Search Count: 1,990
All
Selected
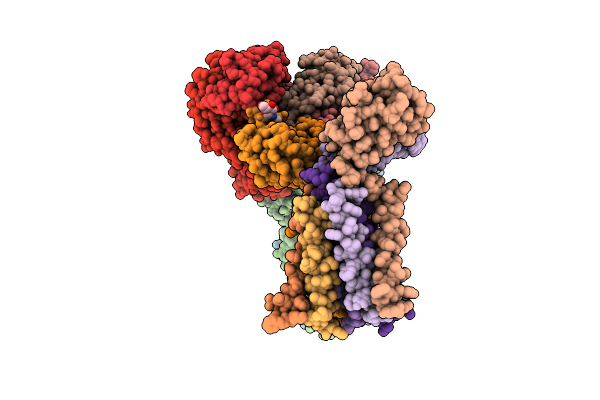 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.35 Å Release Date: 2026-04-15 Classification: IMMUNE SYSTEM Ligands: NAG |
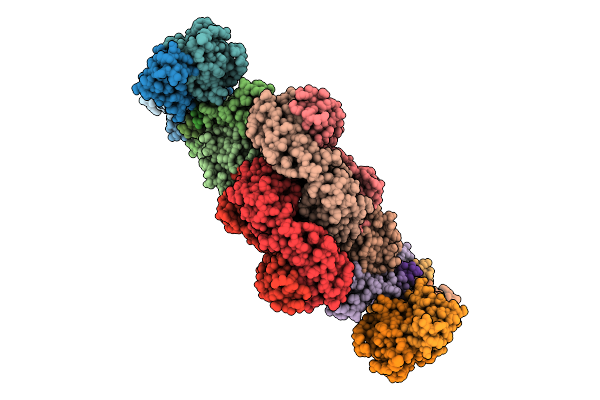 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN/DNA Ligands: GOL, PO4 |
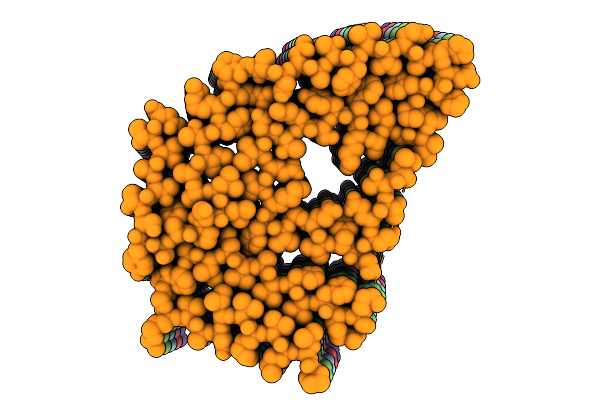 |
Cryo-Em Structure Of Hepatic Amyloid Fibril From A Variant Attrv122Delta, Single Filament Morphology
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
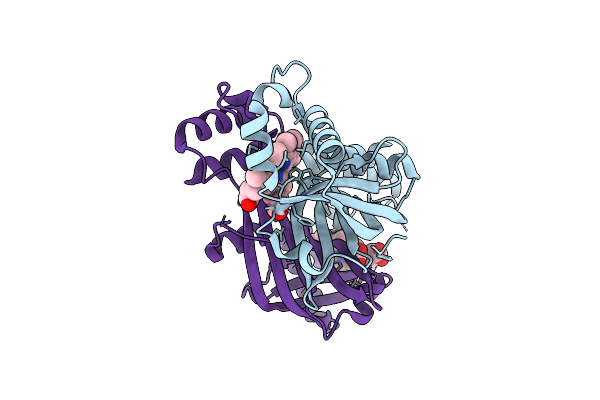 |
The Crystal Structure Of Paib From Bacillus Stearothermophilus Bound To Hem
Organism: Geobacillus kaustophilus hta426
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2026-04-01 Classification: TRANSCRIPTION Ligands: HEM |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: CHAPERONE |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: MLA |
 |
Crystal Structure Of Topbp1Brct7-8 Domain In Complex With 1-Adamantaneacetic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: A1ENP |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: DQC |
 |
Organism: Mus musculus, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: GGL, A1EA4 |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN |
 |
The Crystal Structure Of Human Dead-Box Rna Helicase Ddx28 Reca1 Domain In Complex With Adp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-03-04 Classification: RIBOSOMAL PROTEIN Ligands: PO4, ADP |
 |
Chemical Inhibition Of The N-Acetyltaurine Amidohydrolase Pter Reduces Food Intake And Obesity
Organism: Amphimedon queenslandica
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: ZN |
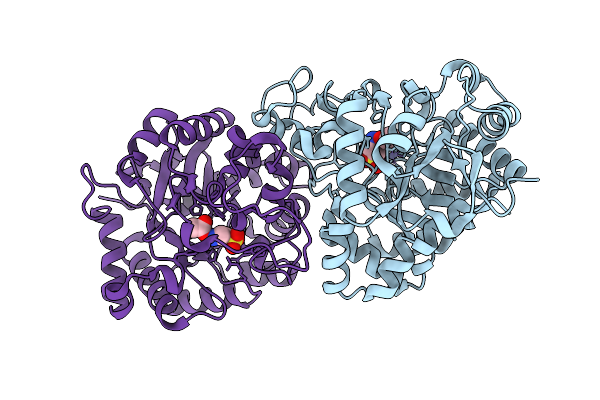 |
Chemical Inhibition Of The N-Acetyltaurine Amidohydrolase Pter Reduces Food Intake And Obesity
Organism: Amphimedon queenslandica
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: TAU, ACT, ZN |

