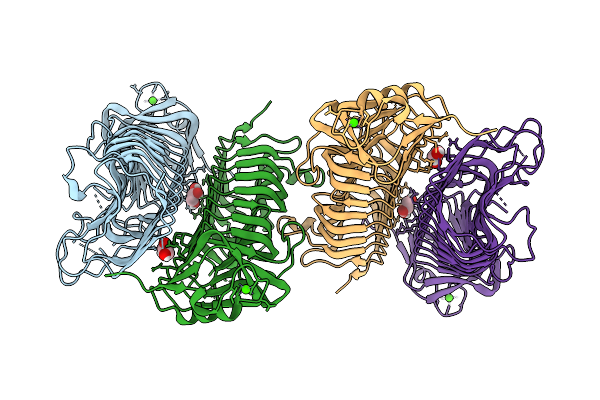 |
Organism: Vibrio cholerae c6706
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: LYASE Ligands: GOL, CA |
 |
Organism: Prymnesium parvum
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-11 Classification: LIGASE |
 |
Organism: Neisseria gonorrhoeae
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: FE |
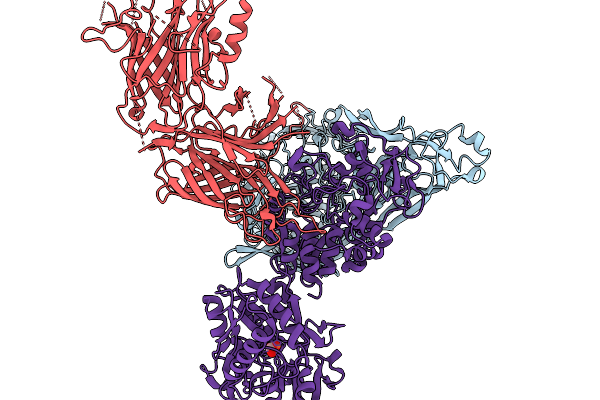 |
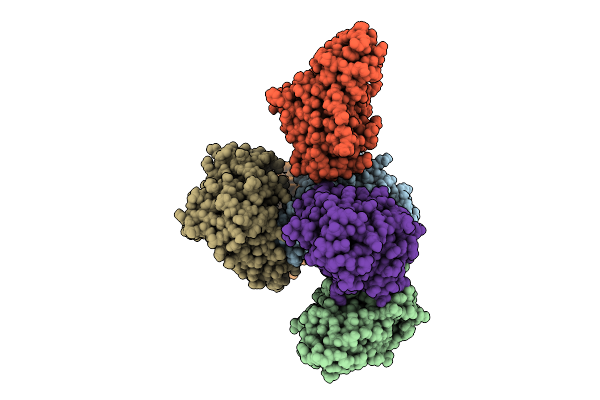
Neisseria Gonorrhoeae Transferrin Binding Protein A In Complex With Transferrin Binding Protein B And Transferrin (Iron Bound In N Lobe Only)
Organism: Homo sapiens, Neisseria gonorrhoeae, Neisseria meningitidis serogroup b
Method: ELECTRON MICROSCOPY Resolution:4.26 Å Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN Ligands: BCT, FE |
 |
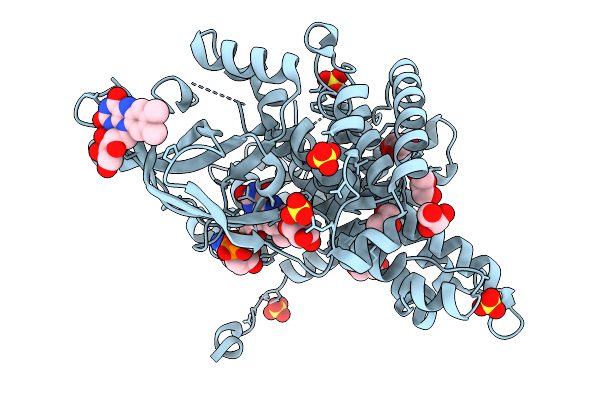
Crystal Structure Of The Transpeptidase Domain Of Pbp2 From The Neisseria Gonorrhoeae Cephalosporin-Resistant Strain H041 In Complex With Boronate Inhibitor Vnrx-14079
Organism: Neisseria gonorrhoeae
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-02-04 Classification: LIGASE Ligands: A1C02 |
 |
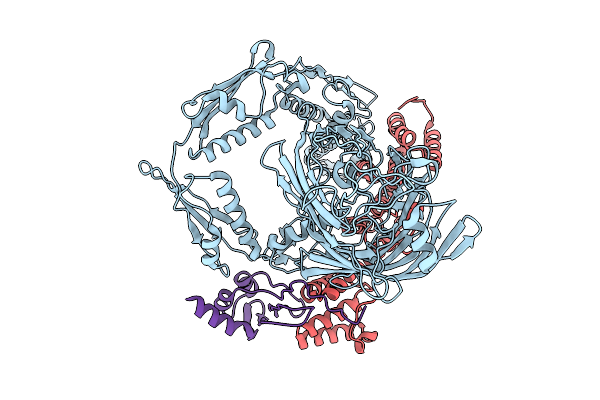
Organism: Goodfellowiella coeruleoviolacea
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-09-10 Classification: OXIDOREDUCTASE Ligands: FAD, SO4, GOL |
 |
Organism: Neisseria gonorrhoeae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN |
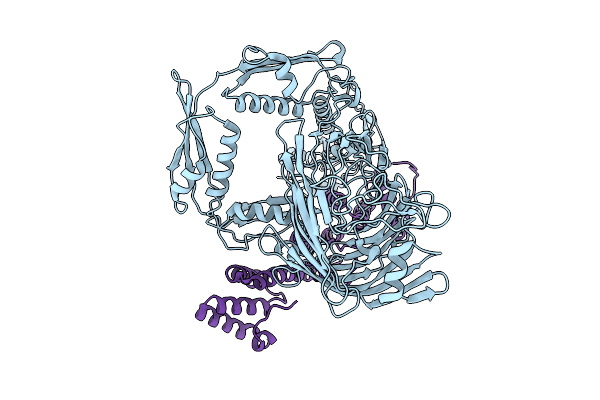 |
Organism: Neisseria gonorrhoeae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN |
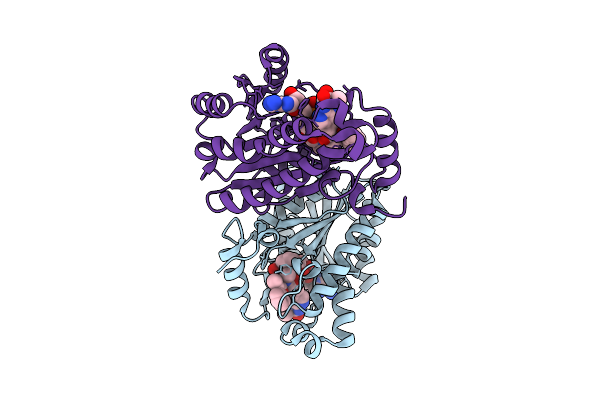 |
Crystal Structure Of Neisseria Gonorrhoeae Fabi In Complex With Nadh And (E)-3-((2R,3S)-3-Hydroxy-2-Methyl-4-Oxo-2,3,4,5-Tetrahydro-1H-Pyrido[2,3-B][1,4]Diazepin-8-Yl)-N-Methyl-N-((3-Methylbenzofuran-2-Yl)Methyl)Acrylamide
Organism: Neisseria gonorrhoeae
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: A1H1H, NAD |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: CL, ETA |
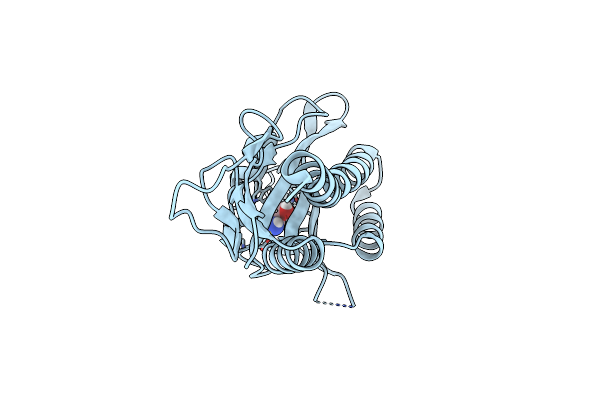 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: 2A1, CL |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: SEL, CL |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN |
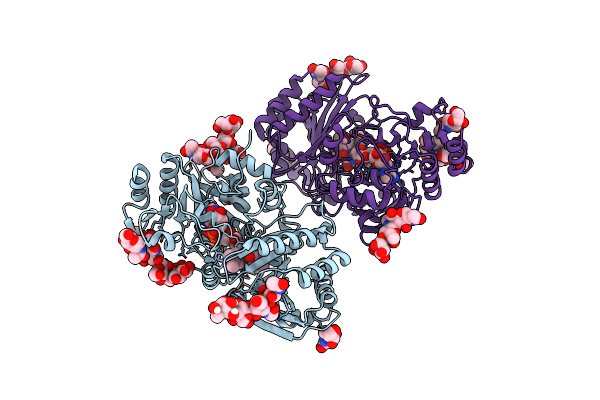 |
Organism: Penicillium rubens wisconsin 54-1255
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: FAD, NAG, A1ITD |
 |
Organism: Penicillium rubens wisconsin 54-1255
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: FAD, A1ITD |
 |
Organism: Penicillium rubens wisconsin 54-1255
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: FAD, NAG, SCN |
 |
Organism: Streptomyces sp. ag109_g2-6
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-02-12 Classification: OXIDOREDUCTASE Ligands: K, SCN, GOL |

