Search Count: 1,277
All
Selected
 |
Organism: Homo sapiens, Trichoplusia ni
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-05-20 Classification: CELL CYCLE |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.54 Å Release Date: 2026-05-20 Classification: CELL CYCLE |
 |
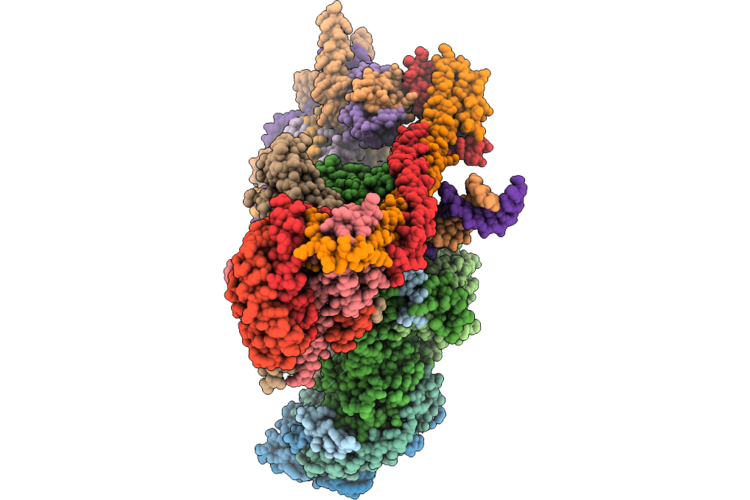
Structure Of The Human Inner Kinetochore Ccan Bound To A Mono-Cenp-A Nucleosome
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.50 Å Release Date: 2026-05-20 Classification: CELL CYCLE |
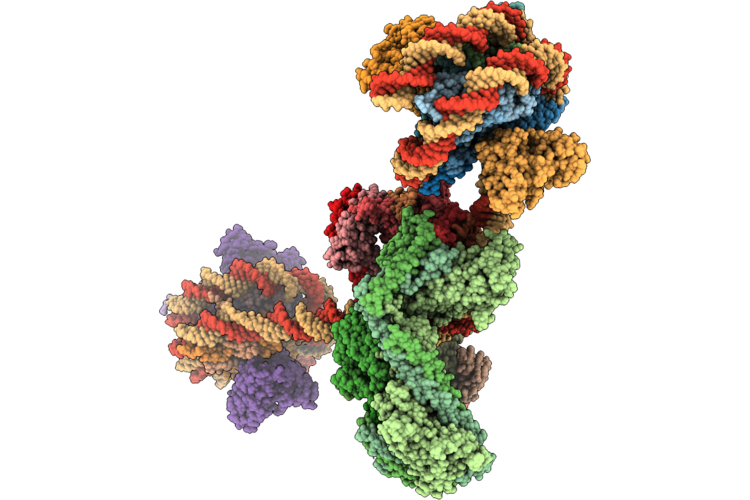 |
Structure Of The Human Inner Kinetochore Ccan Bound To A Di-Cenp-A Nucleosome
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:15.50 Å Release Date: 2026-05-20 Classification: CELL CYCLE |
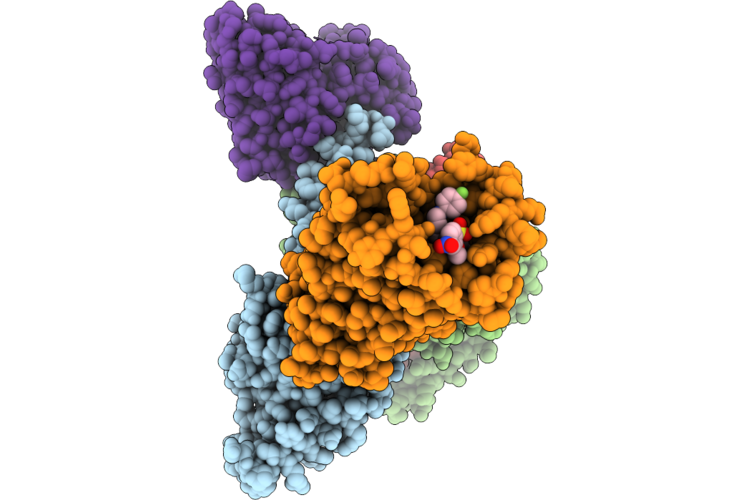 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-05-20 Classification: PEPTIDE BINDING PROTEIN Ligands: A1MCH |
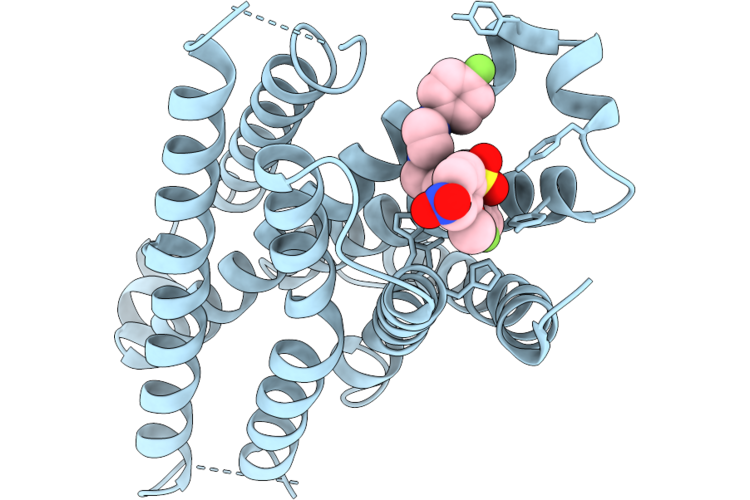 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.31 Å Release Date: 2026-05-20 Classification: PEPTIDE BINDING PROTEIN Ligands: A1MCH |
 |
Organism: Schisandra chinensis
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-05-13 Classification: TRANSFERASE Ligands: SAM |
 |
Organism: Schisandra chinensis
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-05-13 Classification: TRANSFERASE Ligands: SAM, A1E80 |
 |
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: A1L9Y |
 |
Organism: Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: SIGNALING PROTEIN Ligands: A1L9Y |
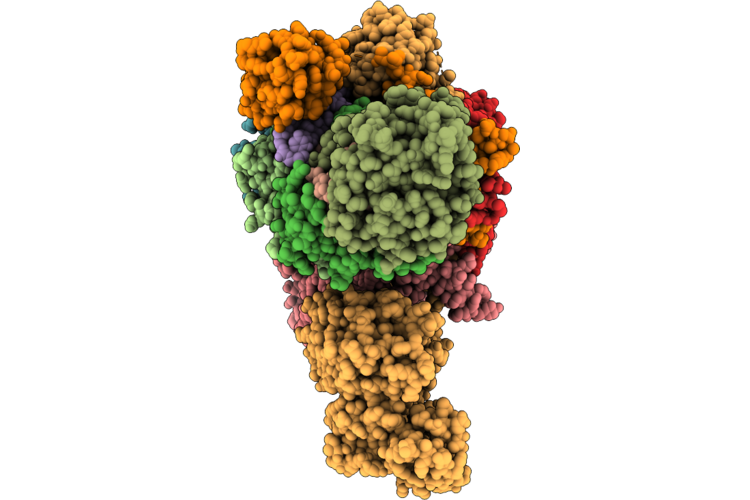 |
Cryo-Em Structure Of Type Vii Crispr-Cas Complex At The Target Engagement State
Organism: Metagenome
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: ANTIVIRAL PROTEIN/RNA Ligands: ZN |
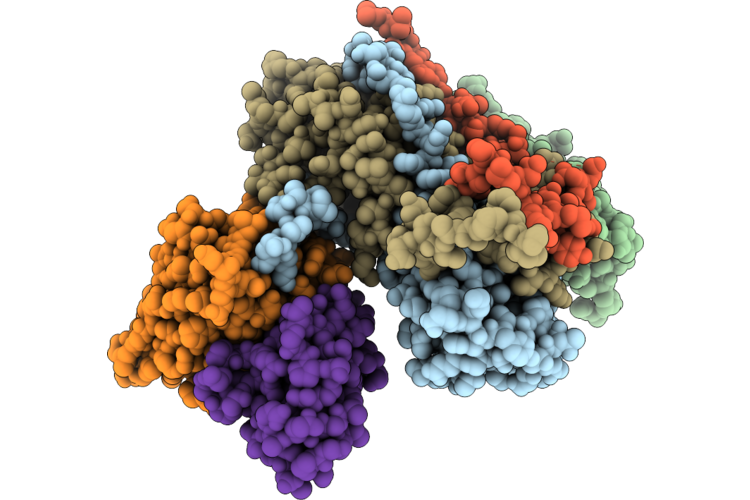 |
Complex Of Fmdv O/18074 And Porcine-Derived Neutralizing Monoclonal Antibody Po18-10
Organism: Sus scrofa, Foot-and-mouth disease virus o
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2026-04-22 Classification: VIRUS |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.77 Å Release Date: 2026-04-15 Classification: PEPTIDE BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.54 Å Release Date: 2026-04-15 Classification: PEPTIDE BINDING PROTEIN |
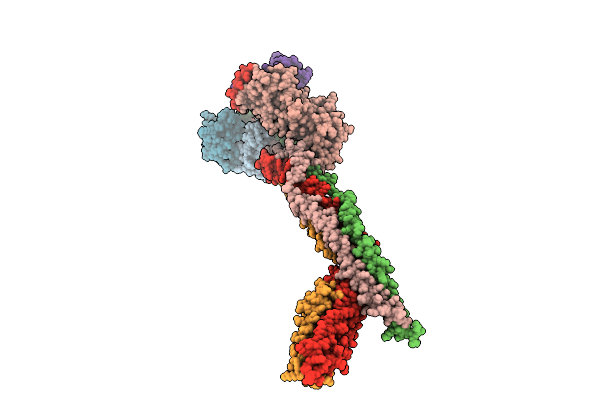 |
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Lacking The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
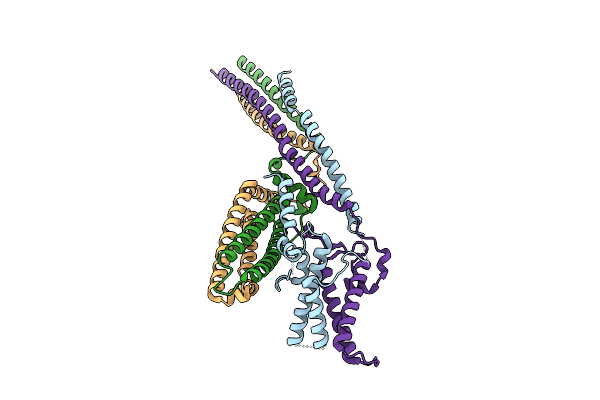 |
Cryo-Em Structure Of The Base Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
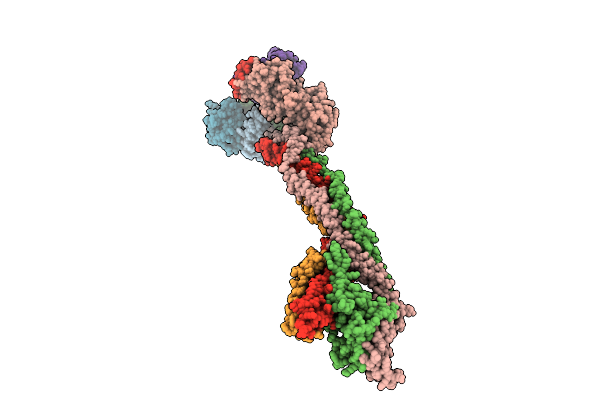 |
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae, Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
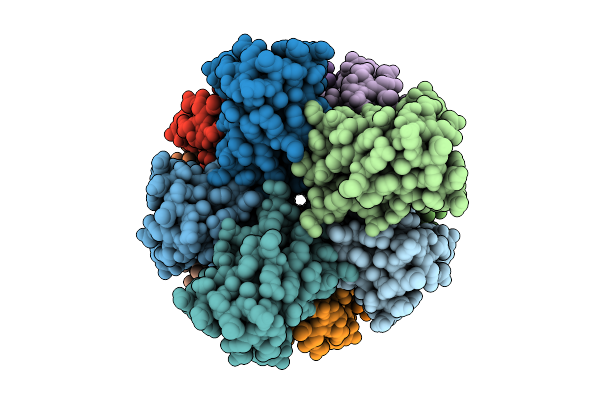 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: PROTEIN FIBRIL |
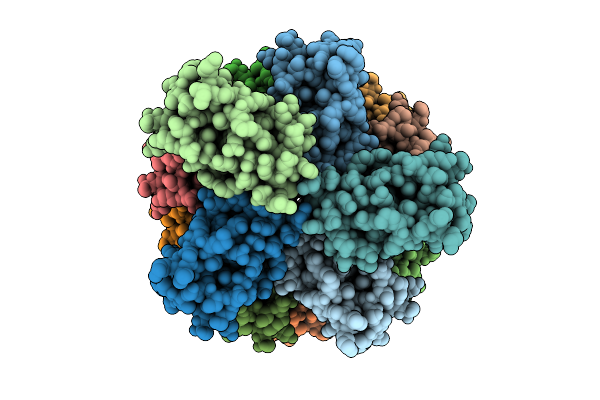 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: PROTEIN FIBRIL |
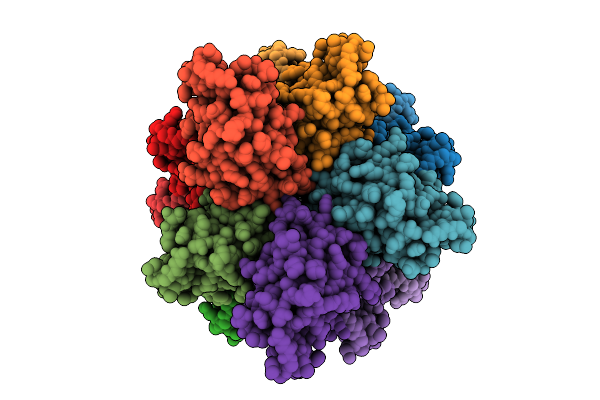 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: APOPTOSIS |

