Search Count: 59,445
All
Selected
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2023-08-09 Classification: TRANSFERASE Ligands: SCA |
 |
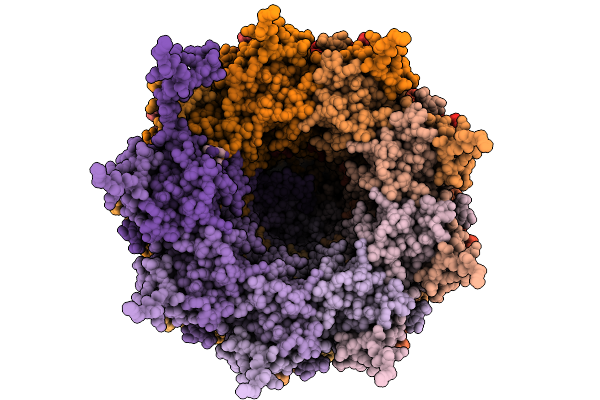
Ssrna-Containing Helical Virus-Like Particle Composed Of Pepmov Coat Protein
Organism: Pepper mottle virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRUS LIKE PARTICLE |
 |
 |
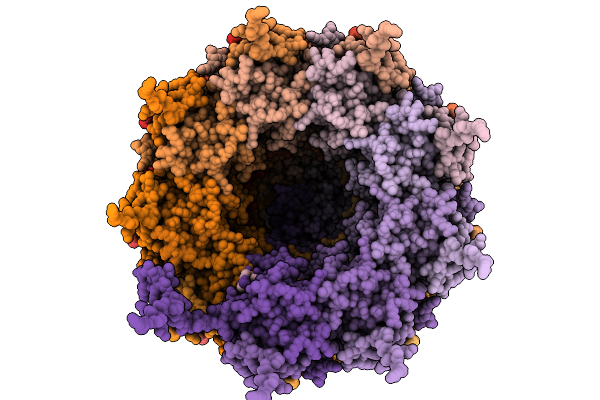
Ssrna-Containing Helical Virus-Like Particle Composed Of Pva (Isolate B11) Coat Protein
Organism: Potato virus a
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRUS LIKE PARTICLE |
 |
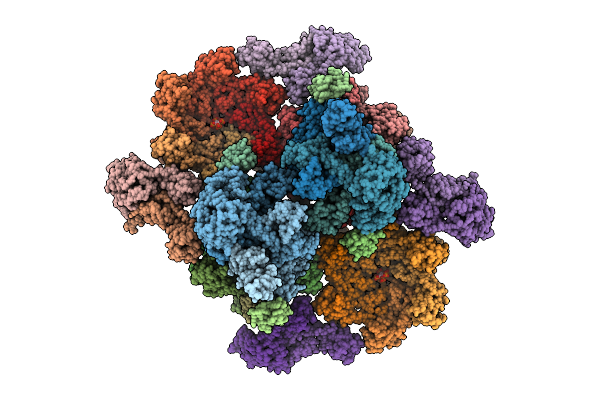 |
Cryoem Reconstruction Of Integrase Filament At The Lumen Of Native Hiv-1 Cores (Box Size 34.2 Nm)
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Resolution:4.63 Å Release Date: 2025-12-10 Classification: VIRAL PROTEIN Ligands: ZN, IHP |
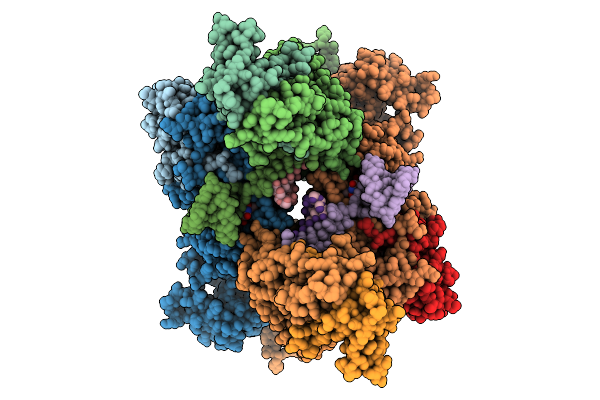 |
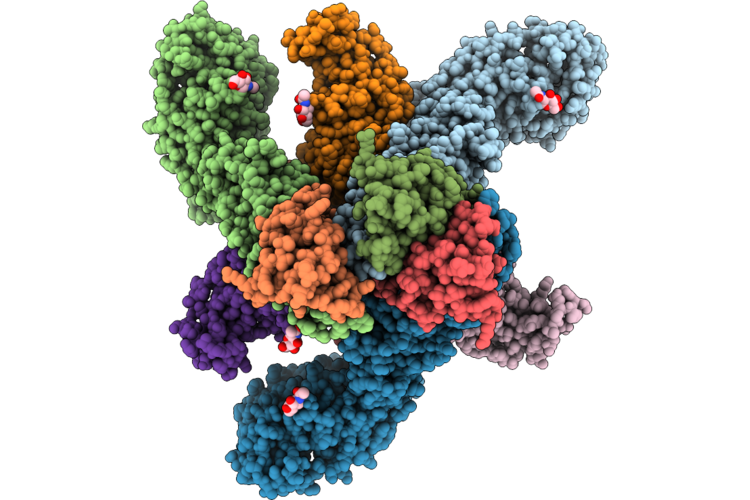
Cryo-Em Structure Of Sars-Cov-2 Nsp10-Nsp14 (E191A) In Complex With T20P14-S Complex, Tetrameric Form
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: VIRAL PROTEIN/RNA Ligands: ZN, MG, K5X |
 |
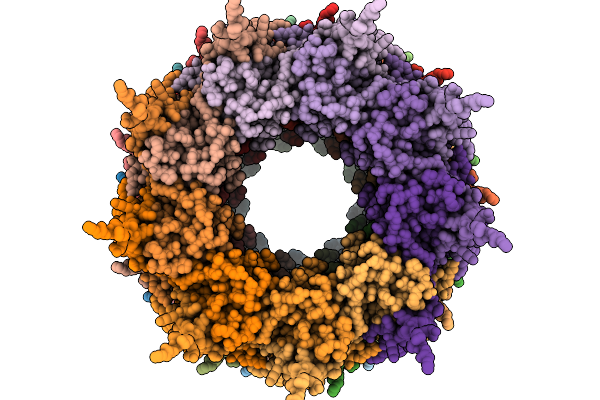
Organism: Tick-borne encephalitis virus
Method: ELECTRON MICROSCOPY Resolution:3.57 Å Release Date: 2026-05-20 Classification: VIRUS Ligands: NAG |
 |
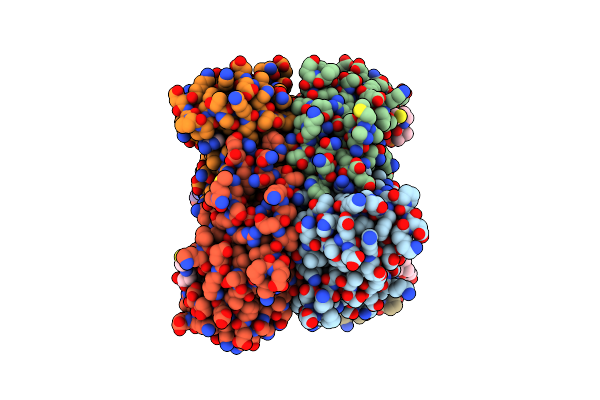
Rna-Free Stacked-Ring (R9) Virus-Like Particle Composed Of Pepmov Coat Protein
Organism: Pepper mottle virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRUS LIKE PARTICLE |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2023-08-09 Classification: TRANSFERASE Ligands: COA |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2023-08-09 Classification: TRANSFERASE Ligands: ADP |
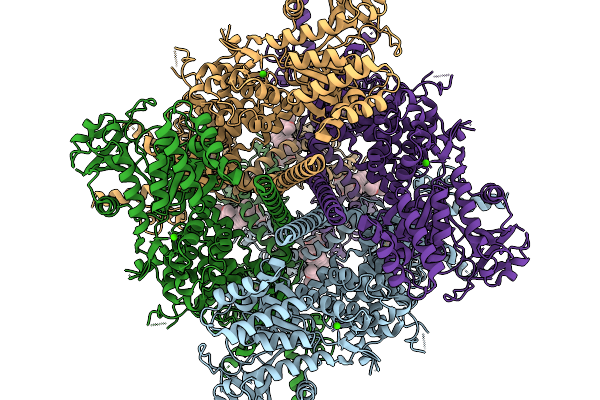 |
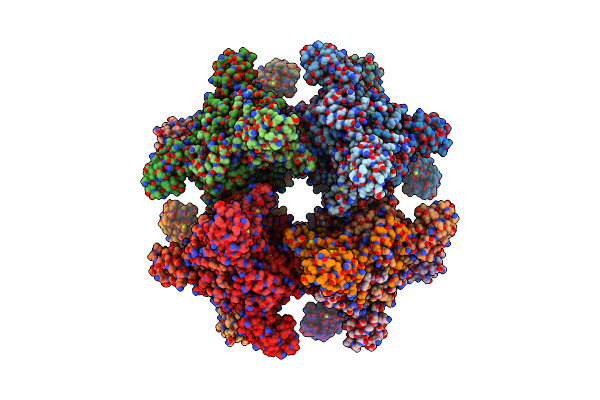
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Resolution:2.75 Å Release Date: 2026-01-07 Classification: TRANSFERASE Ligands: NAG, CA, A1CGZ, A1CGW, A1CG0 |
 |
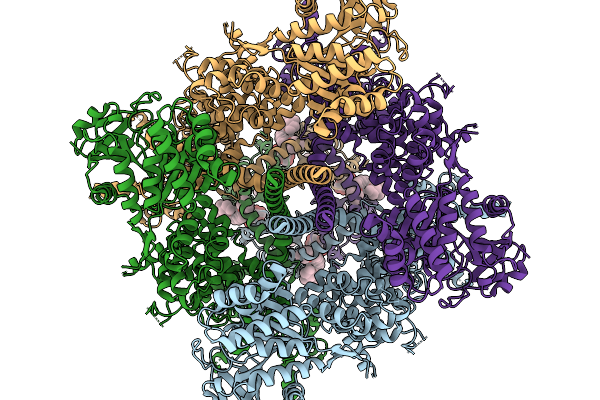
Cryo-Em Structure Of The Dihydrolipoyl Transacetylase Cubic Core Of The E. Coli Pyruvate Dehydrogenase Complex Including Lipoyl Domains
Organism: Escherichia coli (strain k12)
Method: ELECTRON MICROSCOPY Release Date: 2021-08-11 Classification: TRANSFERASE |
 |
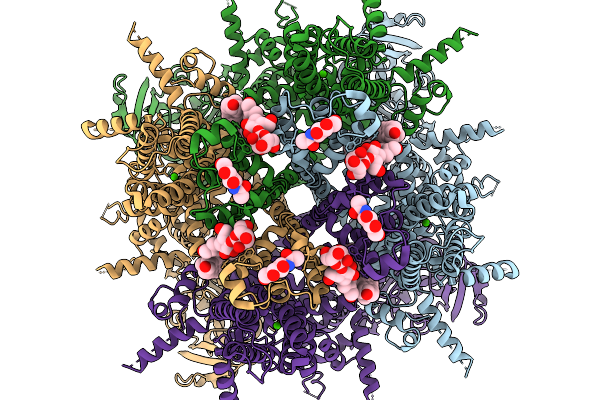
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: TRANSFERASE Ligands: NAG, CA, A1CGZ, A1CGW, A1CG0 |
 |
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Resolution:2.85 Å Release Date: 2026-01-07 Classification: TRANSFERASE Ligands: NAG, CA, A1CGZ, A1CG0 |
 |
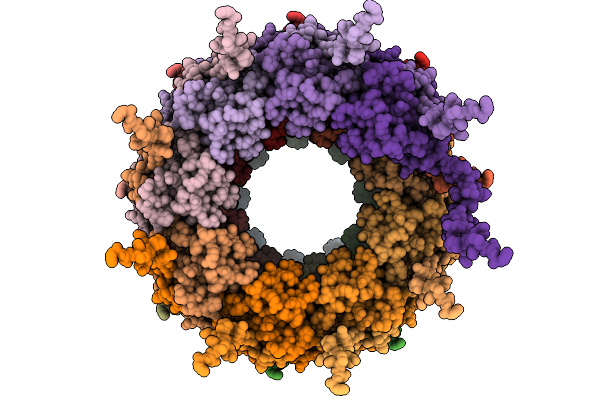
Organism: Tobacco etch virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRUS LIKE PARTICLE |
 |
Rna-Free Helical (H8.5) Virus-Like Particle Composed Of Pva (Isolate B11) Coat Protein
Organism: Potato virus a
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRUS LIKE PARTICLE |
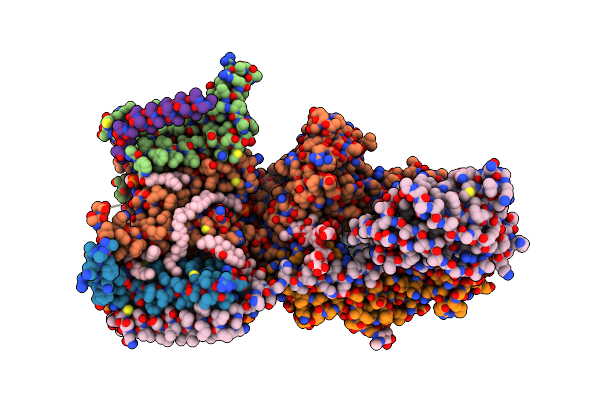 |
Cryo-Em Structure Of Yeast Ost6P Containing Oligosaccharyltransferase Complex
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2022-02-16 Classification: MEMBRANE PROTEIN Ligands: PEE, NAG, AJP, MG, V8K |
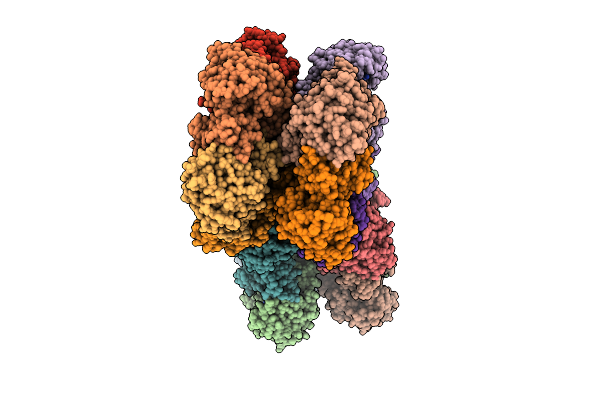 |
Crystal Structure Of Human Glutamine Synthetase In Complex With Adp And Phosphate
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-03-11 Classification: LIGASE Ligands: ADP, MN, PO4, NA |
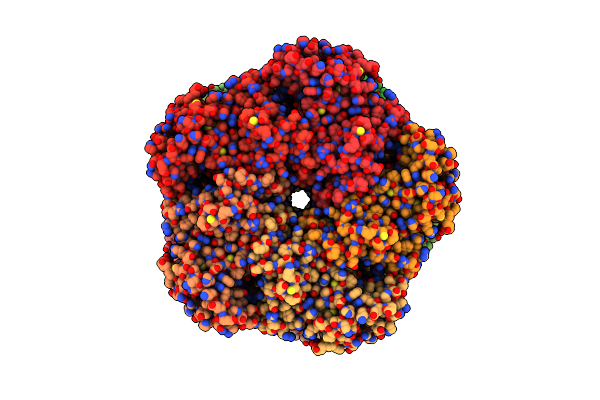 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: LIGASE Ligands: MG |

