Search Count: 2,812
All
Selected
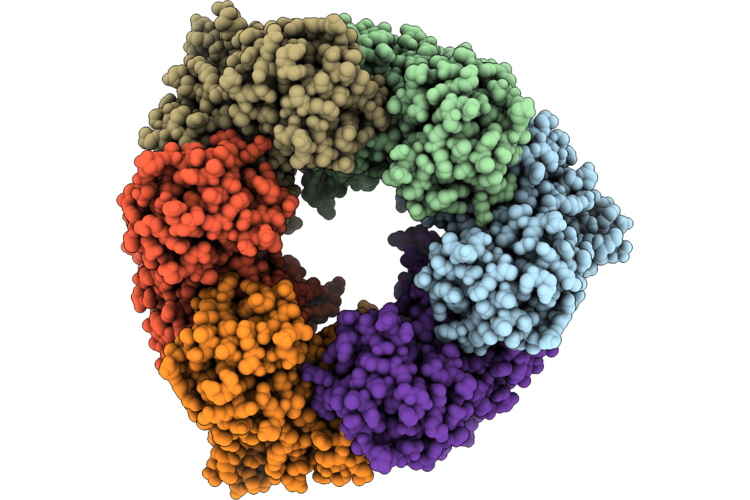 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 1
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
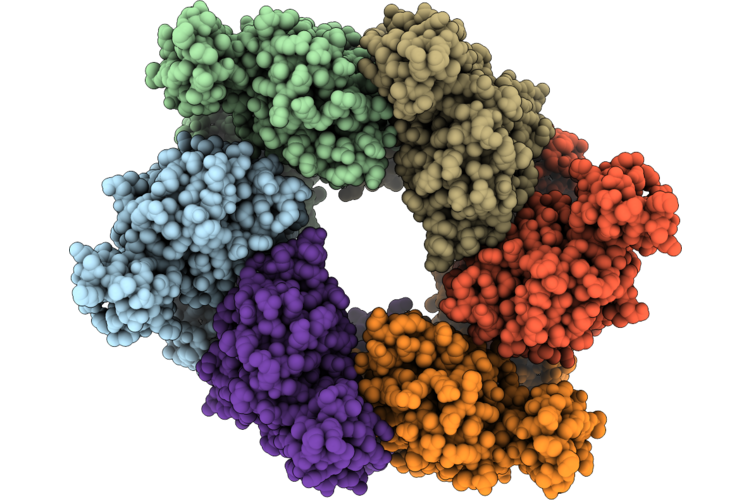 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 2
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
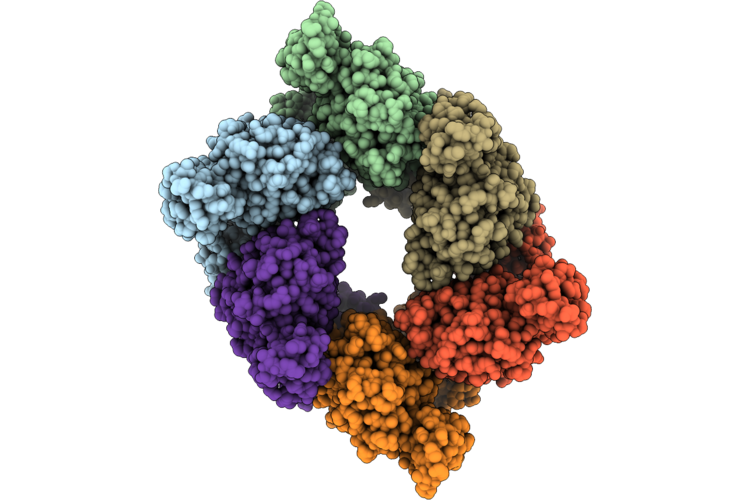 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 3
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
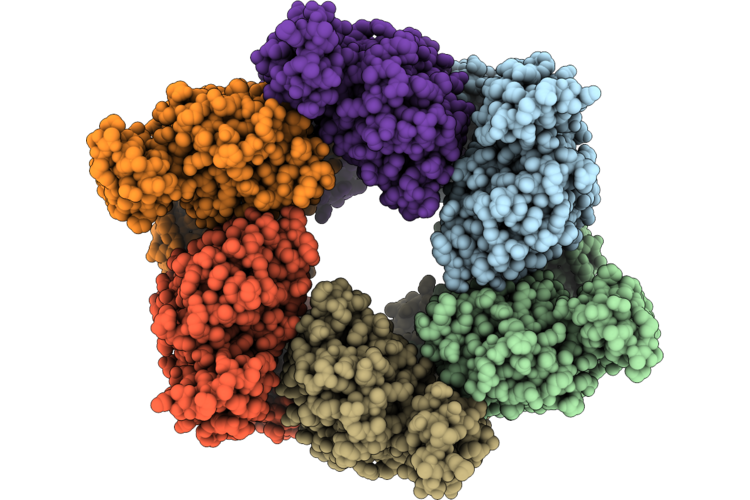 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 4
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
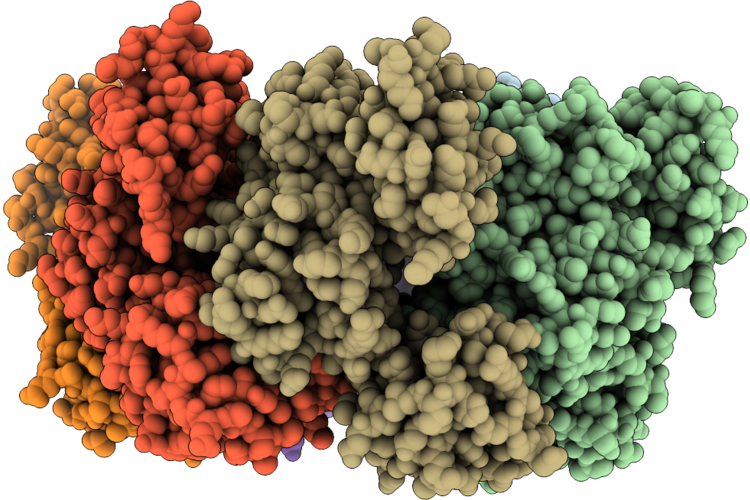 |
Crystal Structure Of Vibrio Cholerae Pilu, A Pilt-Dependent Retraction Atpase - Crystal Form 1.
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
 |
Crystal Structure Of Vibrio Cholerae Pilu, A Pilt-Dependent Retraction Atpase - Crystal Form 2
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-20 Classification: MOTOR PROTEIN Ligands: SO4 |
 |
Organism: Vibrio cholerae, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:3.64 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
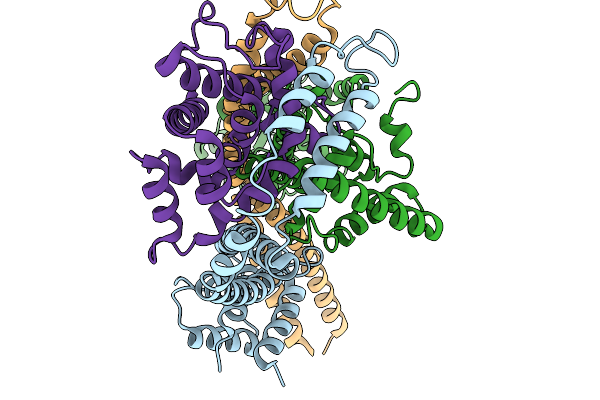 |
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:3.54 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:4.07 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
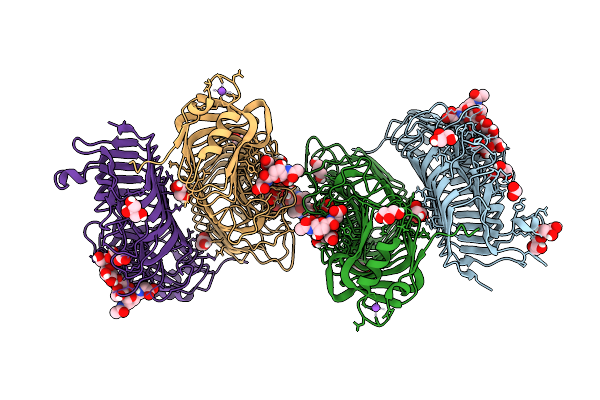 |
Crystal Structure Of The Polysaccharide Lyase Rbmb From Vibrio Cholerae Bound To Vibrio Polysaccharide
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-03-11 Classification: LYASE |
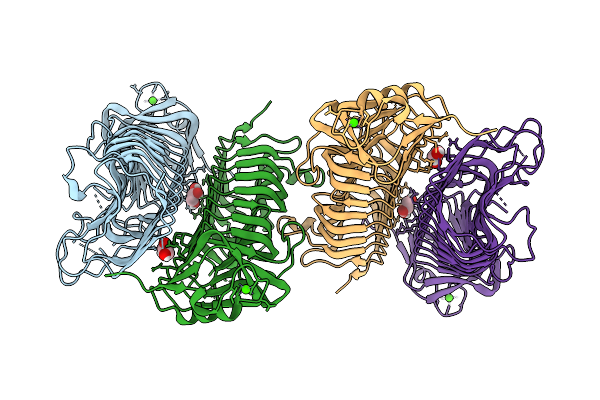 |
Organism: Vibrio cholerae c6706
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: LYASE Ligands: GOL, CA |
 |
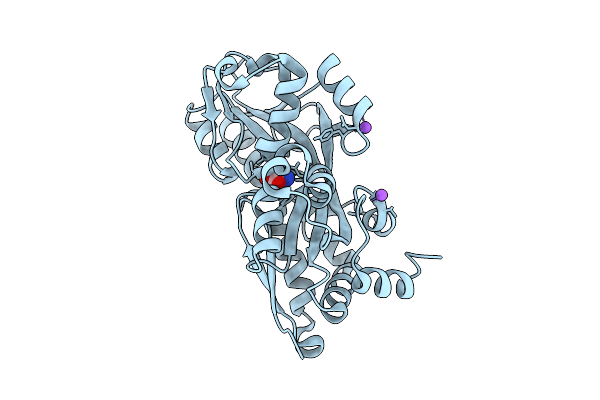
Cryo-Em Structure Of The Glucose-Specific Pts Transporter Iic From V. Cholerae In The Inward-Facing Conformation
Organism: Vibrio cholerae m66-2
Method: ELECTRON MICROSCOPY Resolution:3.68 Å Release Date: 2026-01-14 Classification: TRANSPORT PROTEIN |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2025-12-24 Classification: TRANSPORT PROTEIN |
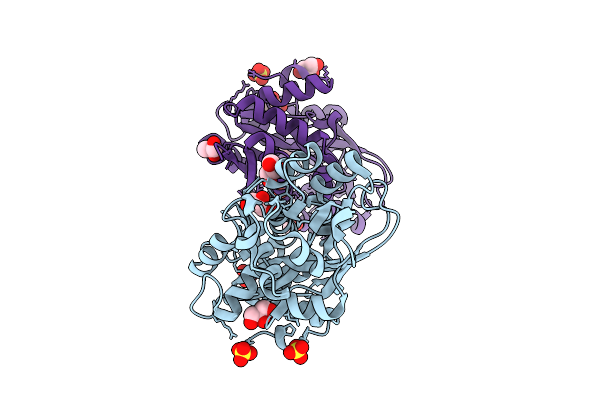 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: TRANSPORT PROTEIN Ligands: GOL, SO4 |
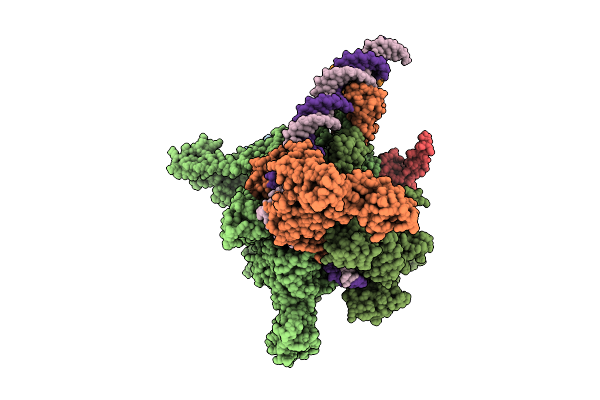 |
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Holoenzyme Bound To An Ompu Promoter Dna Fragment
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN |
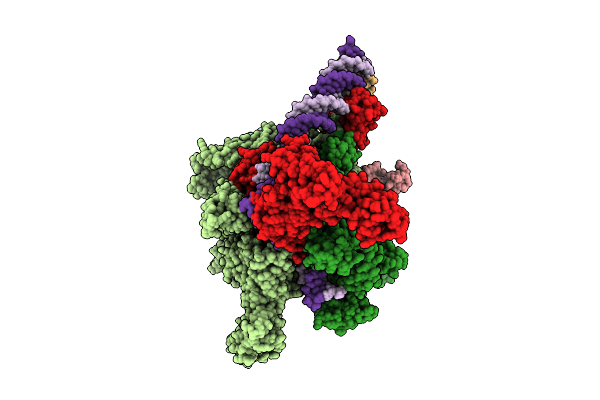 |
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Holoenzyme Bound To An Ompu Promoter Dna Fragment And 5-Mer Rna
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
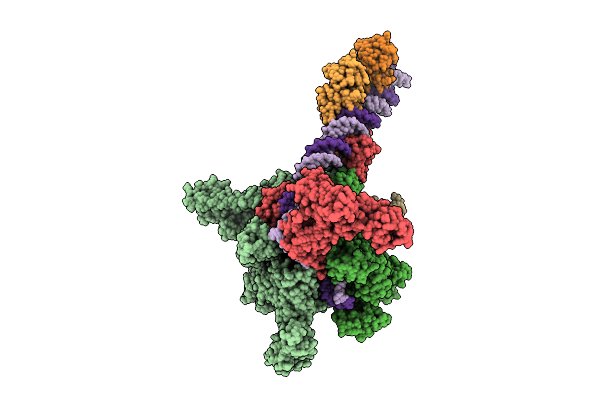
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Toxr Transcription Factor And Ompu Promoter Dna
Organism: Vibrio cholerae o395, Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Tcpp Transcription Factor And A Toxt Promoter Dna Fragment
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Toxr And Tcpp Transcription Factors And A Toxt Promoter Dna Fragment
Organism: Vibrio cholerae o395, Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN |

