Search Count: 59,363
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: CYTOSOLIC PROTEIN Ligands: A1C4S, ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: A1C9A |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: A1C9H |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: GOL, A1C9I |
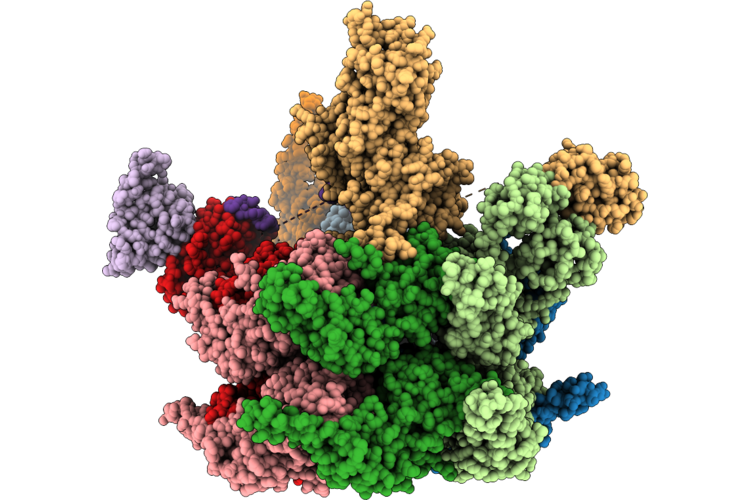 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.85 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ZN, ATP, ADP |
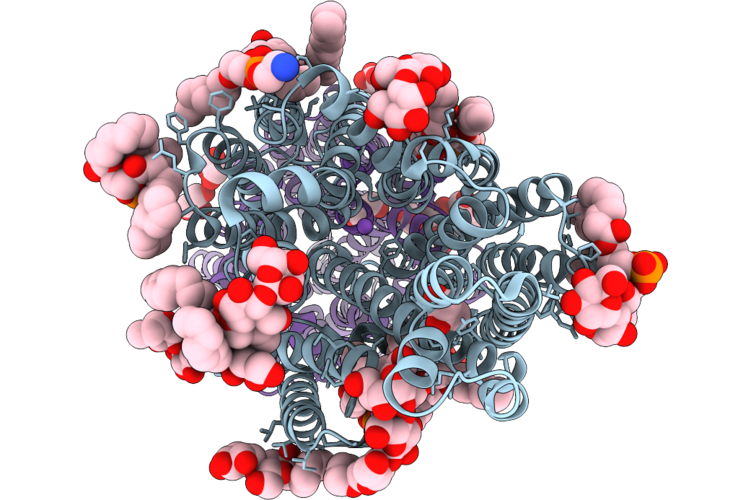 |
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PGT, LMU, PTY, 3PH, GOL, NA |
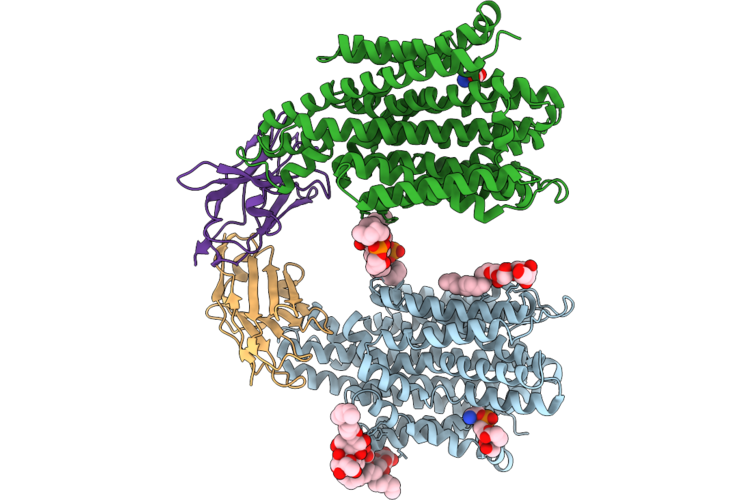 |
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Inward Open State In Complex With Nanobody 89
Organism: Staphylococcus aureus, Lama glama
Method: X-RAY DIFFRACTION Resolution:3.32 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PTY, PGT, LMU, OXM |
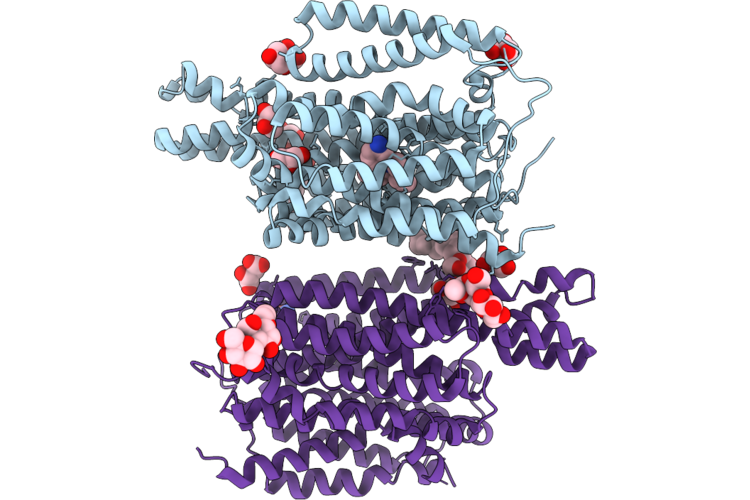 |
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State Bound To Ethidium
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ET, GOL, TLA, CL, LMU, OXM |
 |
Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans.
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Reprocessing And Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Crystal Structure Of A Computationally Designed Protein Bound To A Au-Containing Cofactor ([(Sulfanhc)Au.Trp])
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-22 Classification: DE NOVO PROTEIN Ligands: A1IRM |
 |
Organism: Mycobacterium phage ds6a
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: VIRUS Ligands: ZN, PO4 |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PEG, DSN, CA |
 |
Organism: Mycobacterium phage ds6a
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Crystal Structure Of Gh19 E228Q Domain Of D29-Lysa In Complex With (Glcnac)4
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, MPD |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, EDO, ALA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Crystal Structure Of Ami2B Domain Of Ds6A-Lysa In Complex With L-Ala-D-Iso-Gln-L-Lys- D-Ala-D-Ala
Organism: Mycobacterium phage ds6a, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN |
 |
Crystal Structure Of Gh19 Domain Of D29-Lysa In Complex With Product (Glcnac)
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: NAG |

