Search Count: 2,123
All
Selected
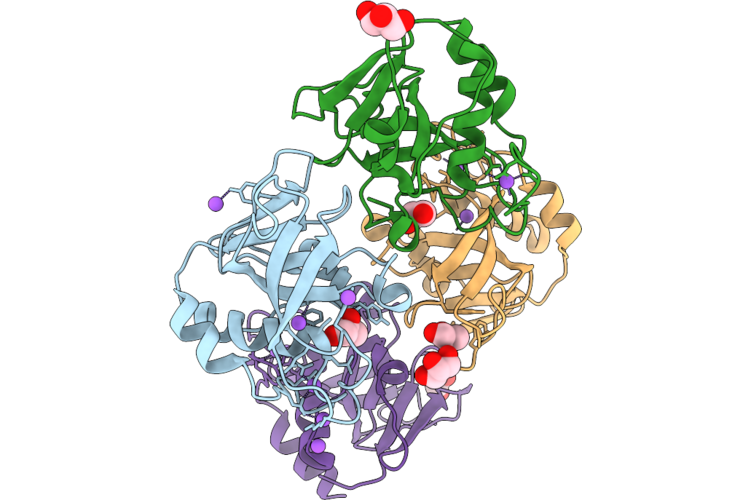 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2026-05-06 Classification: HYDROLASE |
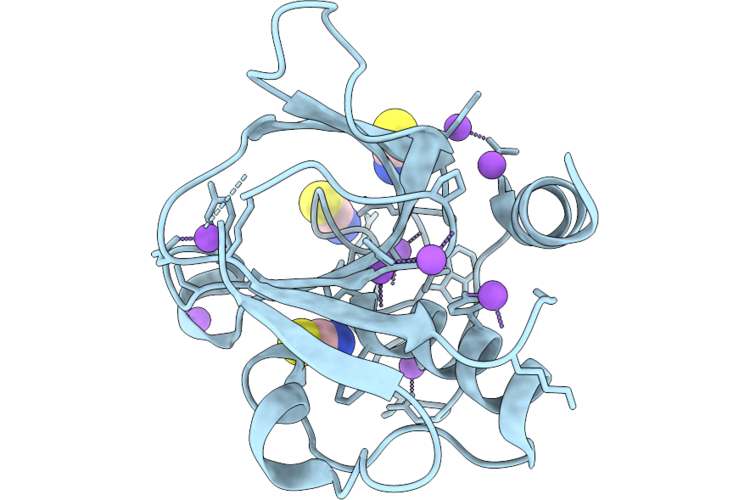 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-05-06 Classification: HYDROLASE |
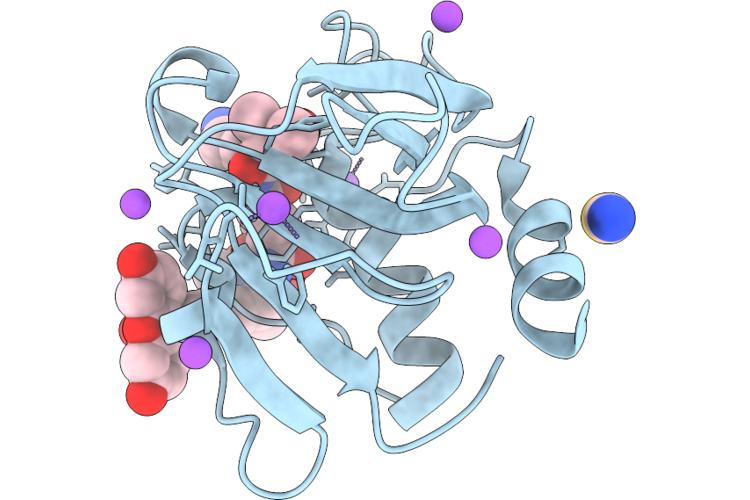 |
Chap Domain Of Staphylococcus Aureus-Specific Lysin L1 Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA, SCN |
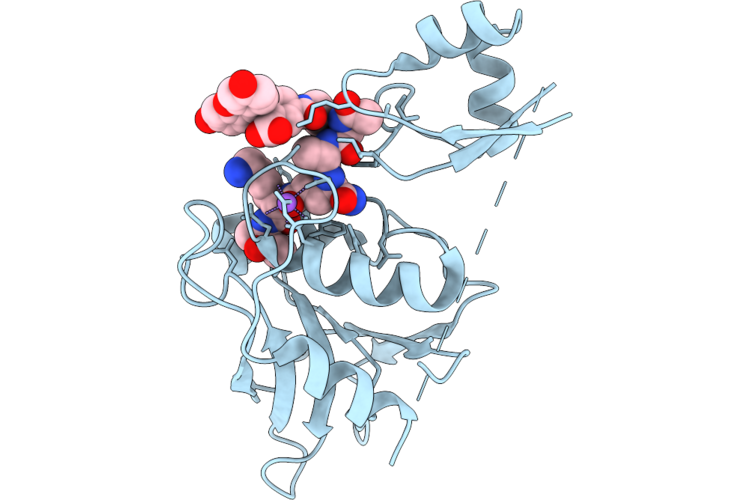 |
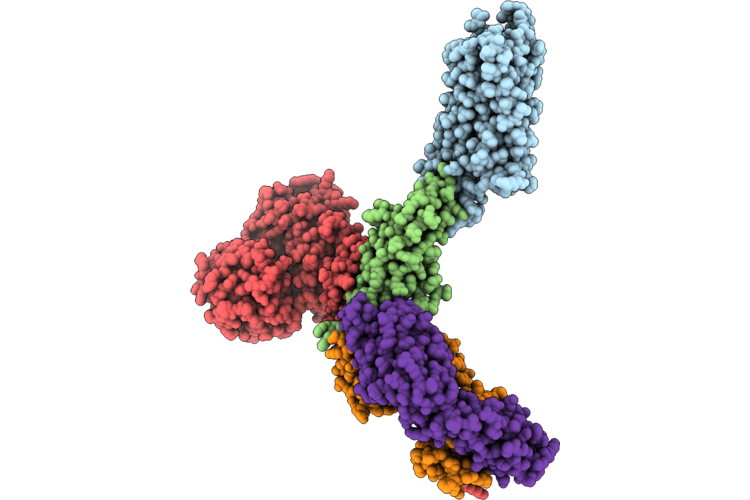
Staphylococcus Aureus-Specific Lysin L1-3 (Lysm-Chap) Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA |
 |
Organism: Escherichia coli o157:h7, Staphylococcus aureus, Staphylococcus aureus subsp. aureus mu50, Streptococcus sp. 'group g', Homo sapiens, Spodoptera frugiperda
Method: ELECTRON MICROSCOPY Resolution:3.08 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN |
 |
Organism: Staphylococcus aureus, Lama glama
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-05-06 Classification: IMMUNE SYSTEM Ligands: ZN |
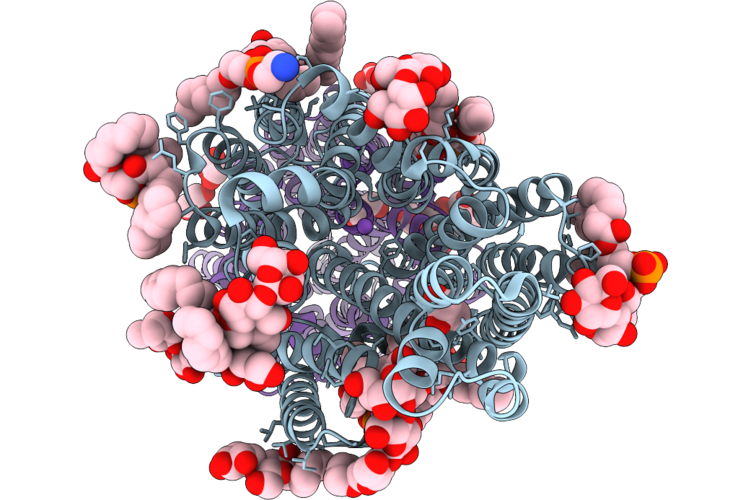 |
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PGT, LMU, PTY, 3PH, GOL, NA |
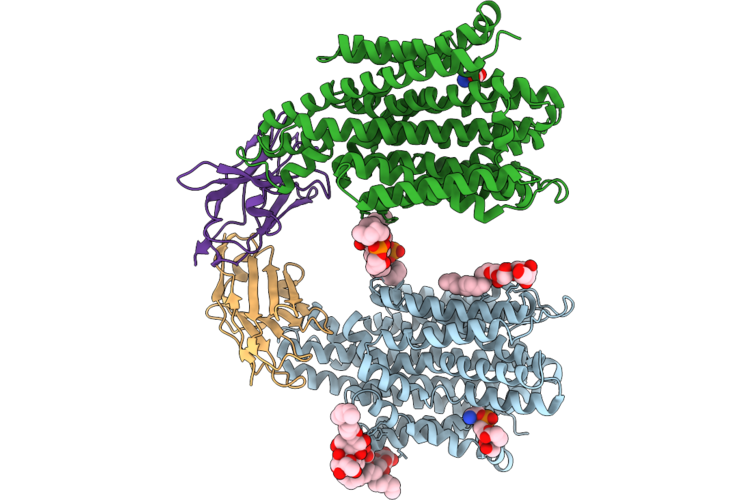 |
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Inward Open State In Complex With Nanobody 89
Organism: Staphylococcus aureus, Lama glama
Method: X-RAY DIFFRACTION Resolution:3.32 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PTY, PGT, LMU, OXM |
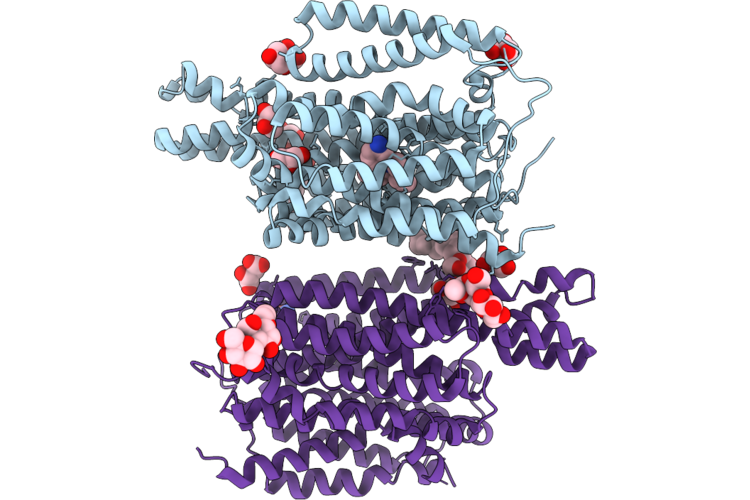 |
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State Bound To Ethidium
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ET, GOL, TLA, CL, LMU, OXM |
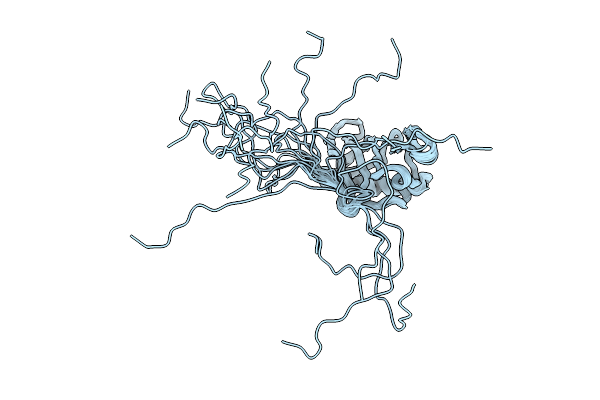 |
Organism: Staphylococcus aureus
Method: SOLUTION NMR Release Date: 2026-04-15 Classification: PROTEIN BINDING |
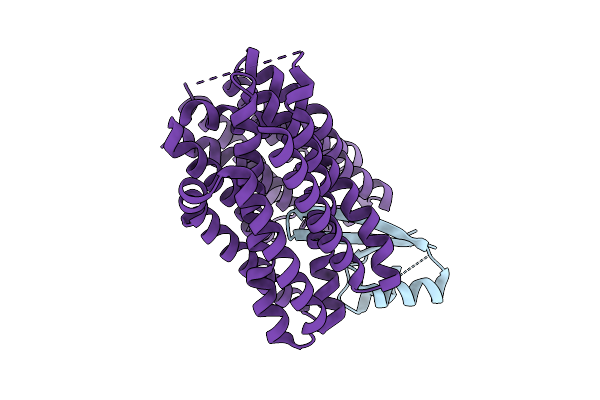 |
Organism: Synthetic construct, Staphylococcus aureus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN |
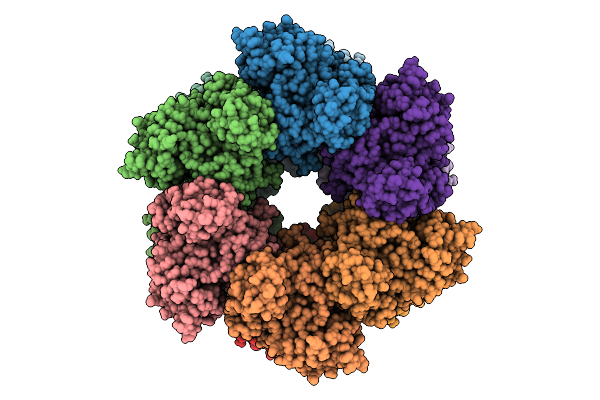 |
Organism: Staphylococcus aureus
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN |
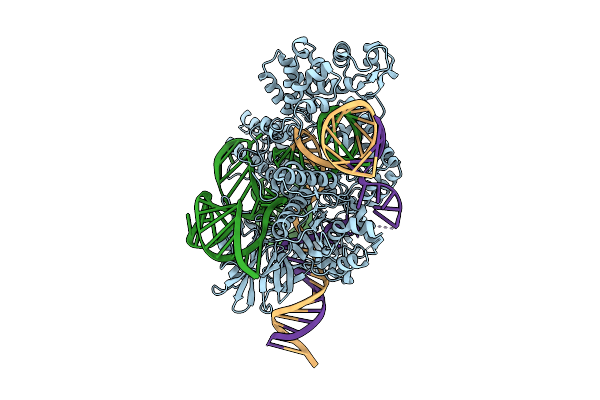 |
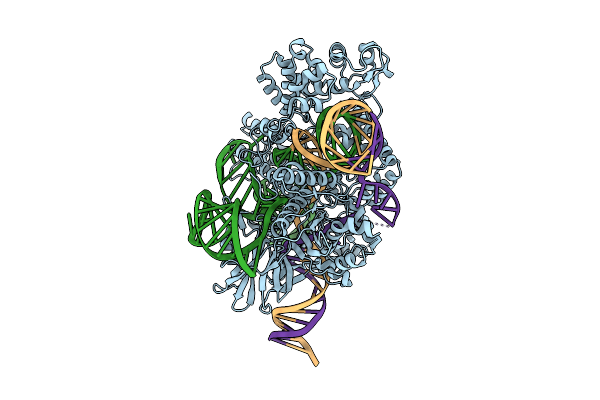
Cryo-Em Structure Of Wild-Type Sacas9-Guide Rna-Mismatched Target Dna Complex
Organism: Staphylococcus aureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN |
 |
Organism: Staphylococcus aureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN Ligands: MG |
 |
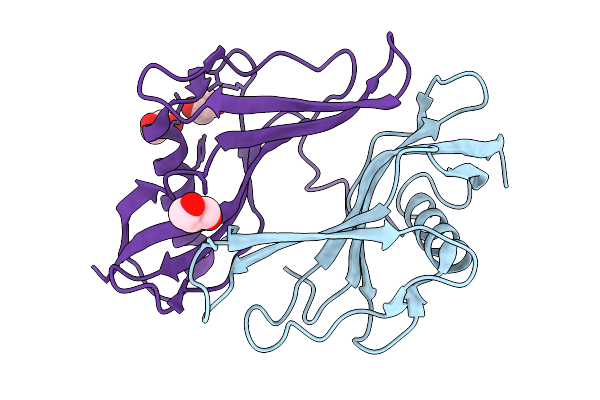
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: PGE, FLC |
 |
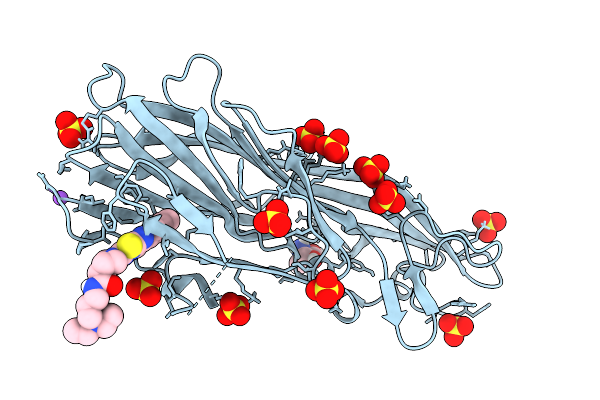
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: GOL, EDO |
 |
Alpha-Hemolysin Of S.Aureus (Monomer) In Complex With The Small Molecule N-(3-(Diethylamino)Propyl)-1-(6-(P-Tolyl)Imidazo[2,1-B][1,3,4]Thiadiazol-2-Yl)Piperidine-4-Carboxamide
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-04 Classification: TOXIN Ligands: A1JN0, TRS, SO4, NA |
 |
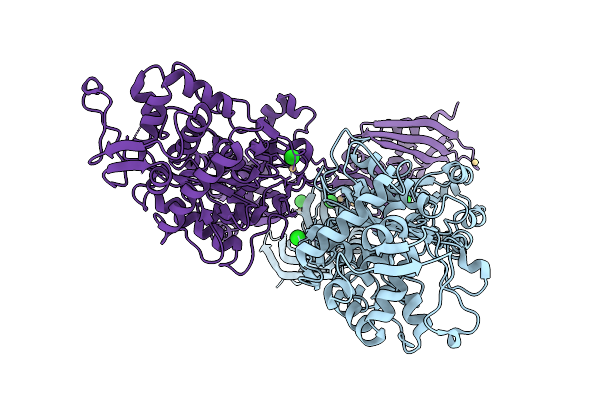
Organism: Staphylococcus aureus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: RIBOSOME |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |

