Search Count: 35,378
All
Selected
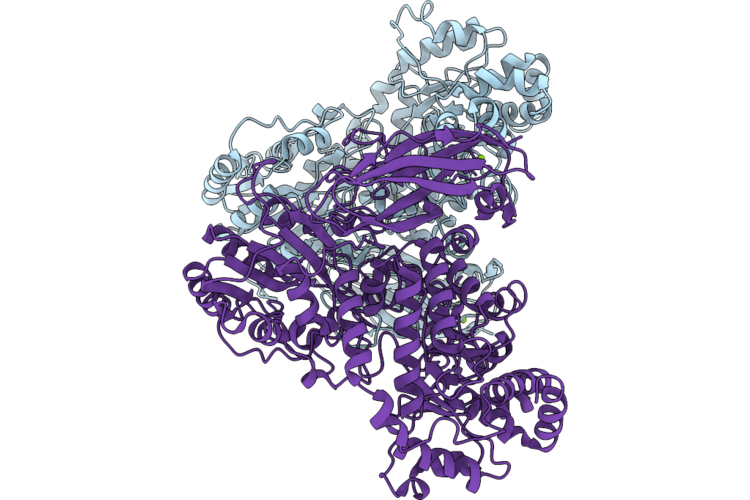 |
Organism: Bacteroides ovatus atcc 8483
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-05-20 Classification: HYDROLASE Ligands: MG |
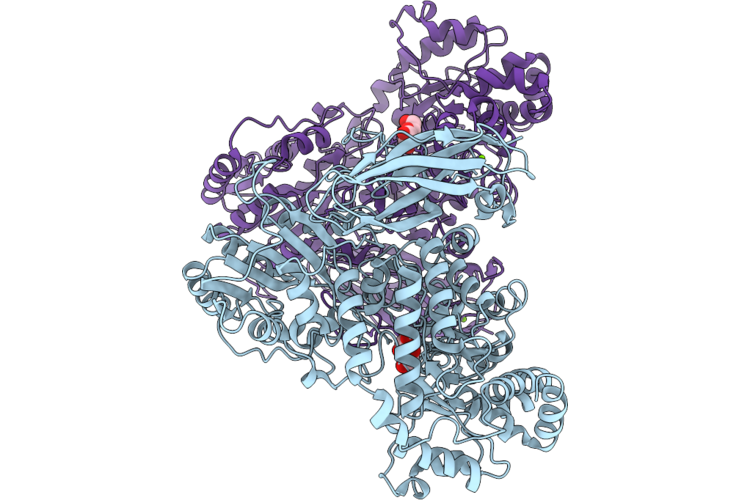 |
Crystal Structure Of Beta-Glucosidase From Bacteroides Ovatus Bound To Beta-D-Glucose
Organism: Bacteroides ovatus atcc 8483
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-05-20 Classification: HYDROLASE Ligands: BGC, MG |
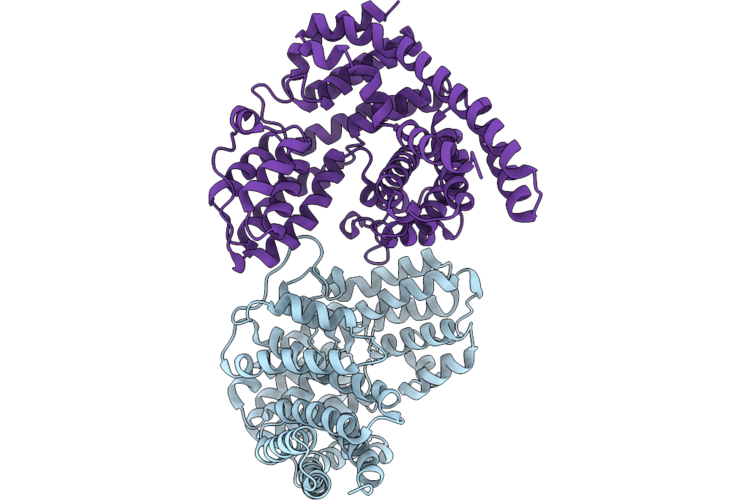 |
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-05-13 Classification: CYTOSOLIC PROTEIN |
 |
Crystal Structure Of Juvenile Hormone Acid Methyltransferase Jhamt From Choristoneura Fumiferana (Cfjhamt) In Complex With Sah (Crystal Form 1)
Organism: Choristoneura fumiferana
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-05-06 Classification: BIOSYNTHETIC PROTEIN Ligands: SAH, IOD |
 |
Crystal Structure Of Juvenile Hormone Acid Methyltransferase Cfjhamt In Complex With Sah (Crystal Form 2)
Organism: Choristoneura fumiferana
Method: X-RAY DIFFRACTION Resolution:2.84 Å Release Date: 2026-05-06 Classification: BIOSYNTHETIC PROTEIN Ligands: SAH, CA, CL |
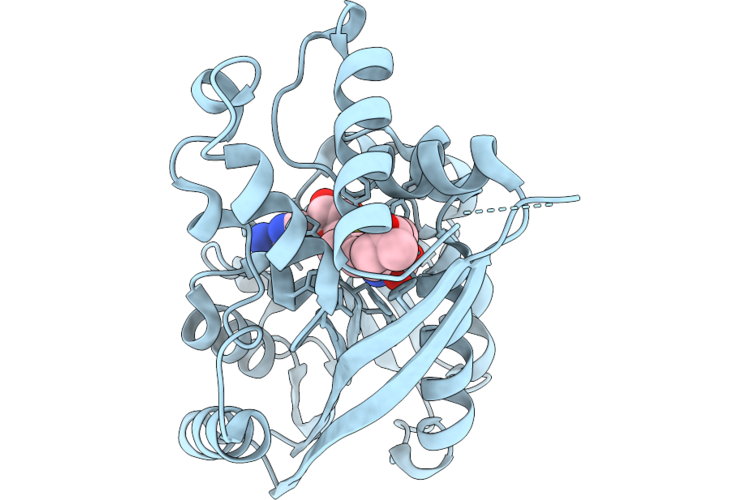 |
Crystal Structure Of Juvenile Hormone Acid Methyltransferase Cfjhamt In Complex With Sah And Juvenile Hormone Iii Acid
Organism: Choristoneura fumiferana
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-05-06 Classification: BIOSYNTHETIC PROTEIN Ligands: SAH, JH3 |
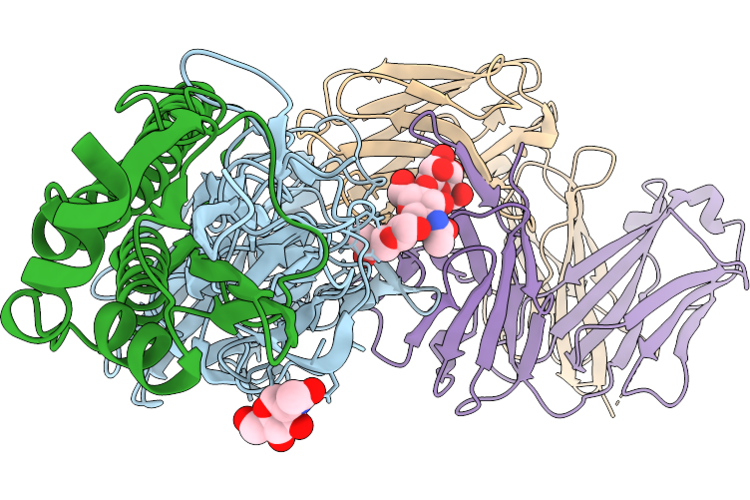 |
Crystal Structure Of Hemagglutinin From H1N1 Influenza A Virus A/California/04/2009 Bound To The 3_H2 Antibody
Organism: Influenza a virus (a/california/04/2009(h1n1)), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.39 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: NAG |
 |
Crystal Structure Of Hemagglutinin From H1N1 Influenza A Virus A/California/04/2009 Bound To The 49_C09 Antibody
Organism: Influenza a virus (a/california/04/2009(h1n1)), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.80 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: NAG |
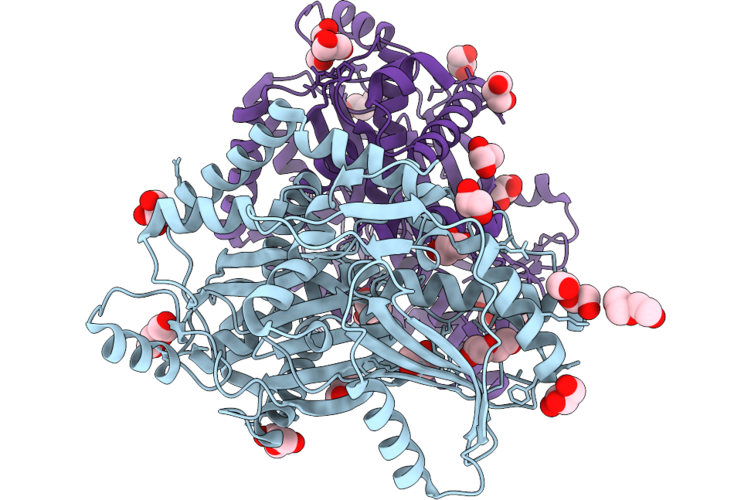 |
Organism: Coprococcus eutactus atcc 27759
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-29 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL, PEG |
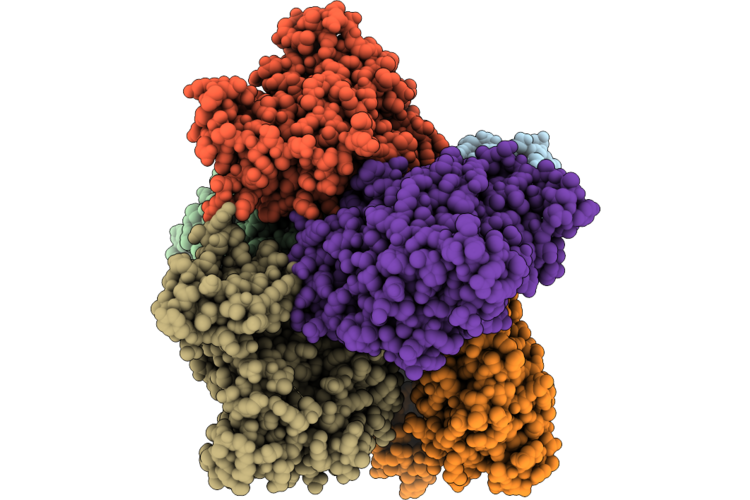 |
Crystal Structure Of The Catalytic Subunit Of The Circadian Regulator Casein Kinase 2 From Neurospora Crassa
Organism: Neurospora crassa
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-29 Classification: TRANSFERASE |
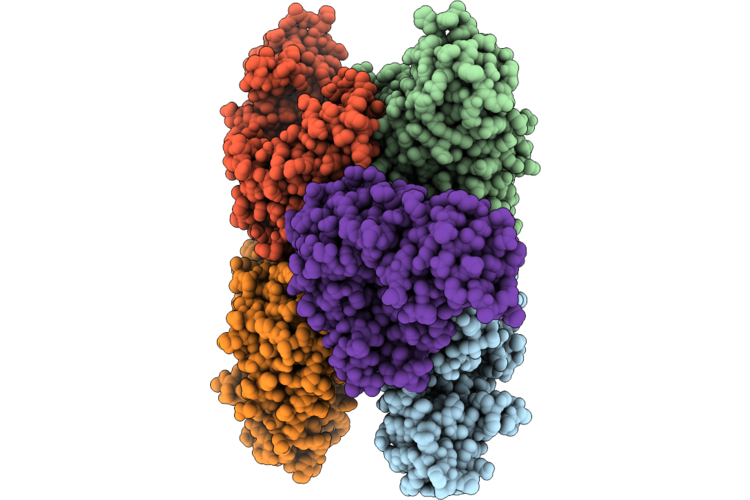 |
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-04-29 Classification: TRANSFERASE |
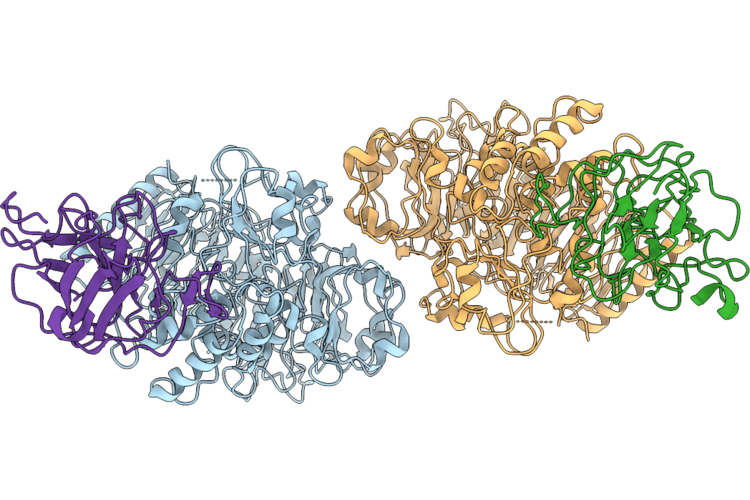 |
Cryo-Em Structure Of Plant Resistance Protein Nrc2 Dimer Bound To Nematode Effector Sprysec-15
Organism: Nicotiana benthamiana, Globodera rostochiensis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: PLANT PROTEIN |
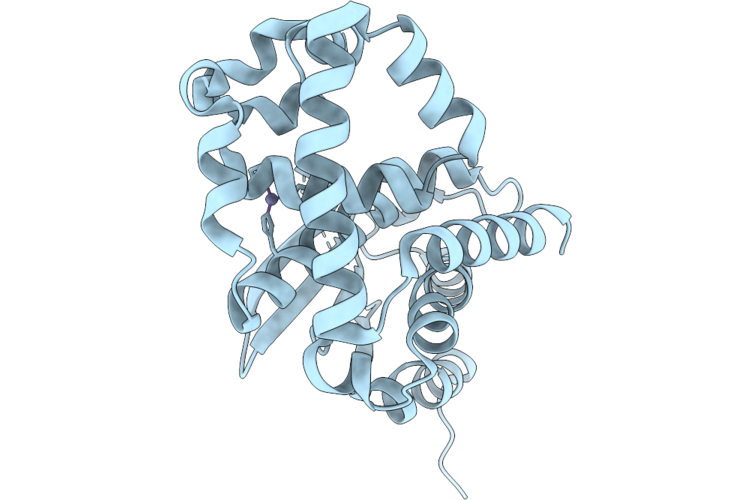 |
Crystal Structure Of Zinc-Binding Metallopeptidase (Uniprot: A0Aan3A8D2) From Bacteroides Ovatus
Organism: Bacteroides ovatus atcc 8483
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ZN |
 |
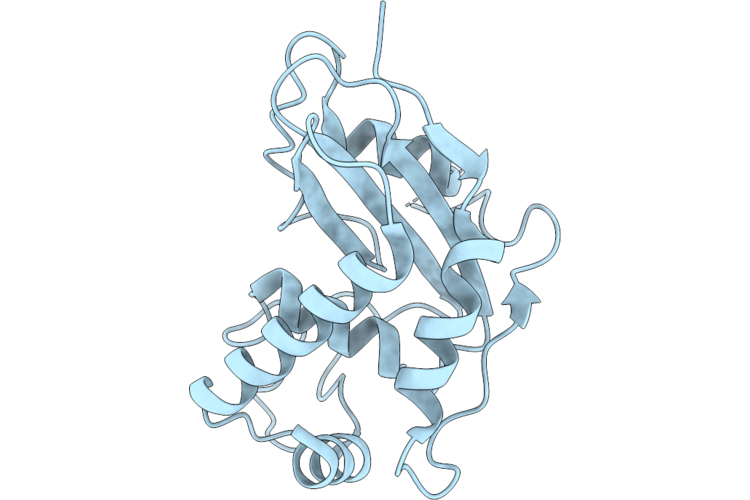
Crystal Structure Of Halide Methyltransferase From Dichomitus Squalens In Complex With Sah
Organism: Dichomitus squalens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: SAH, ACT |
 |
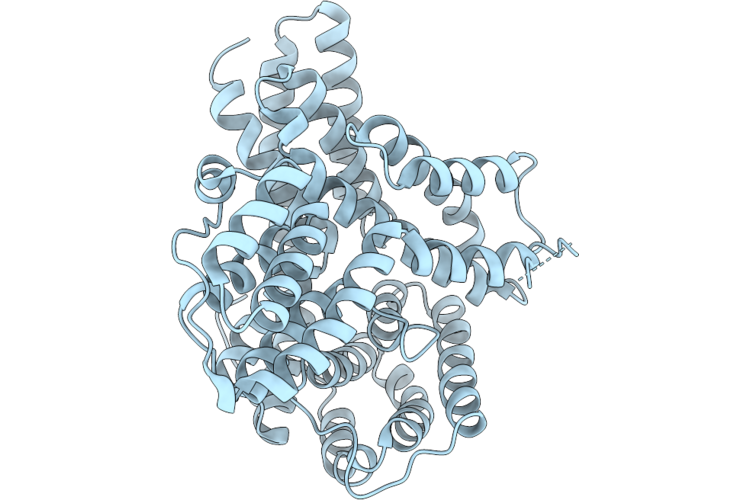
Organism: Eimeria tenella strain houghton
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-04-22 Classification: UNKNOWN FUNCTION |
 |
Central Domain Of Glucan Water Dikinase-1 In Alternative Closed Conformation
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:3.25 Å Release Date: 2026-04-22 Classification: CYTOSOLIC PROTEIN |
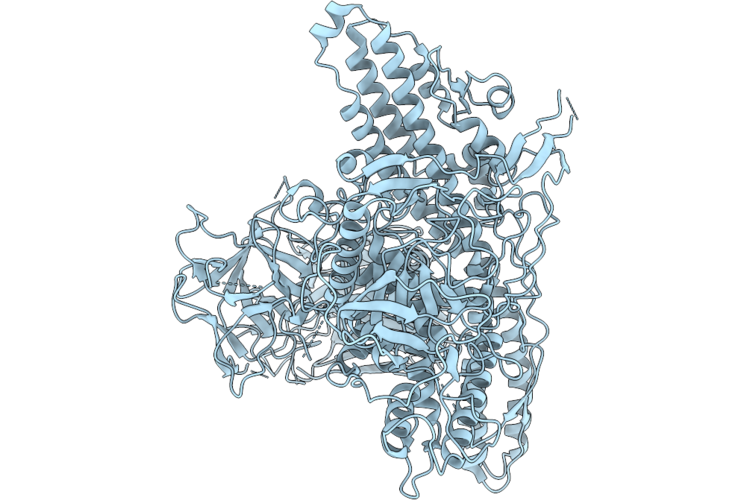 |
Cryo-Em Structure Of Semi-Closed State Of Botulinum Neurotoxin Serotype A At Ph 7.4
Organism: Clostridium botulinum (strain hall / atcc 3502 / nctc 13319 / type a)
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-22 Classification: TOXIN |
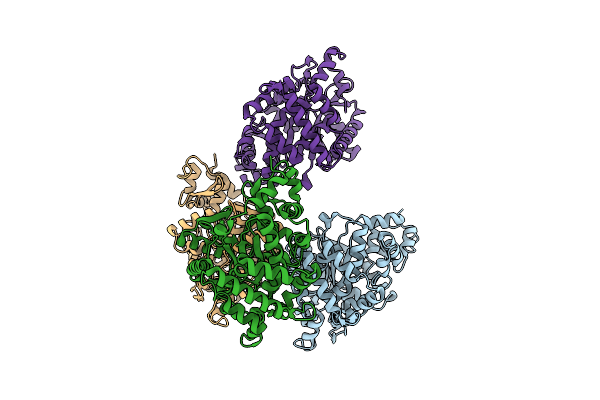 |
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2026-04-15 Classification: PLANT PROTEIN |
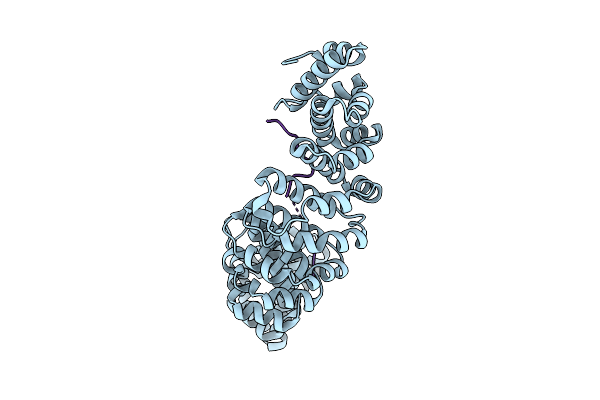 |
Organism: Mus musculus, Black sea bass polyomavirus 1
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-01 Classification: PROTEIN TRANSPORT |
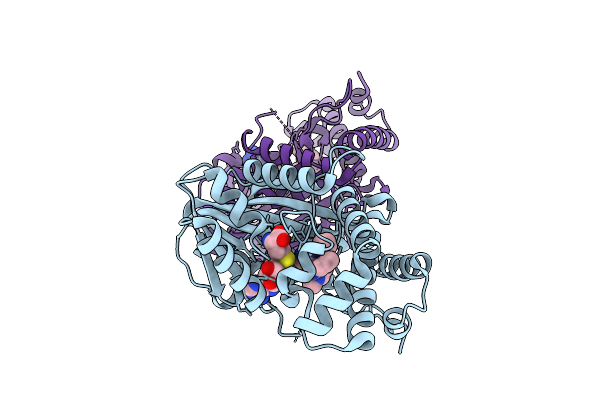 |
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: A1EWC, SAH |

