Search Count: 17
All
Selected
 |
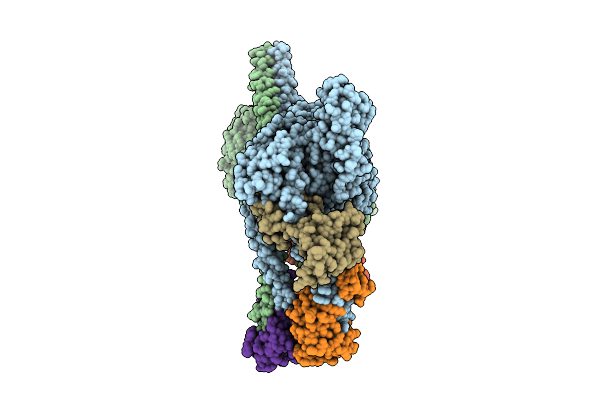
Cryo-Em Structure Of Human Beta-Cardiac Myosin In The Interacting-Heads Motif And S2-Fh Docked State
Organism: Homo sapiens, Saccharomyces cerevisiae, Aequorea victoria, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CONTRACTILE PROTEIN Ligands: ADP, PO4 |
 |
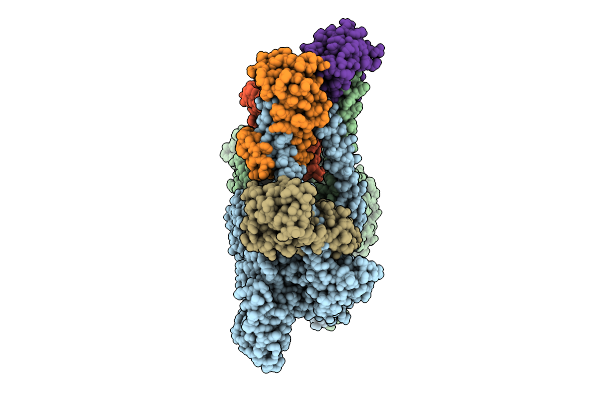
Cryo-Em Structure Of Human Beta-Cardiac Myosin Bound To Omecamtiv Mecarbil In The Interacting-Heads Motif And S2-Fh Docked State
Organism: Homo sapiens, Saccharomyces cerevisiae, Aequorea victoria, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:4.50 Å Release Date: 2026-04-08 Classification: CONTRACTILE PROTEIN Ligands: 2OW, ADP, PO4 |
 |
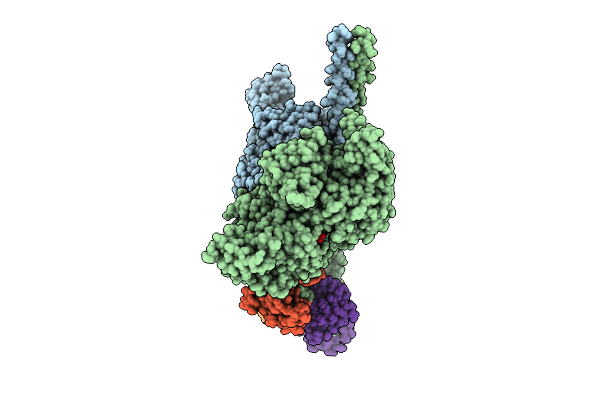
Cryo-Em Structure Of Human Beta-Cardiac Myosin Bound To Mavacamten In The Interacting-Heads Motif And S2-Fh Docked State
Organism: Homo sapiens, Saccharomyces cerevisiae, Aequorea victoria, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CONTRACTILE PROTEIN Ligands: PO4, XB2, ADP |
 |
Cryo-Em Structure Of Human Beta-Cardiac Myosin Bound To Mavacamten In The Interacting-Heads Motif And S2-Fh Undocked State
Organism: Homo sapiens, Saccharomyces cerevisiae, Aequorea victoria, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CONTRACTILE PROTEIN Ligands: XB2, ADP, PO4 |
 |
Cryo-Em Structure Of Human Beta-Cardiac Myosin In The Interacting-Heads Motif And S2-Fh Undocked State
Organism: Homo sapiens, Saccharomyces cerevisiae, Aequorea victoria, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-04-08 Classification: CONTRACTILE PROTEIN Ligands: ADP, PO4 |
 |
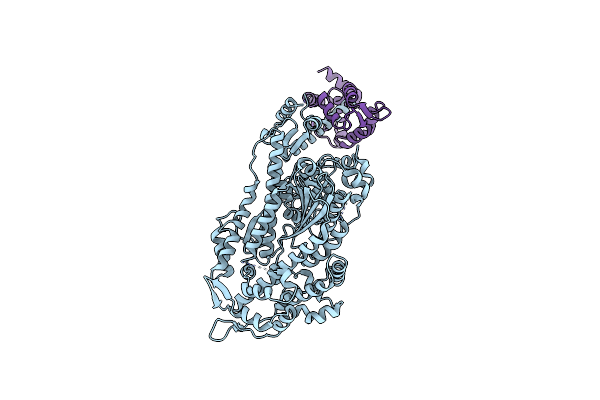
Cryo-Em Structure Of Human Beta-Cardiac Myosin Bound To Omecamtiv Mecarbil In The Interacting-Heads Motif And S2-Fh Undocked State
Organism: Homo sapiens, Saccharomyces cerevisiae, Aequorea victoria, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2026-04-08 Classification: CONTRACTILE PROTEIN Ligands: 2OW, ADP, PO4 |
 |
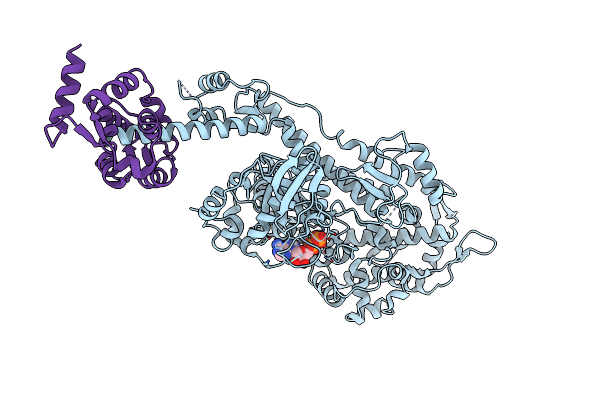
Human Cardiac Myosin Ii And Associated Essential Light Chain In The Rigor Conformation
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2023-01-11 Classification: CONTRACTILE PROTEIN |
 |
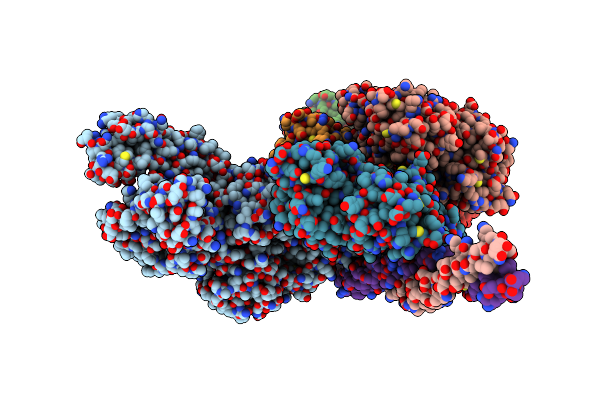
Human Beta-Cardiac Myosin Ii Bound To Adp-Mg2+ And The Associated Essential Light Chain
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2023-01-11 Classification: CONTRACTILE PROTEIN Ligands: MG, ADP |
 |
Helical Reconstruction Of The Human Cardiac Actin-Tropomyosin-Myosin Complex In Complex With Adp-Mg2+
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2023-01-11 Classification: MOTOR PROTEIN Ligands: MG, ADP |
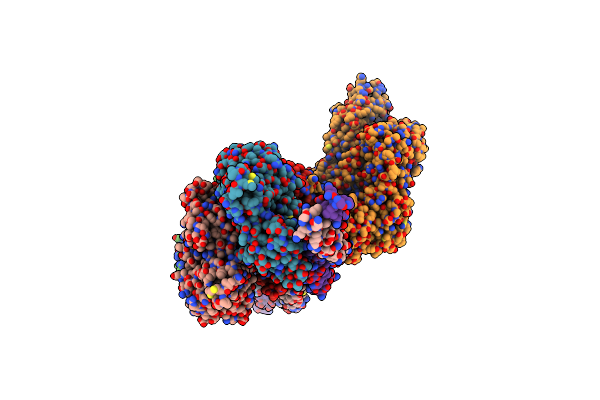 |
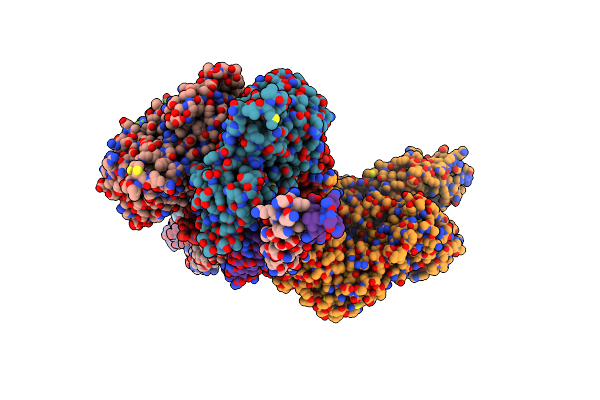
Helical Reconstruction Of The Human Cardiac Actin-Tropomyosin-Myosin Complex In The Rigor Form
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2022-11-23 Classification: MOTOR PROTEIN Ligands: ADP, MG |
 |
Helical Reconstruction Of The Human Cardiac Actin-Tropomyosin-Myosin Loop 4 7G Mutant Complex
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2022-11-23 Classification: MOTOR PROTEIN Ligands: ADP, MG |
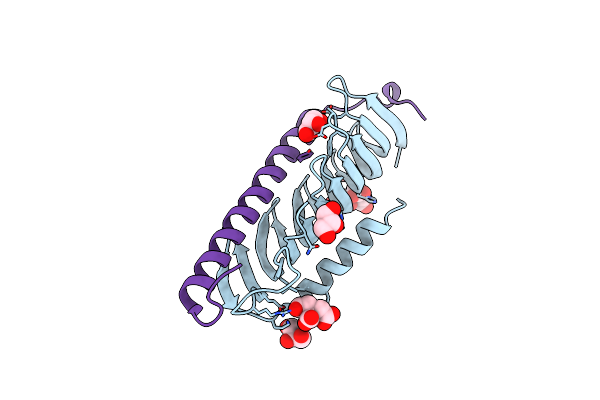 |
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2019-07-24 Classification: MOTOR PROTEIN Ligands: GOL, CIT |
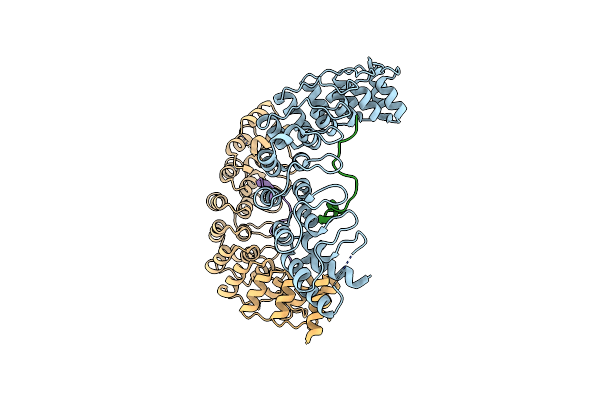 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2016-02-03 Classification: PROTEIN BINDING/MOTOR PROTEIN |
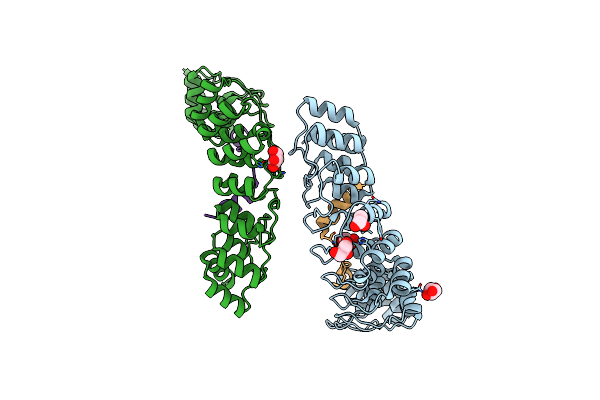 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2016-02-03 Classification: PROTEIN BINDING/MOTOR PROTEIN Ligands: GOL |
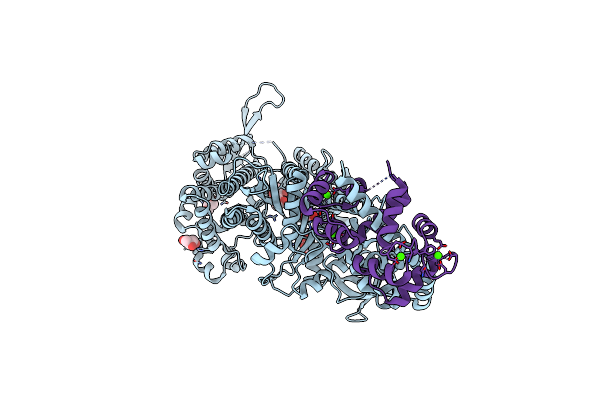 |
Organism: Sus scrofa, Drosophila melanogaster
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2005-06-07 Classification: MOTOR PROTEIN/METAL-BINDING PROTEIN Ligands: GOL, CA |
 |
Organism: Sus scrofa, Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2005-06-07 Classification: MOTOR PROTEIN/METAL-BINDING PROTEIN Ligands: SO4, CA |
 |
Crystal Structure Of Myosin V Motor With Essential Light Chain-Nucleotide-Free
Organism: Gallus gallus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2003-09-26 Classification: ATPASE/MYOSIN Ligands: SO4 |

