Search Count: 20
All
Selected
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica Bound To C-Terminal Region Of Collagen Model Peptide (Pro-Hyp-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
Organism: Hathewaya histolytica
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: ZN, CA |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica In Complex With Collagen Model Peptide (Pro-Hyp-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica In Complex With Collagen Model Peptide (Pro-Hyp-Gly)12
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica Bound To C-Terminal Region Of The Collagen-Binding Protein Colh (Pro-Pro-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
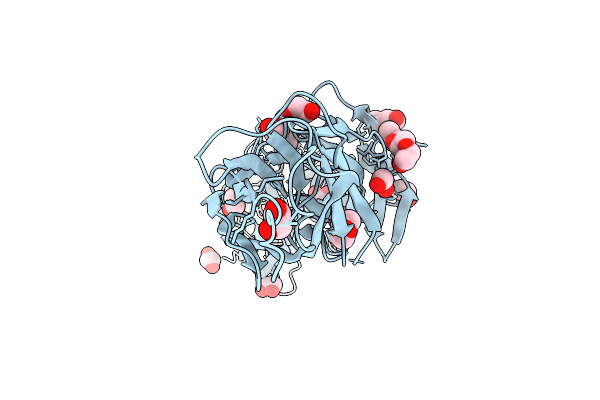 |
Organism: Streptococcus sanguinis sk36
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-21 Classification: CELL ADHESION Ligands: EDO, PEG, PGE |
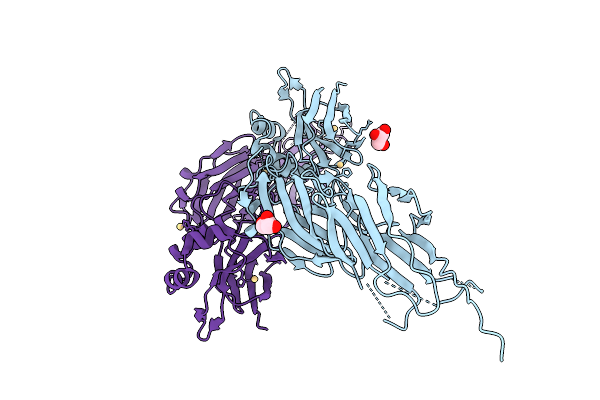 |
Organism: Streptococcus sanguinis
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-01-21 Classification: CELL ADHESION Ligands: CD, GOL |
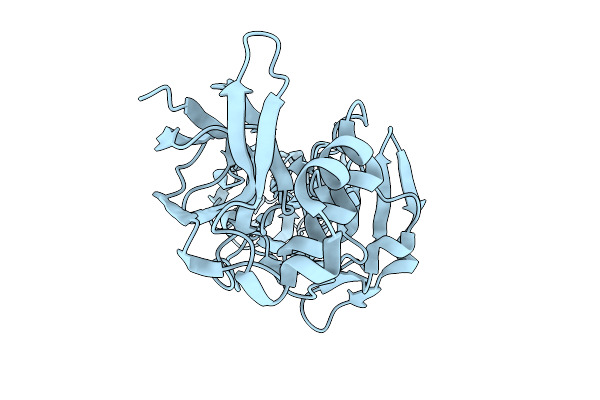 |
Crystal Structure Of Se-Met Ssa1633 From Streptococcus Sanguinis (Phase Determination Model)
Organism: Streptococcus sanguinis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-01-21 Classification: CELL ADHESION |
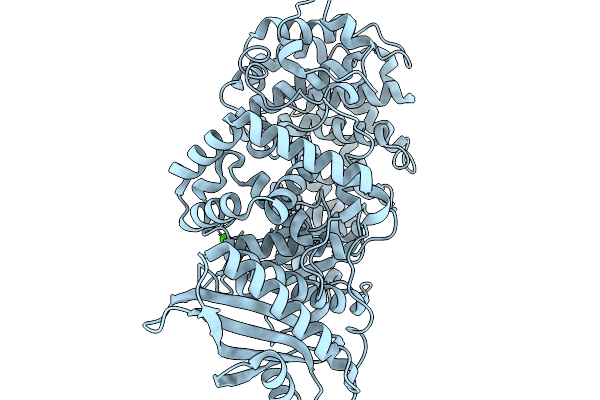 |
Room Temperature Crystal Structure Of Collagenase Colh From Hathewaya Histolytica At 2.7 Angstrom Resolution
Organism: Hathewaya histolytica
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ZN, CA |
 |
Organism: Streptococcus pyogenes serotype m3
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2023-12-06 Classification: PROTEIN FIBRIL Ligands: GOL |
 |
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2023-05-31 Classification: TRANSFERASE Ligands: SAM, DNI, MG, CL, NA |
 |
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-04-12 Classification: TRANSFERASE Ligands: SAH, DNI, MG |
 |
Organism: Hericium erinaceus
Method: X-RAY DIFFRACTION Resolution:1.29 Å Release Date: 2022-11-23 Classification: DNA BINDING PROTEIN Ligands: GMP, PEG |
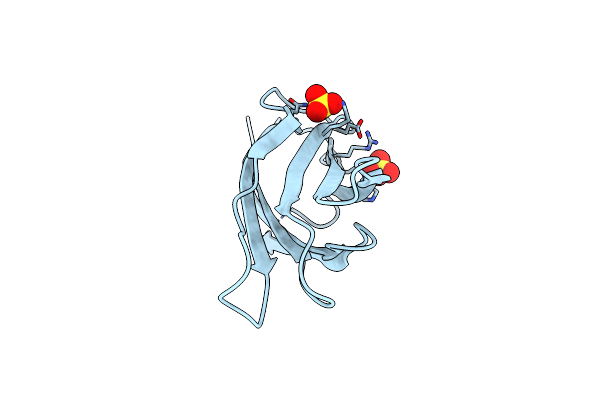 |
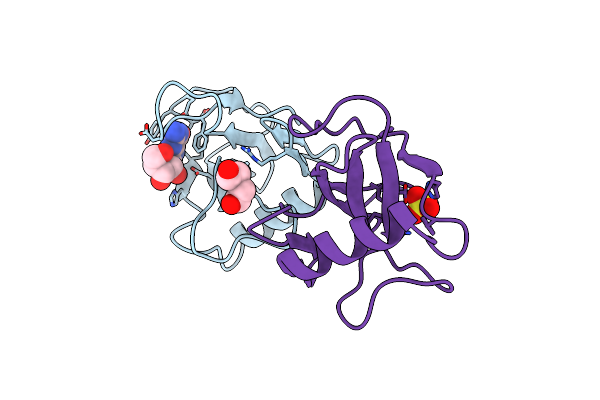
Organism: White spot syndrome virus
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2021-06-23 Classification: VIRAL PROTEIN Ligands: SO4 |
 |
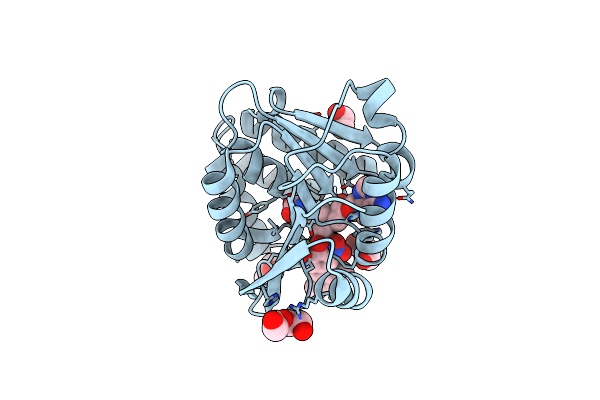
Organism: Hericium erinaceus
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2021-01-20 Classification: RNA BINDING PROTEIN Ligands: GMP, PEG, SO4 |
 |
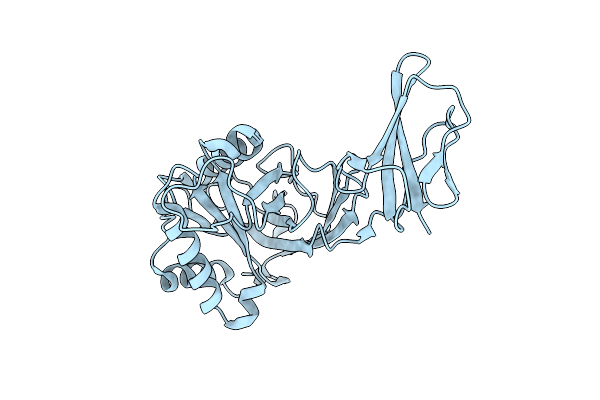
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2020-03-04 Classification: TRANSFERASE Ligands: SAM, EAO, MG, IPA, PEG |
 |
Organism: Porphyromonas gingivalis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2018-07-25 Classification: ELECTRON TRANSPORT |
 |
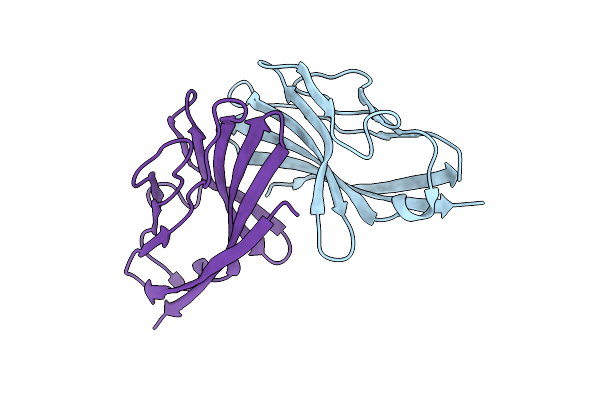
Organism: Hericium erinaceus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2017-09-27 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2016-12-28 Classification: CELL ADHESION |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2016-12-28 Classification: CELL ADHESION |

