Search Count: 40,336
All
Selected
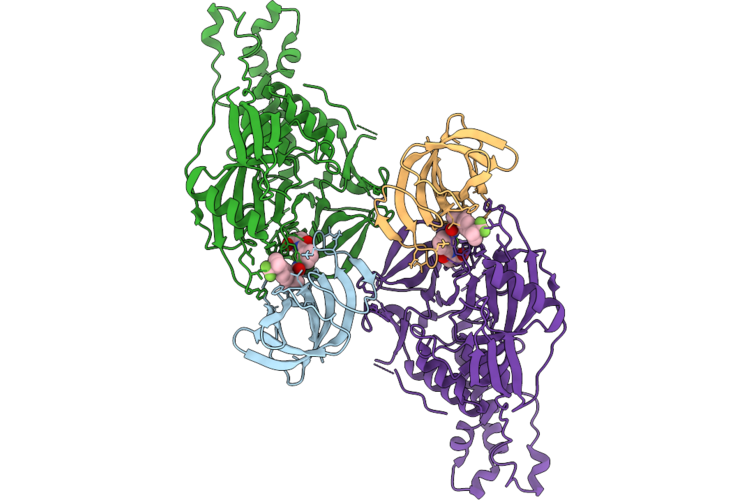 |
Cryo-Em Structure Of Crbn In Complex With Hbs1L And Tng-4857 (Focused Refinement)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: CYTOSOLIC PROTEIN Ligands: ZN, A1C9W |
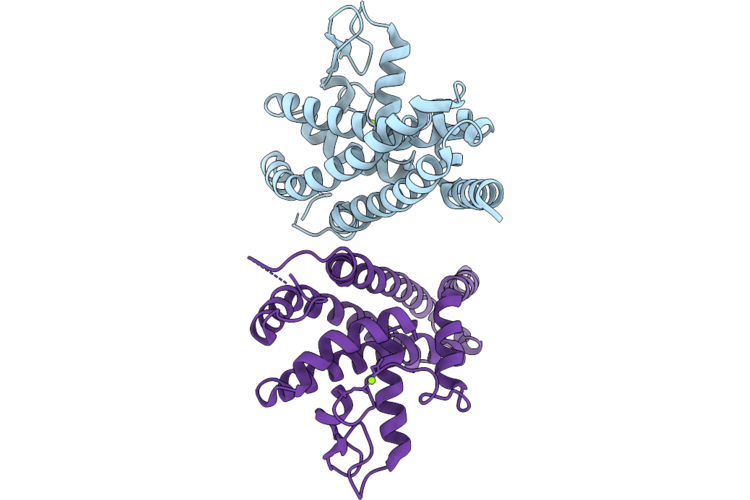 |
Crystal Structure Of Complement-Inhibiting Protein Chia Of Borrelia Recurrentis
Organism: Borrelia recurrentis
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-05-06 Classification: PROTEIN BINDING Ligands: MG |
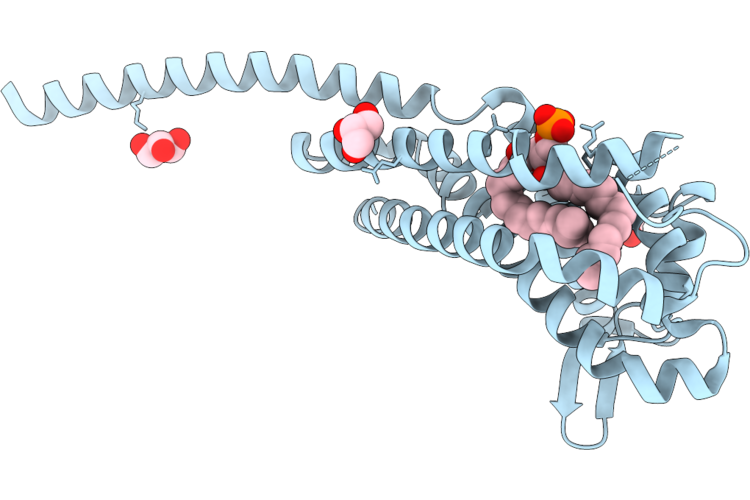 |
Crystal Structure Of Complement-Inhibiting Protein Chib Of Borrelia Recurrentis
Organism: Borrelia recurrentis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-06 Classification: PROTEIN BINDING Ligands: LPP, GOL |
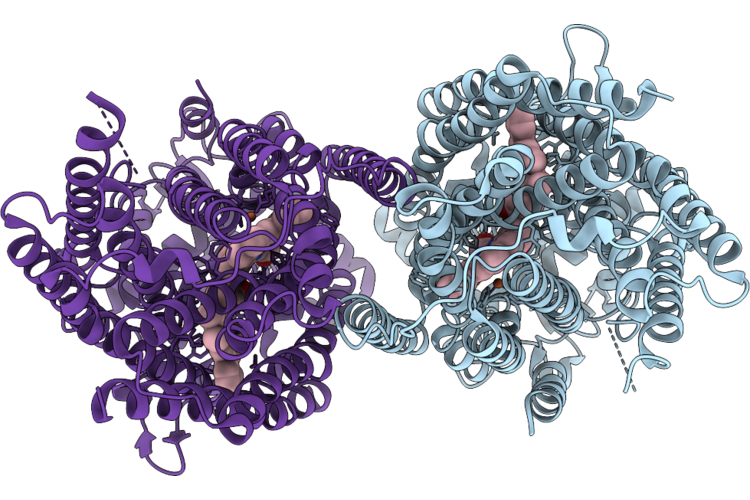 |
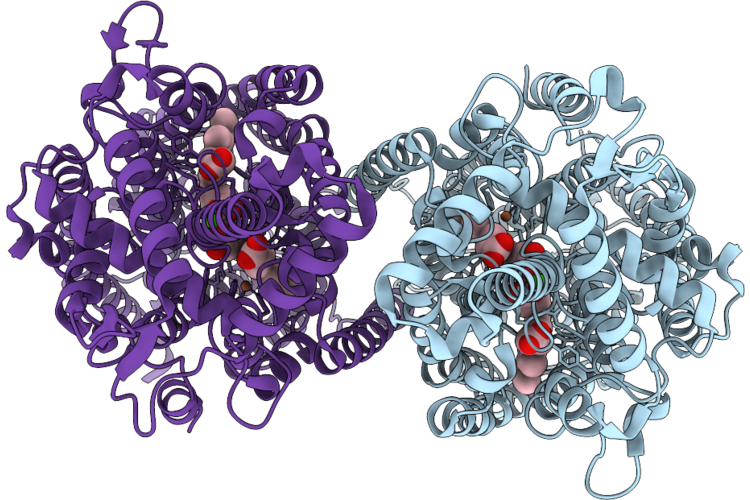
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 8.0 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
 |
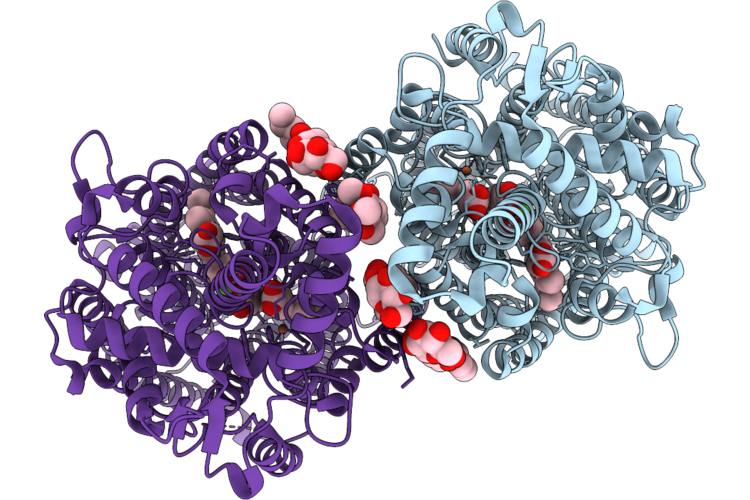
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Arg720Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
 |
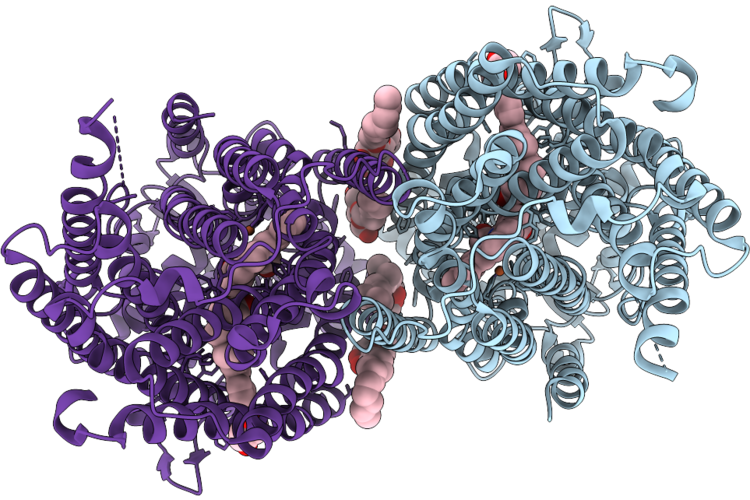
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, UQ5 |
 |
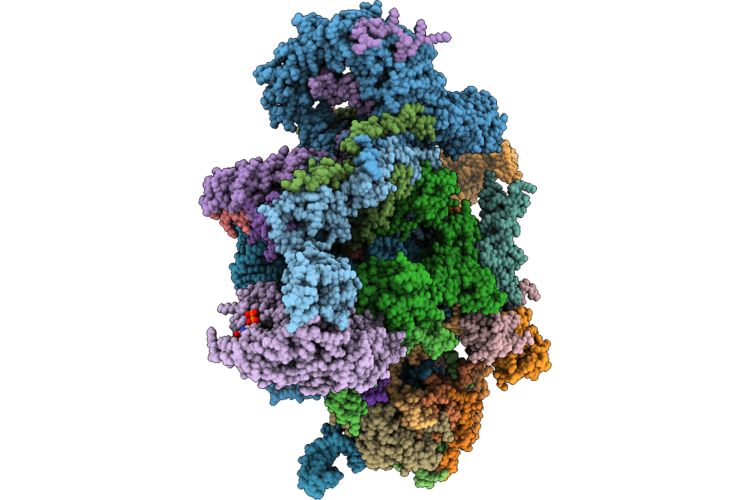
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant With Quino At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, UQ5, HQE, LMT |
 |
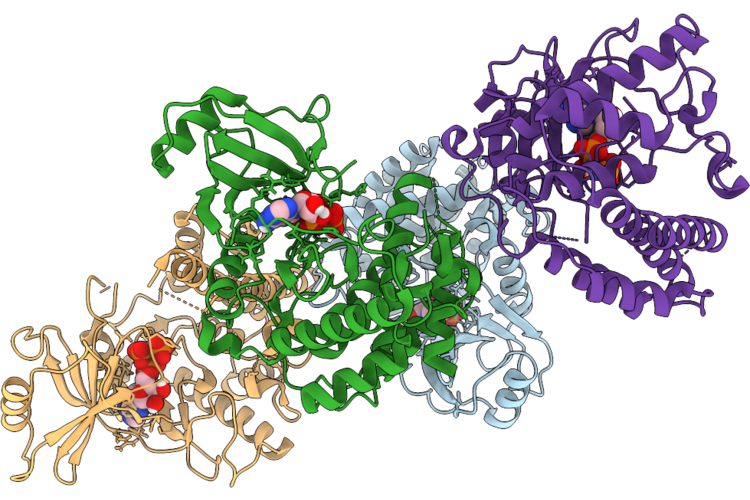
Assembly Intermediate Of Human Mitochondrial Ribosome Small Subunit In Complex With Noa1 And Partial Rbfa (State N2)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RIBOSOME Ligands: MG, K, ZN, FES, ATP, GDP |
 |
Caldalkalibacillus Thermarum Acyl-Coa Dehydrogenase Member 10 Bound To Anp And Magnesium
Organism: Caldalkalibacillus thermarum
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: ANP |
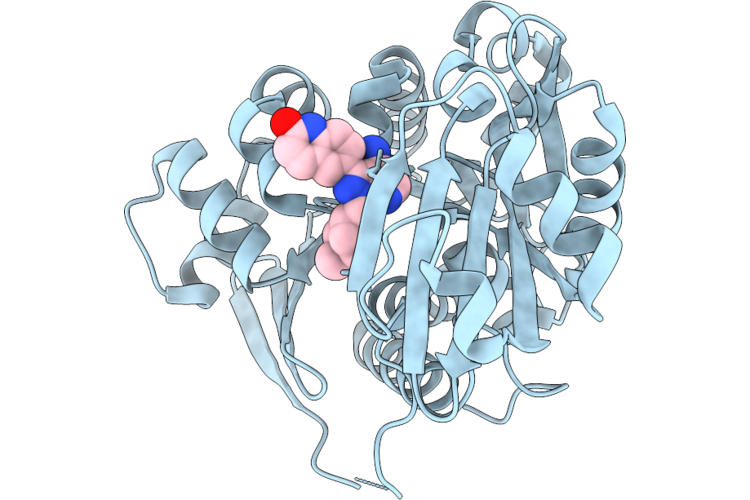 |
Crystal Structure Of Human Adenosine Kinase (Adk) In Complex With Inhibitor Bki-1817
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-05-06 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: CL, SO4, A1B96 |
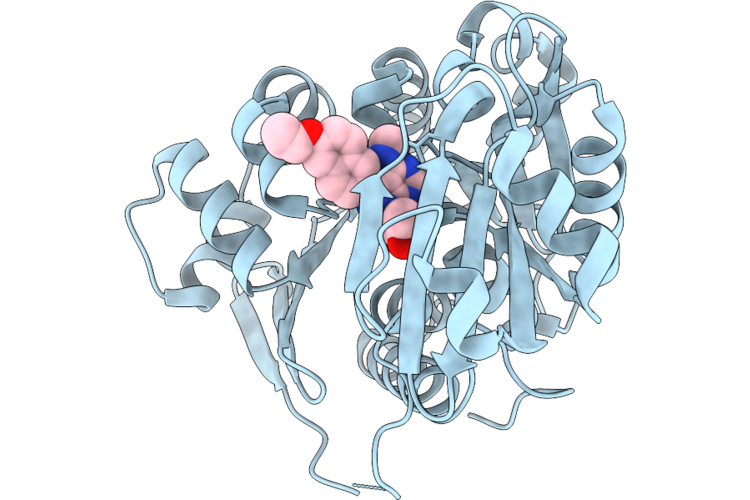 |
Crystal Structure Of Human Adenosine Kinase (Adk) In Complex With Inhibitor Bki-1676
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.81 Å Release Date: 2026-05-06 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1B95, SO4, CL |
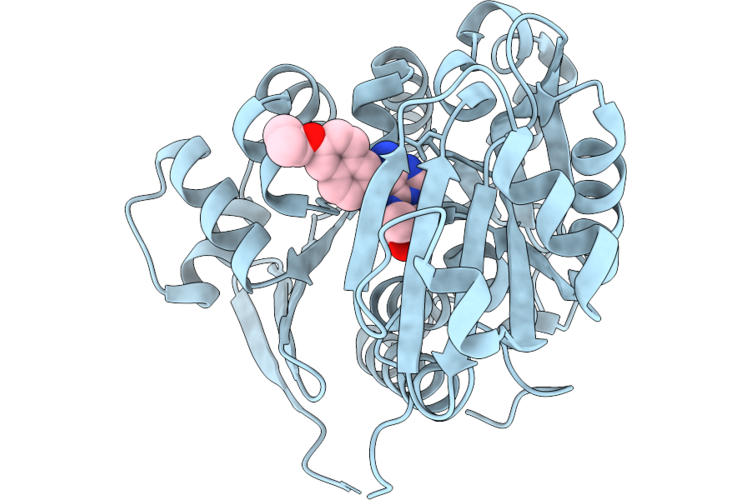 |
Crystal Structure Of Human Adenosine Kinase (Adk) In Complex With Inhibitor Bki-1553
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2026-05-06 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: CL, UW5, SO4 |
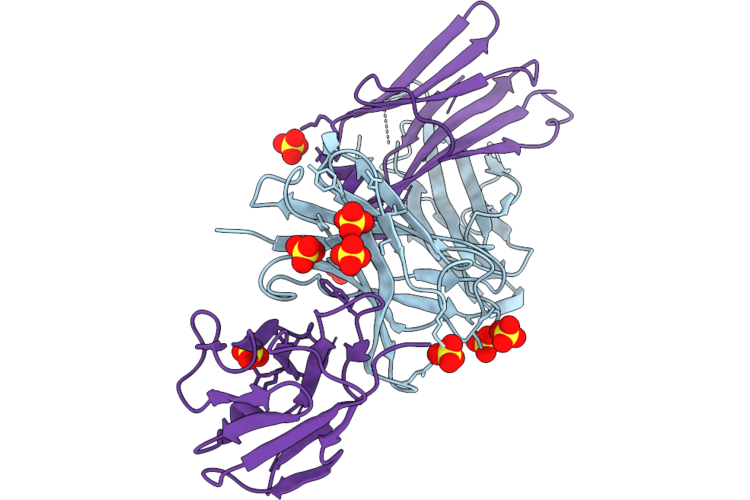 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-05-06 Classification: APOPTOSIS Ligands: SO4 |
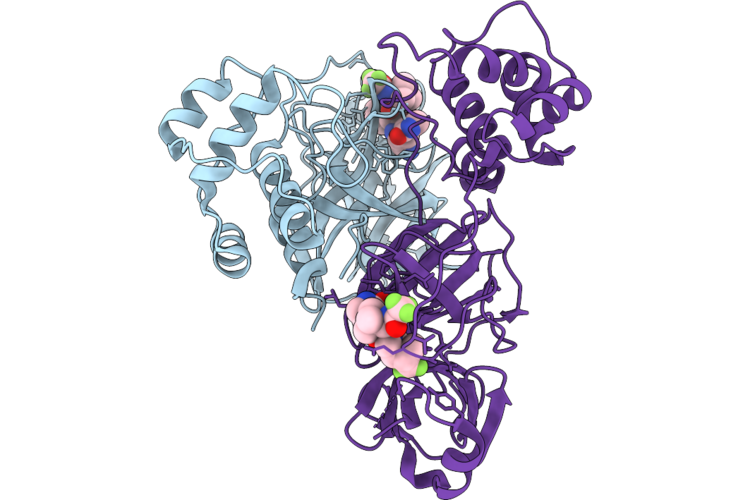 |
Organism: Human coronavirus hku1
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHF |
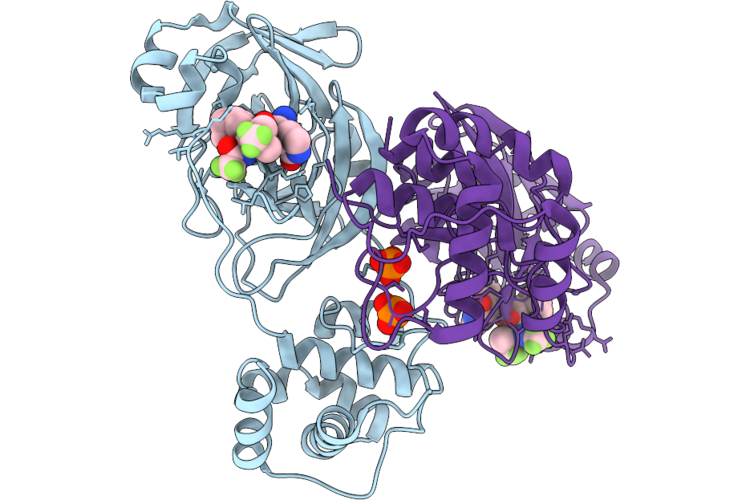 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHJ, PO4 |
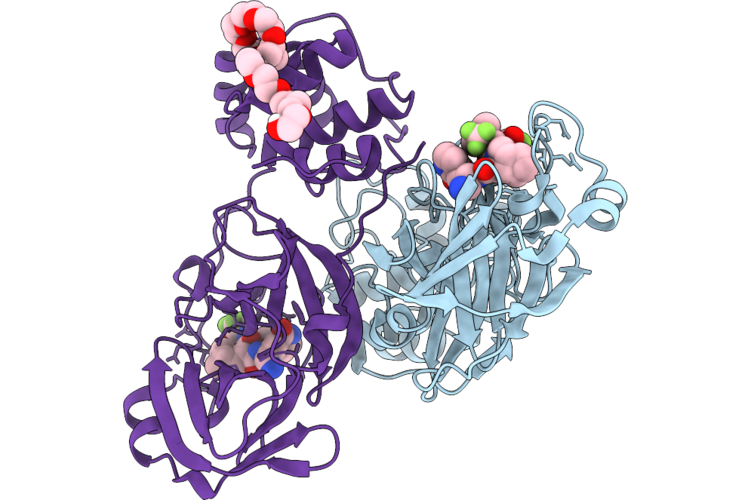 |
Organism: Human coronavirus 229e
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHJ, 12P |
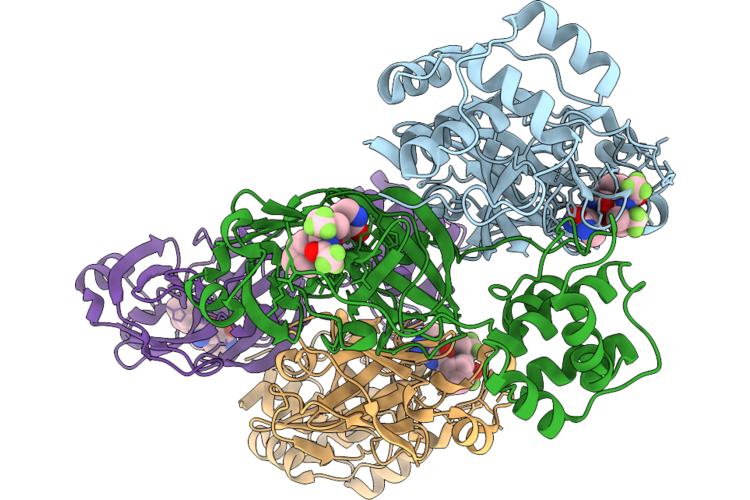 |
Organism: Middle east respiratory syndrome-related coronavirus
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHJ |
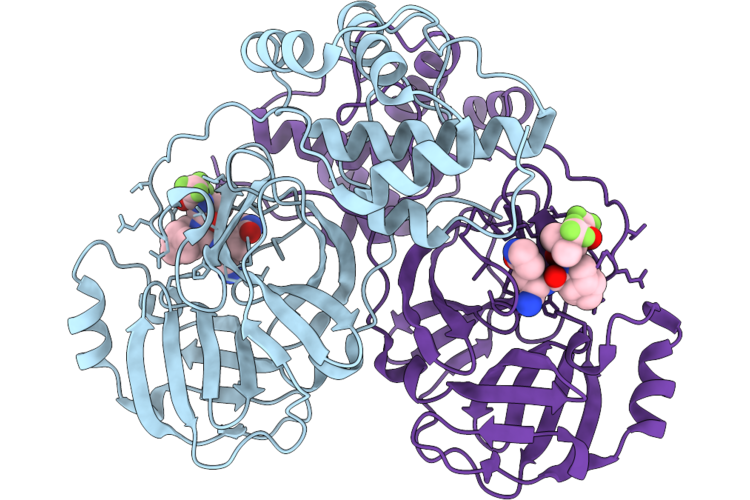 |
Organism: Infectious bronchitis virus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHJ |
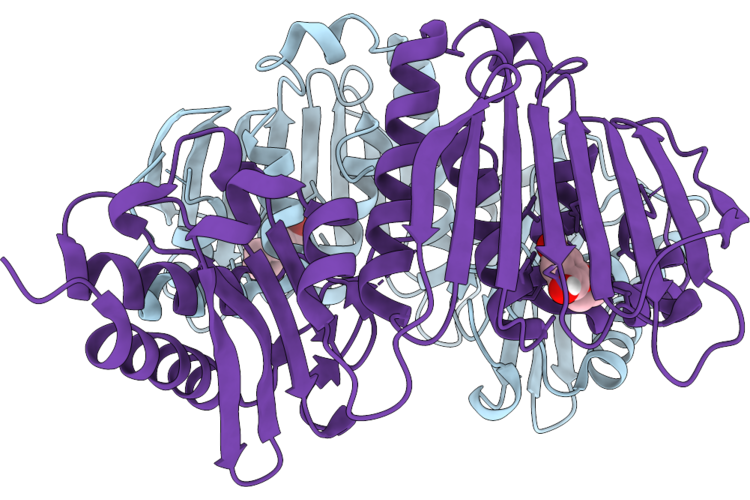 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: A1CKR |
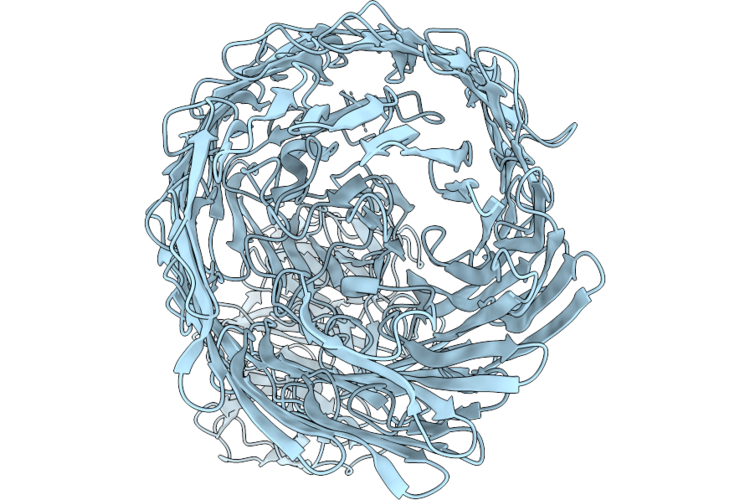 |
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TOXIN |

