Search Count: 4,545
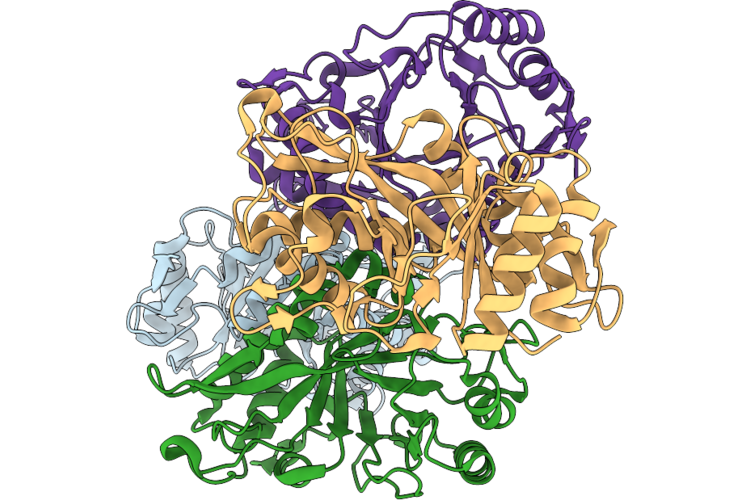 |
Organism: Streptomyces plicatus
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-05-13 Classification: CARBOHYDRATE Ligands: CL |
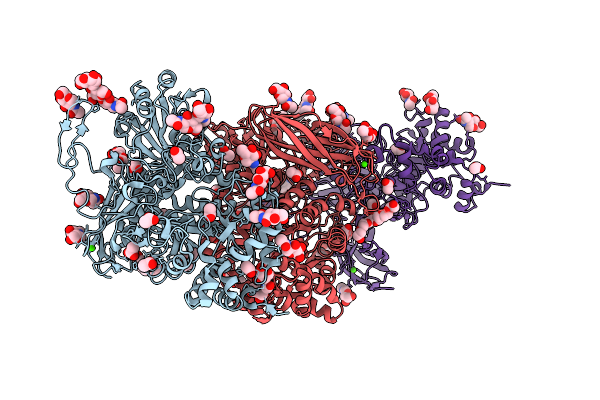 |
Crystal Structure Of Isoprimeverose-Producing Enzyme From Phaeoacremonium Minimum
Organism: Rattus rattus, Phaeoacremonium minimum
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAG, ACT, PG4, CA, PGE, P6G |
 |
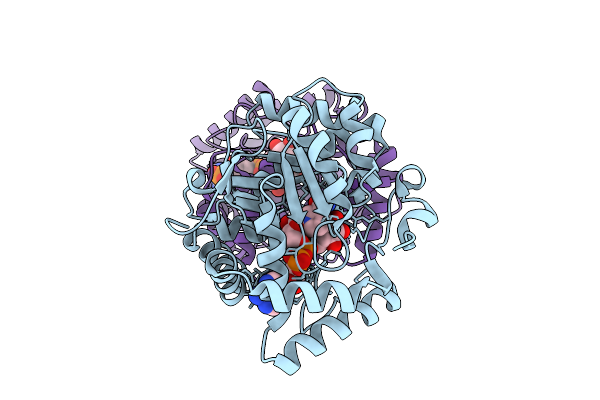
Crystal Structure Of L-Galactose Dehydrogenase From Luteolibacter Sp. Strain Lg18 In Complex With L-Galactose And Nadp+
Organism: Luteolibacter sp. lg18
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: GIV, NAP |
 |
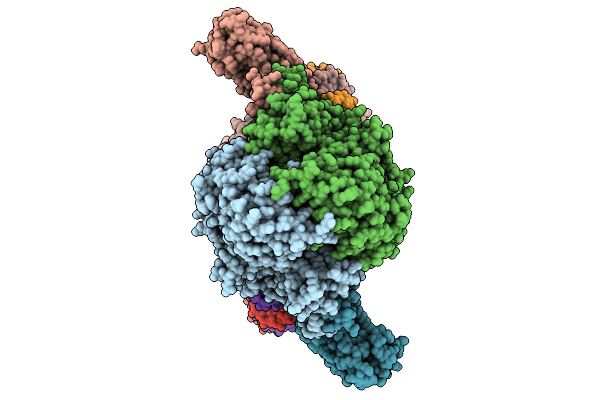
Crystal Structure Of L-Galactose Dehydrogenase From Luteolibacter Sp. Strain Lg18 In Complex With L-Glucose And Nadp+
Organism: Luteolibacter sp. lg18
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: Z8T, NAP |
 |
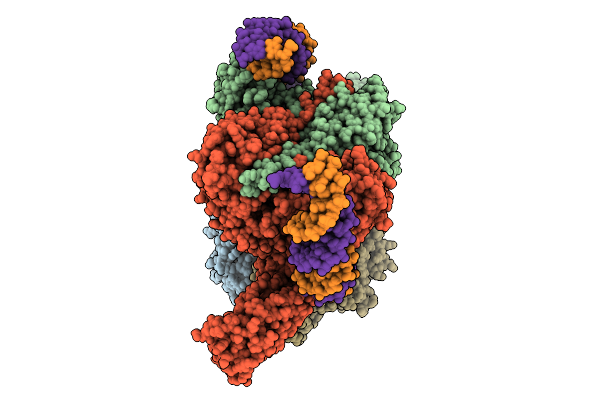
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Cleavage State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: CA, ANP, K |
 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, CA, K |
 |
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
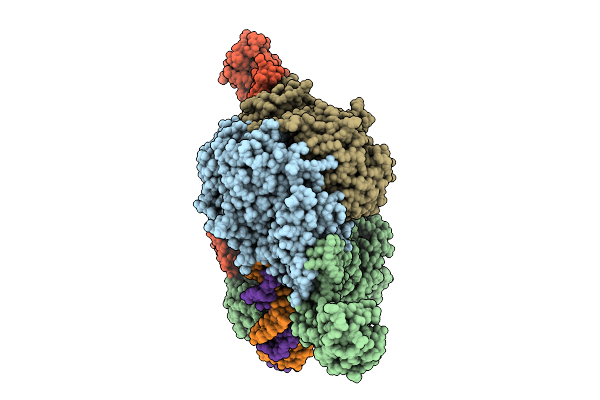
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Partially Unfolded Transducer State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: MG, ANP, K |
 |
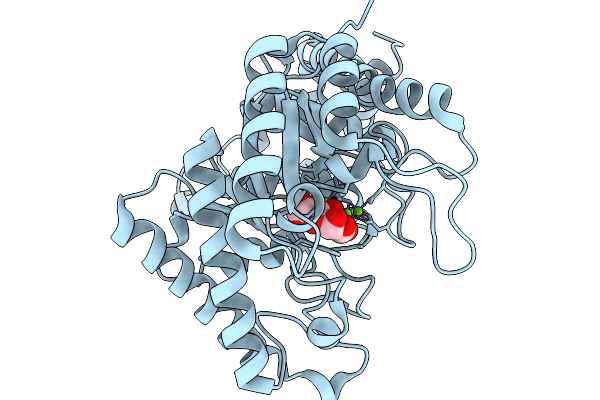
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
Xylose Isomerase Collected At 20C Using Time-Resolved Serial Synchrotron Crystallography With Glucose At 180 Seconds
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-18 Classification: ISOMERASE Ligands: GLC, GLO, MG |
 |
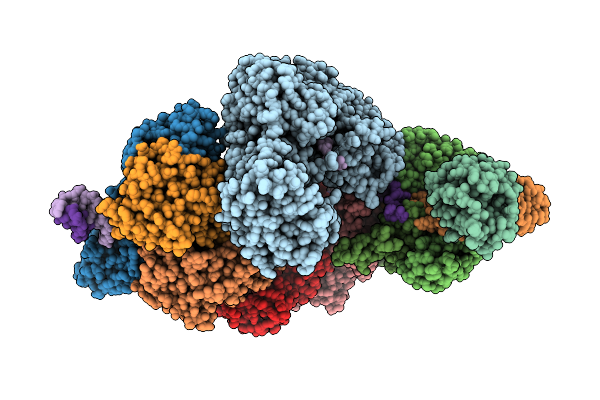
Structure Of Murj In Complex With Single Gene Lysis Protein From Phage Changjiang3
Organism: Escherichia coli k-12, Changjiang levi-like virus 3
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN |
 |
Organism: Shewanella putrefaciens cn-32
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: ANTIVIRAL PROTEIN Ligands: MG |
 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1JAP, L1P, A1JAO, NA, A1JAN |
 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: B13, A1JAP, L1P, A1JAO, NA, A1JAN |
 |
Spitrobot-2 Advances Time-Resolvedcryo-Trapping Crystallography To Under 25 Ms: Xylose Isomerase (Apo State)
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2025-12-03 Classification: ISOMERASE Ligands: MN |
 |
Spitrobot-2 Advances Time-Resolvedcryo-Trapping Crystallography To Under 25 Ms: Xylose Isomerase Bound With Glucose (25 Ms Soaking)
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-12-03 Classification: ISOMERASE Ligands: MG, MN, BGC |
 |
Spitrobot-2 Advances Time-Resolvedcryo-Trapping Crystallography To Under 25 Ms: Xylose Isomerase Bound With Glucose (50 Ms Soaking)
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-03 Classification: ISOMERASE Ligands: MG, MN, BGC |
 |
Organism: Salpingoeca macrocollata
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2025-10-01 Classification: IMMUNE SYSTEM Ligands: 1SY |
 |
Organism: Marinactinospora thermotolerans dsm 45154
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2025-09-03 Classification: HYDROLASE |
 |
Organism: Fasciola hepatica
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-08-27 Classification: CELL ADHESION |

