Search Count: 10,606
All
Selected
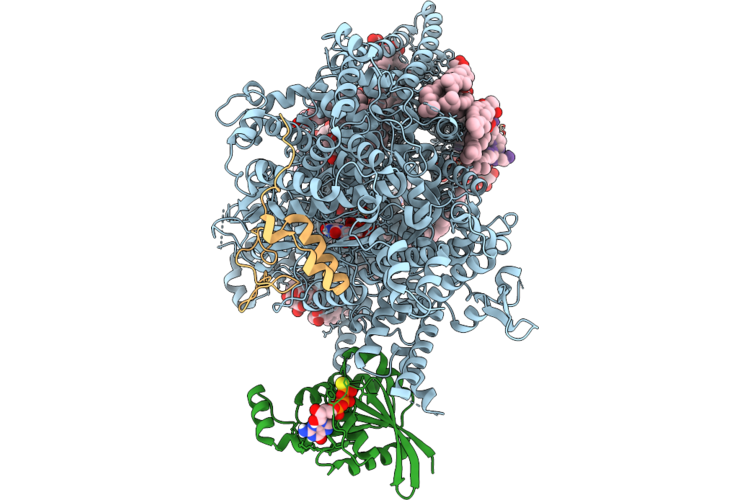 |
Structure Of Beta-1,3-Glucan Synthase In Complex With Caspofungin, Rho1 And Long Glucan
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP, GSP, A1CHR |
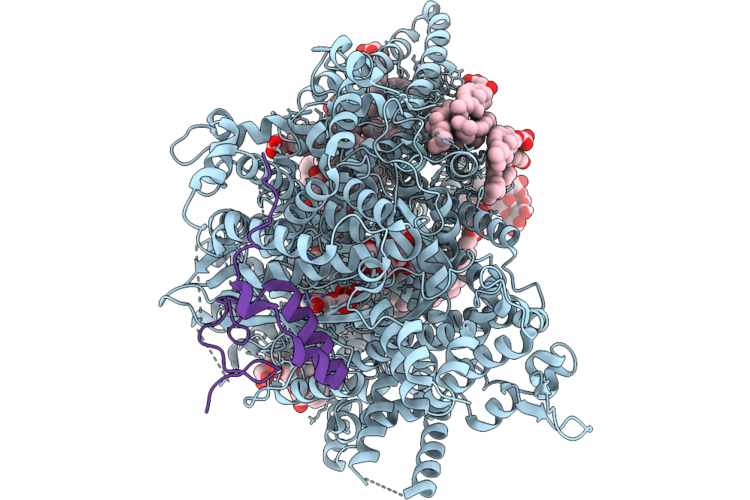 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) In Complex With Short Glucan
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
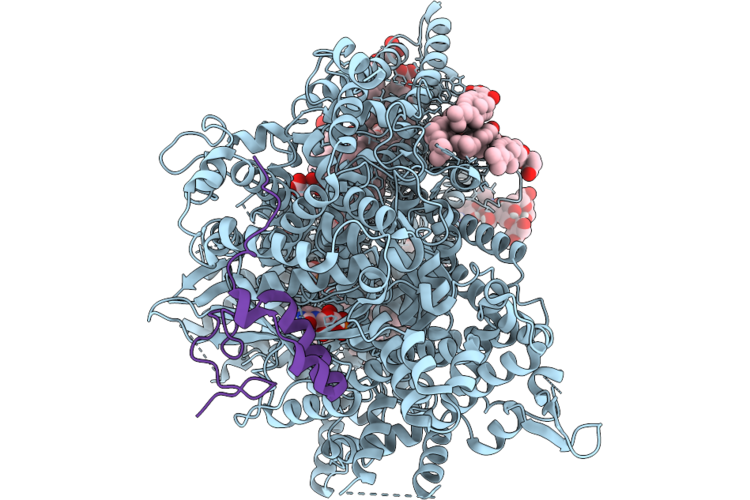 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Relevant Ground State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
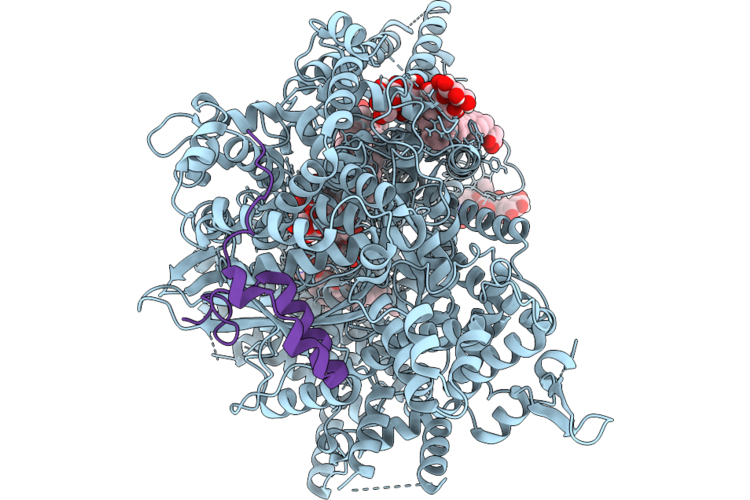 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Less Relevant L2 State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01, 3PE, LMN |
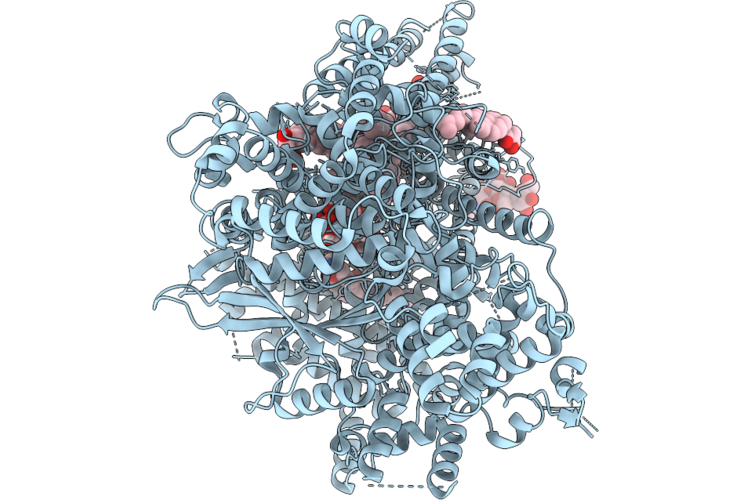 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Less Relevant L1 State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01, 3PE, LMN |
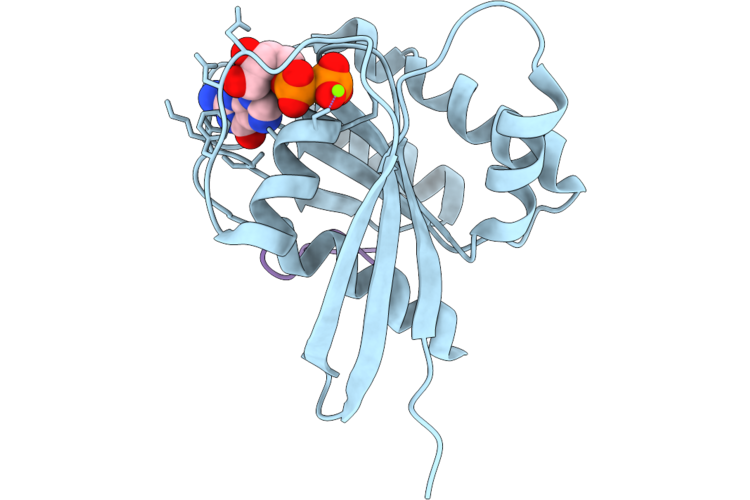 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
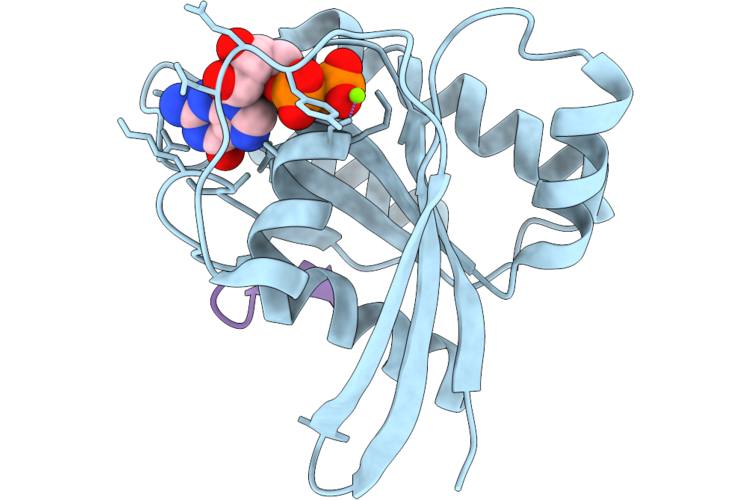 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
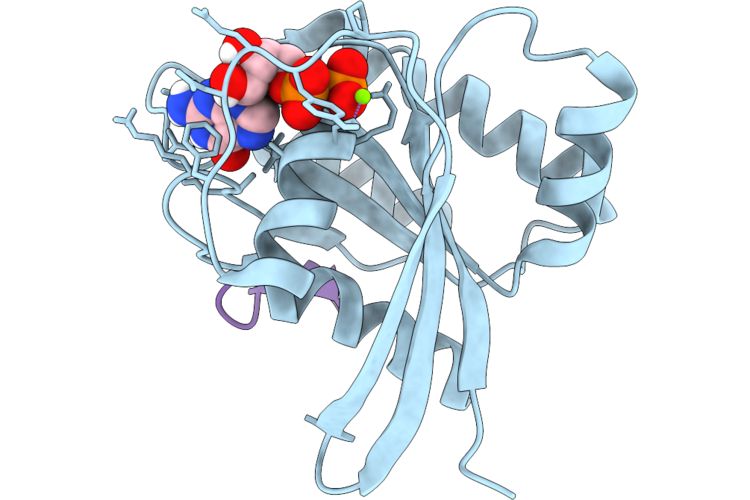 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: MG, GDP |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: MG, GDP, PO4 |
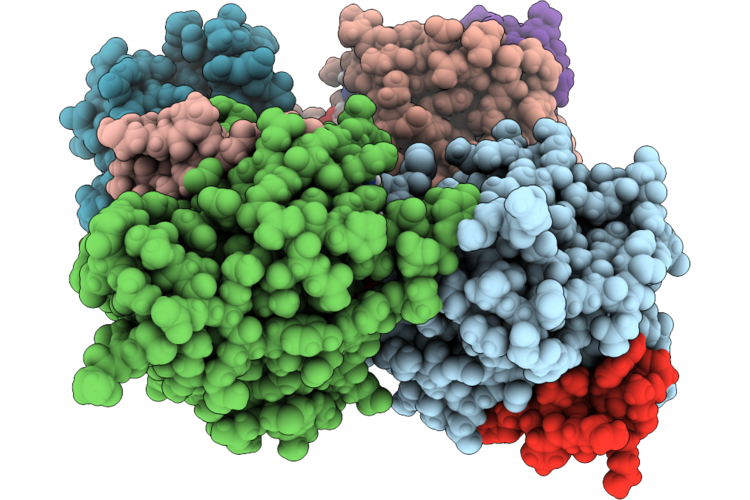 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP |
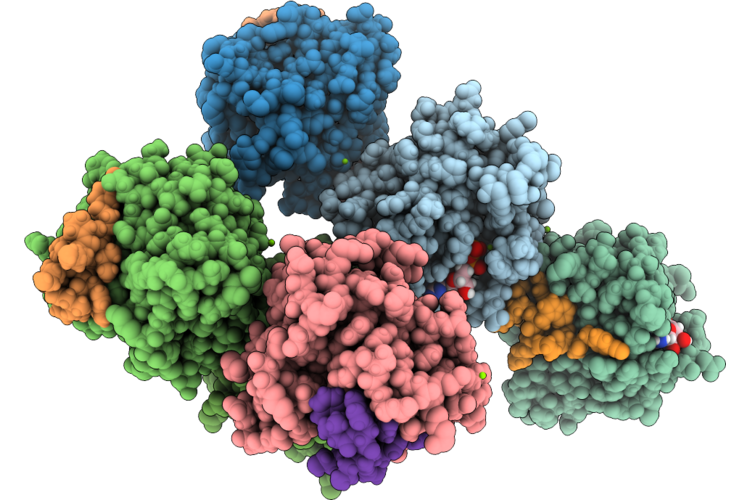 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
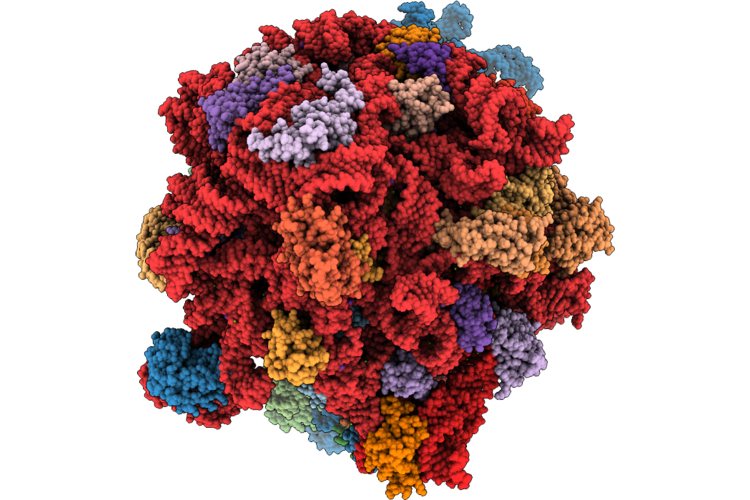 |
Doxycycline Bound E. Coli Ribosome With Rearranged Peptidyl Transferase Centre
Organism: Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RIBOSOME Ligands: ZN, MG, K, DXT |
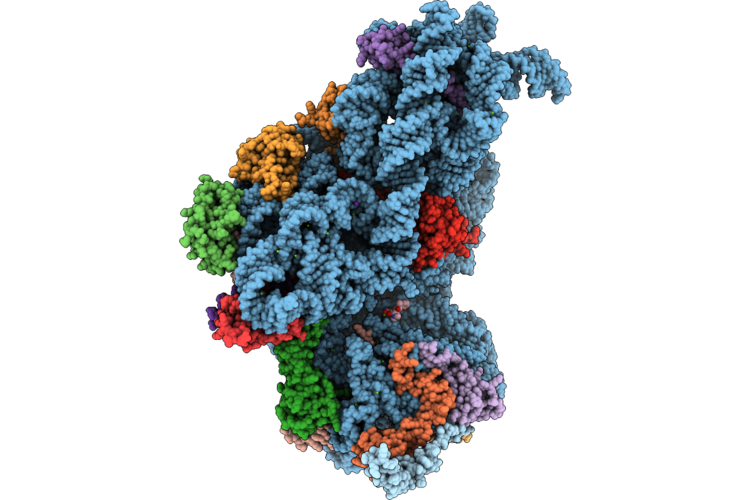 |
Organism: Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RIBOSOME Ligands: ZN, DXT, MG, K |
 |
Organism: Coxiella burnetii
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-04-22 Classification: RIBOSOME Ligands: ZN, K, MG |
 |
Organism: Coxiella burnetii
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RIBOSOME Ligands: DXT, K, MG, ZN |
 |
Triple Stack Doxycycline Bound 50S Subunit Of The Coxiella Burnetii Ribosome
Organism: Coxiella burnetii
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: ANTIBIOTIC Ligands: ZN, MG, K, DXT |

