Search Count: 6,124
All
Selected
 |
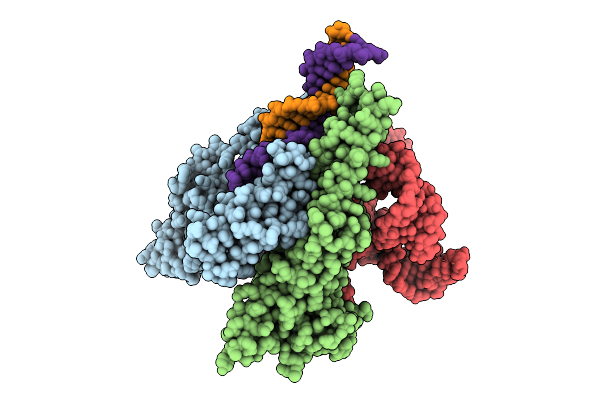
Organism: Homo sapiens, Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
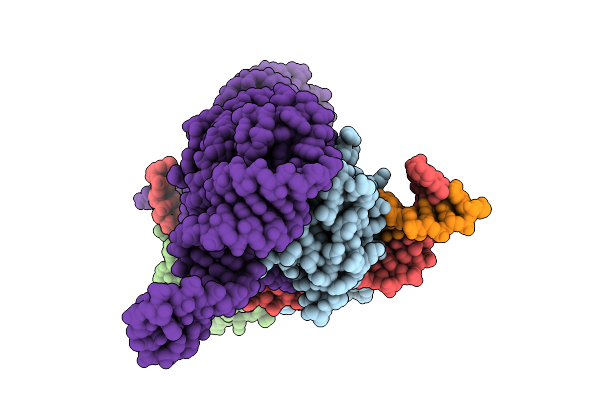
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-04-08 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Ruminiclostridium herbifermentans
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-04-08 Classification: RNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Oscillibacter sp.
Method: ELECTRON MICROSCOPY Resolution:3.67 Å Release Date: 2026-04-08 Classification: RNA BINDING PROTEIN/DNA/RNA Ligands: ZN |
 |
Organism: Oscillibacter sp.
Method: ELECTRON MICROSCOPY Resolution:3.68 Å Release Date: 2026-04-08 Classification: RNA BINDING PROTEIN/DNA/RNA Ligands: ZN |
 |
Organism: Oscillibacter sp.
Method: ELECTRON MICROSCOPY Resolution:3.27 Å Release Date: 2026-04-08 Classification: RNA BINDING PROTEIN/DNA/RNA Ligands: ZN |
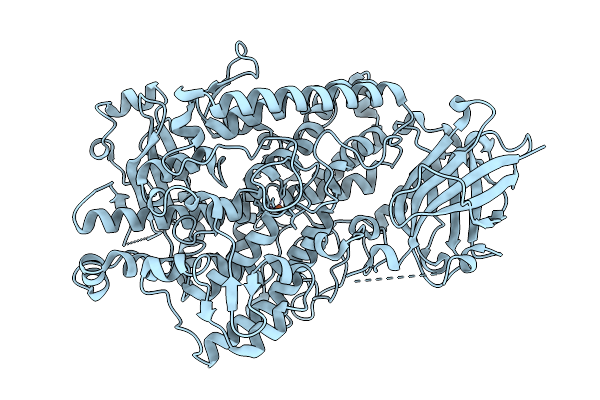 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE |
 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE |
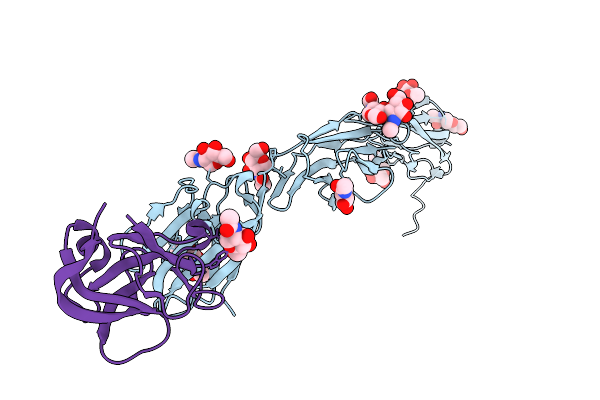 |
Cardioderma Bat Coronavirus Ky43 Receptor Binding Domain In Complex With Human Ceacam6
Organism: Homo sapiens, Cardioderma bat coronavirus
Method: X-RAY DIFFRACTION Resolution:3.01 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN Ligands: NAG |
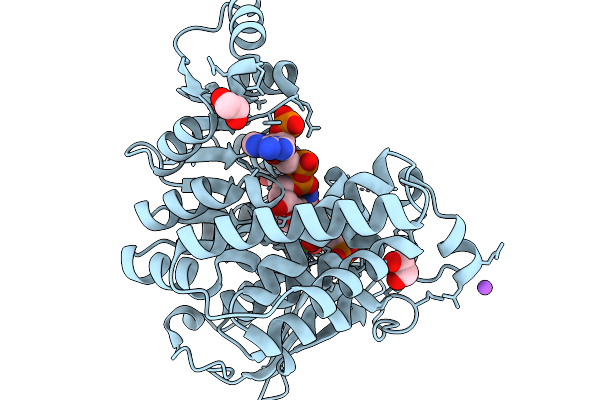 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: EDO, NDP, A1CFA, MG, CL, NA |
 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: NDP, A1CFB, EDO, MG, CL |
 |
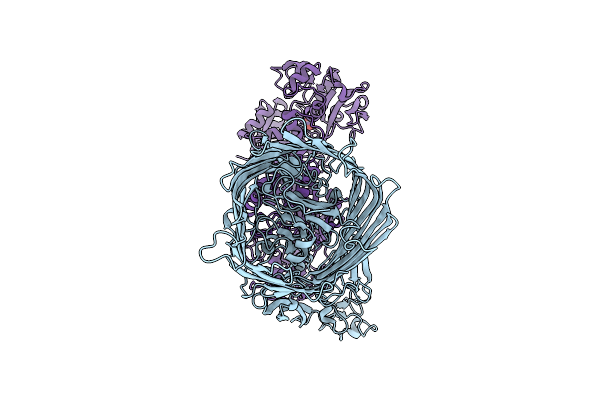
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: NDP, A1CE6, EDO, MG |
 |
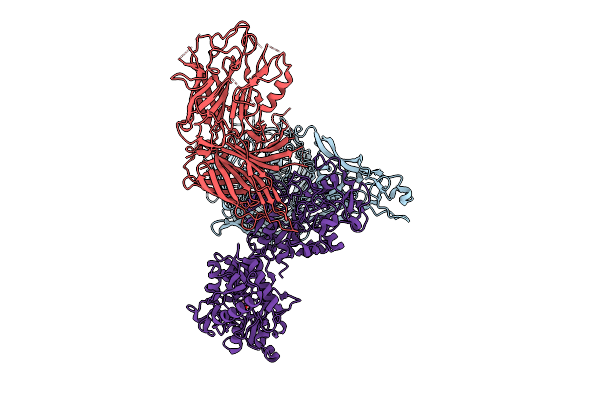
Transferrin Binding Protein A In Complex With Transferrin (Iron Bound In N Lobe Only)
Organism: Neisseria meningitidis serogroup b, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: TRANSPORT PROTEIN Ligands: BCT, FE |
 |
Transferrin Binding Protein A In Complex With Transferrin Binding Protein B And Transferrin (Iron Bound To N Lobe Only)
Organism: Neisseria meningitidis, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: BCT, FE |
 |
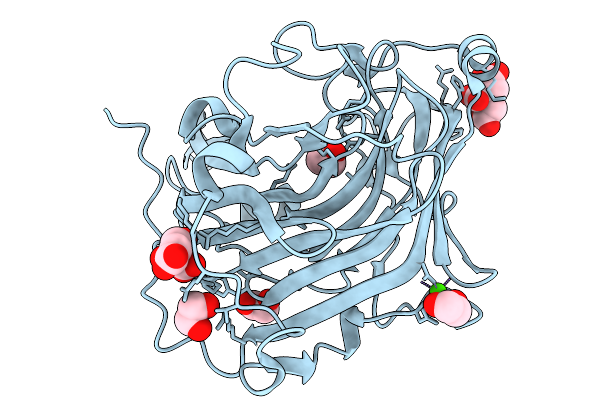
Crystal Structure Of Kirsten Rat Sarcoma G12C Complexed With Gmppnp And Covalently Bound To 1-[(2R,3R)-3-{[(7P)-7-(8-Ethynyl-7-Fluoronaphthalen-1-Yl)-8-Fluoro-2-{ [(2R,4R,7As)-2-Fluorotetrahydro-1H-Pyrrolizin-7A(5H)-Yl]Methoxy}Pyrido[4,3-D] Pyrimidin-4-Yl](Methyl)Amino}-2-Methylpyrrolidin-1-Yl]-3-(Pyrazin-2-Yl)Propan-1-One
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: GDP, A1C5K, CA |
 |
Organism: Bacterium
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: CIT, EDO, CA |
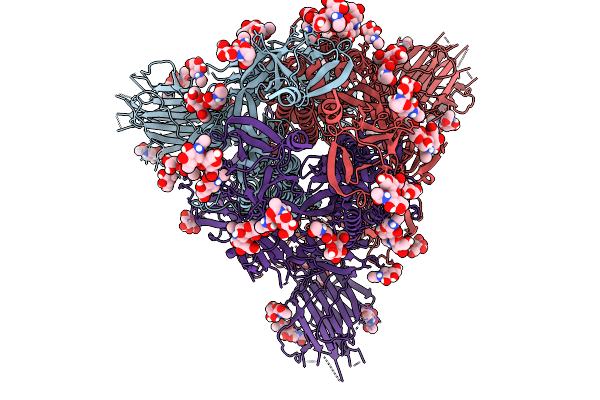 |
Cryo-Em Consensus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 1 Up & 2 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
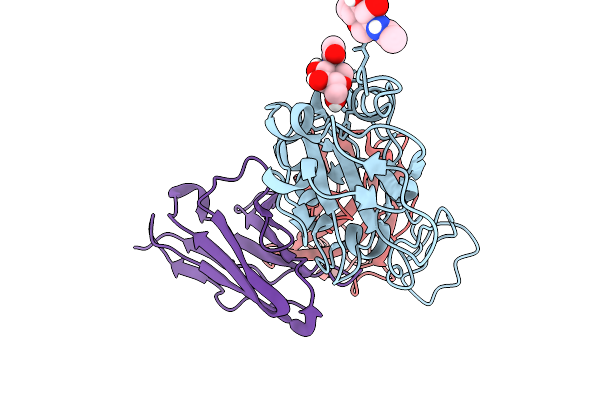 |
Cryo-Em Focus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 1 Up & 2 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
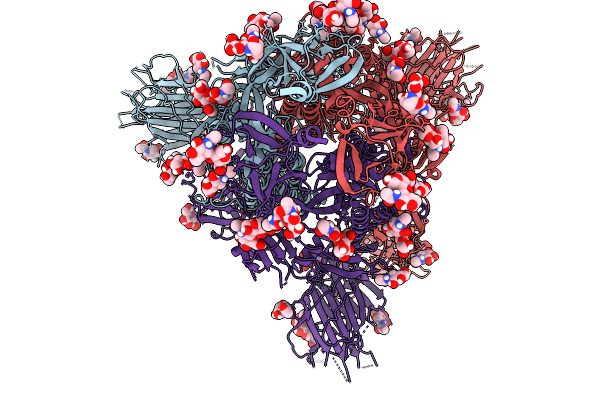 |
Cryo-Em Consensus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 2 Up & 1 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
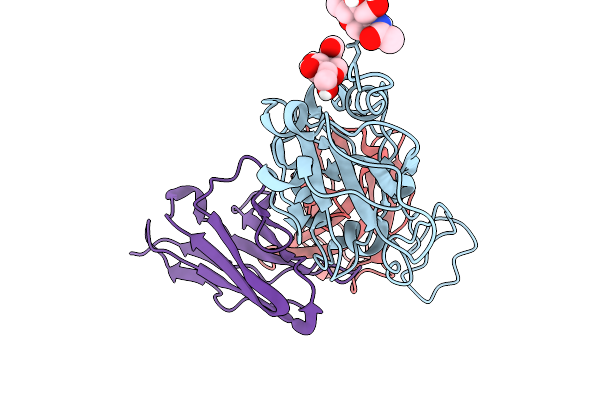 |
Cryo-Em Focus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 2 Up & 1 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |

