Search Count: 4,214
All
Selected
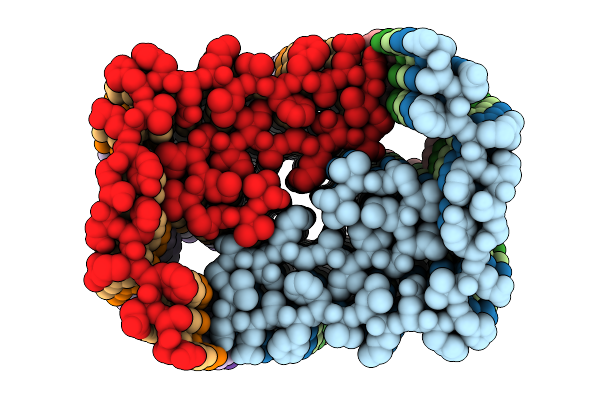 |
Population B Fibril Generated From The Heterotypic Interaction Of Abeta40 And Medin.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: PROTEIN FIBRIL |
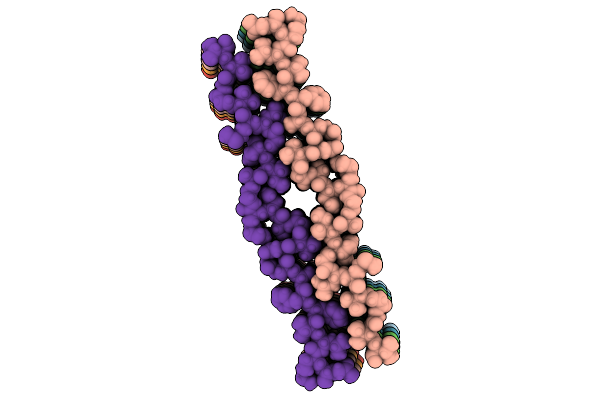 |
Population A Fibril Generated From The Heterotypic Interaction Of Abeta40 And Medin.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-04-08 Classification: PROTEIN FIBRIL |
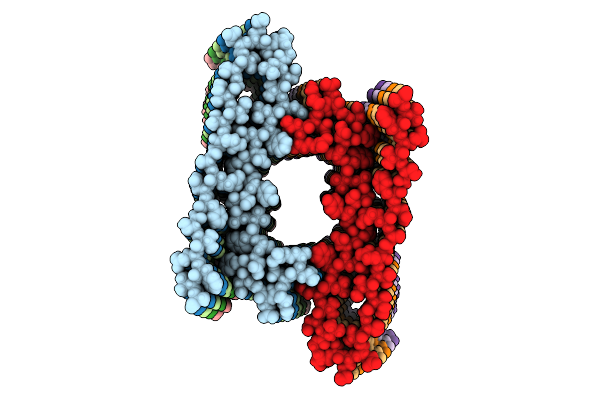 |
Medin Fibril Generated From The Heterotypic Interaction Of Abeta40 And Medin.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-04-08 Classification: PROTEIN FIBRIL |
 |
Crystal Structure Of The Inactive Mutant Of Rice Protein Disulfide Isomerase-Like Protein Ospdil2-3 A-B Domains
Organism: Oryza sativa subsp. japonica
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: BTB |
 |
Crystal Structure Of Rice Protein Disulfide Isomerase-Like Protein Ospdil2-3 A-B Domain
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: BTB |
 |
Crystal Structure Of Rice Protein Disulfide Isomerase-Like Protein Ospdil2-3 A0 Domain
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
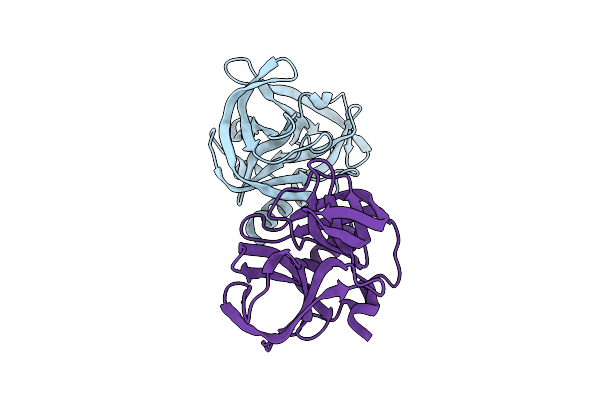 |
Crystal Structure Of Enterovirus D68 3C Protease Determined Via Sulfur Phasing
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-01 Classification: VIRAL PROTEIN Ligands: CL |
 |
Crystal Structure Of Zika Virus Ns2B-Ns3 Protease Determined Via Sulfur Phasing
Organism: Zika virus
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-04-01 Classification: VIRAL PROTEIN |
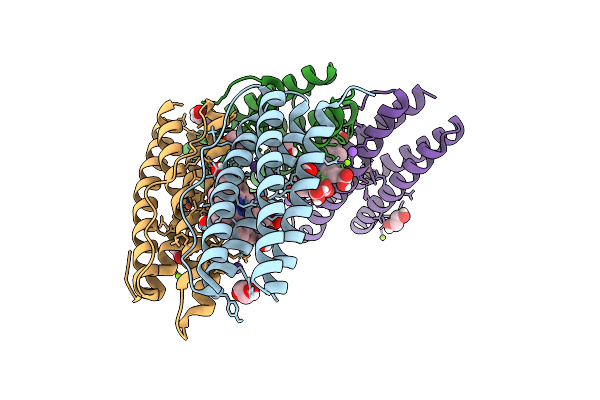 |
Mini-Bacterioferritin From Candidatus Methanoperedens Species Blz2 As Isolated Form At 1.07-A Resolution
Organism: Candidatus methanoperedens sp. blz2
Method: X-RAY DIFFRACTION Resolution:1.07 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: EDO, FE, FEC, MG, NA, GOL |
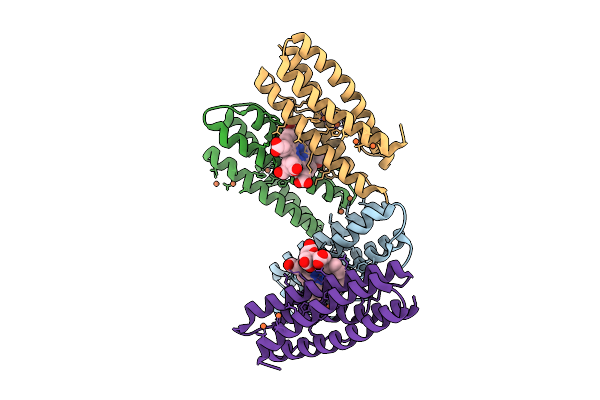 |
Mini-Bacterioferritin From Candidatus Methanoperedens Species Blz2 In A Partially Oxidized State
Organism: Candidatus methanoperedens sp. blz2
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FE, FEC |
 |
Mini-Bacterioferritin From Candidatus Methanoperedens Species Blz2 Oxidized Then Chemically Reduced
Organism: Candidatus methanoperedens sp. blz2
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FE, FEC |
 |
Mini-Bacterioferritin From Candidatus Methanoperedens Species Blz2 Untreated Control Of A Redox Cycling Experiment
Organism: Candidatus methanoperedens sp. blz2
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FEC, FE |
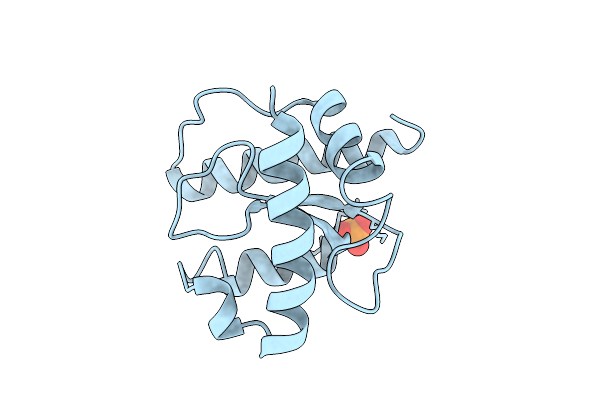 |
X-Ray Structure Of The C-Terminal Domain (Residues 366-485) Of S. Pombe Threonylcarbamoyladenosine Dehydratase At 1.55 A Resolution
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN Ligands: PO4 |
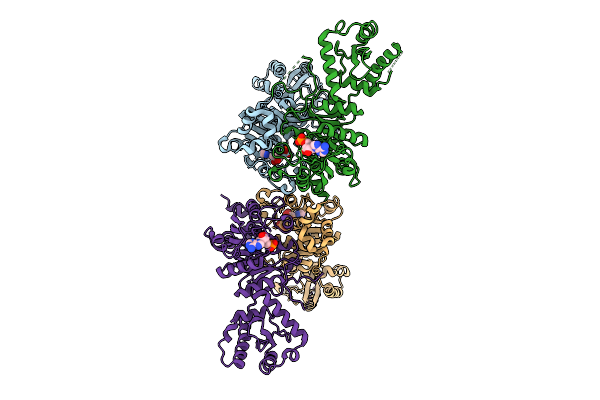 |
X-Ray Structure Of S. Cerevisiae Threonlycarbamoyladenosine Dehydratase 1 (Residues 50-429) In Complex With Amp At 3.7 A Resolution
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:3.70 Å Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN Ligands: AMP, MG |
 |
Crystal Structure Of Coxsackievirus A16 (G-10) 2A Protease Determined Via Sulfur Phasing
Organism: Coxsackievirus a16
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: GOL, ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-03-18 Classification: TRANSPORT PROTEIN Ligands: GOL |
 |
Organism: Prymnesium parvum
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-11 Classification: LIGASE |
 |
Investigating The Binding Mechanism Of Interferon Regulatory Factor 4 To Dna In The Context Of Multiple Myeloma
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN Ligands: CA |
 |
Organism: Chromobacterium sphagni
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: ZN |
 |
Biocatalytic Regioselective C-Formylation Of Resorcinol Derivatives (Csatase C88S)
Organism: Chromobacterium sphagni
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: GOL, ACT, ZN |

