Search Count: 267
All
Selected
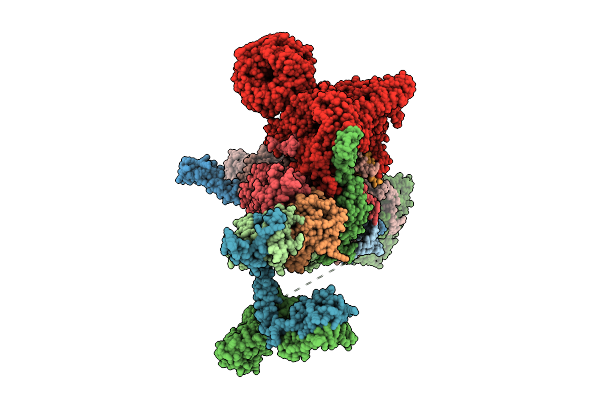 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Proximal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
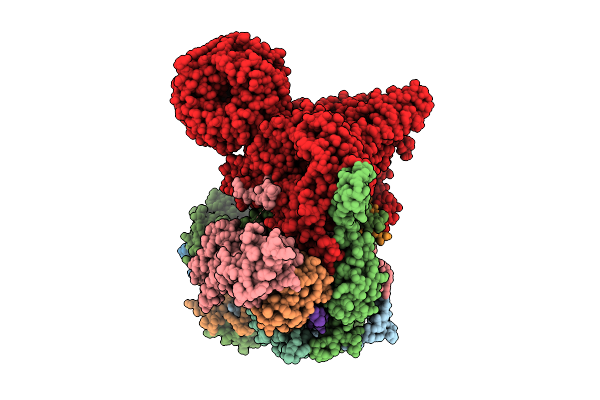 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Core)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
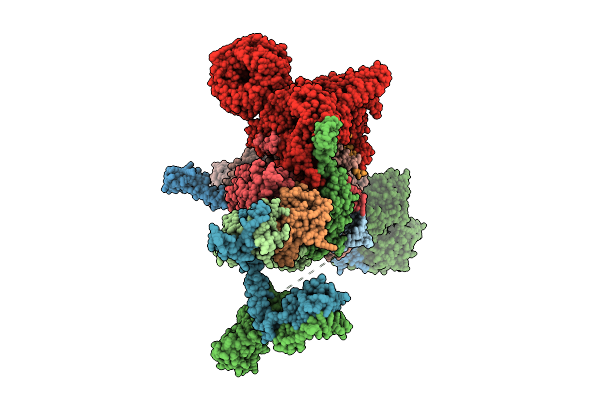 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Distal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
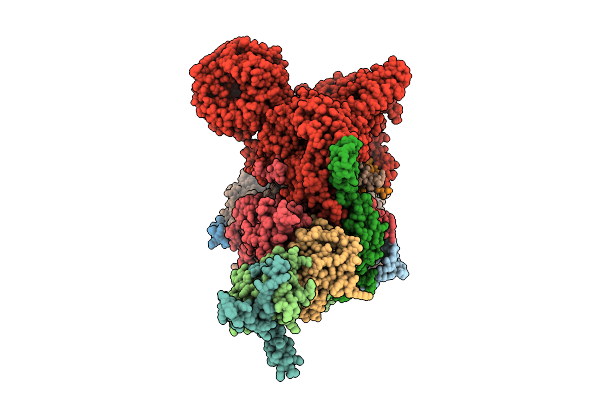 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Sf3A State 1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
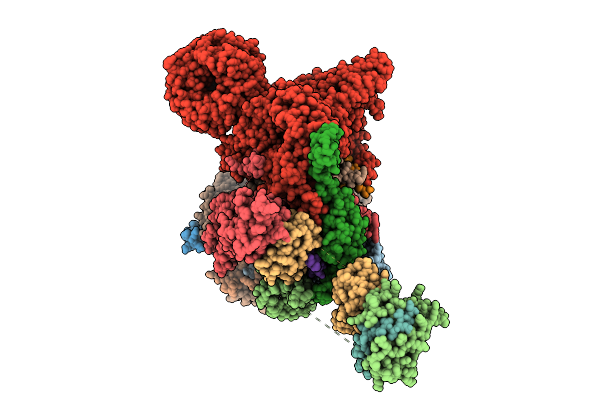 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Sf3A State 2)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
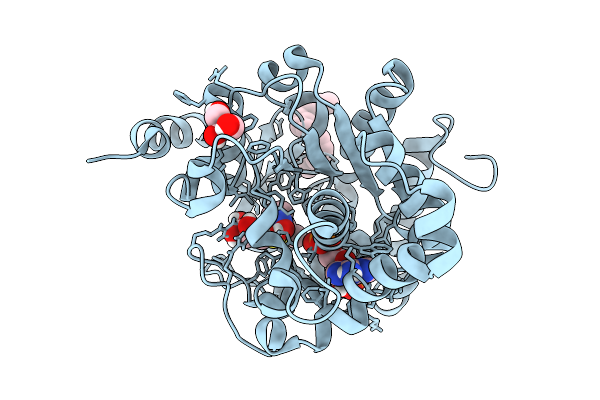 |
Crystal Structure Of Arabidopsis Thaliana Sulfotransferase Sot18 Complexed With Glucoraphanin Precursor Involved In Glucosinolate Biosynthesis
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2026-03-04 Classification: CYTOSOLIC PROTEIN Ligands: A3P, GOL, A1L7V, PLM |
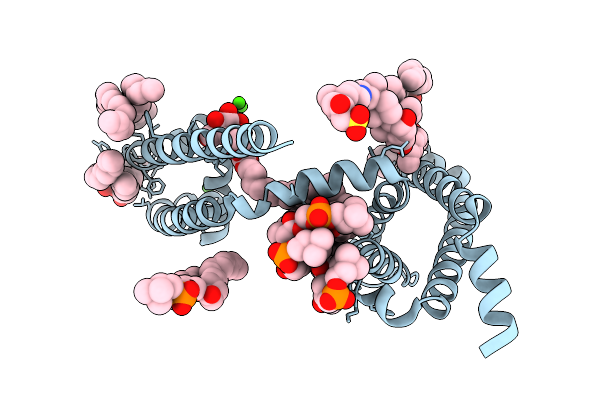 |
Organism: Aliarcobacter butzleri
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: CA, 1N7, LMT, PX4 |
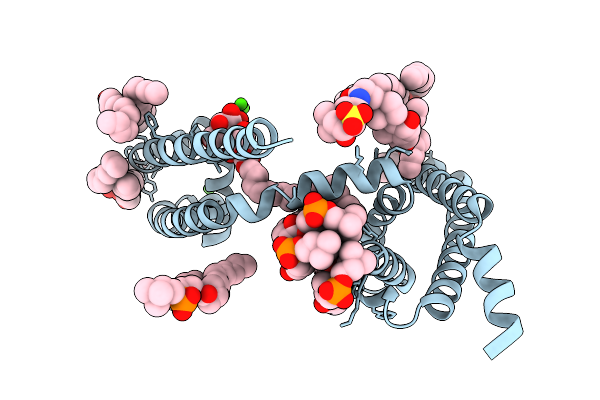 |
Organism: Aliarcobacter butzleri
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: CA, 1N7, LMT, PX4 |
 |
Organism: Aliarcobacter butzleri
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: CA, 1N7, LMT, PX4 |
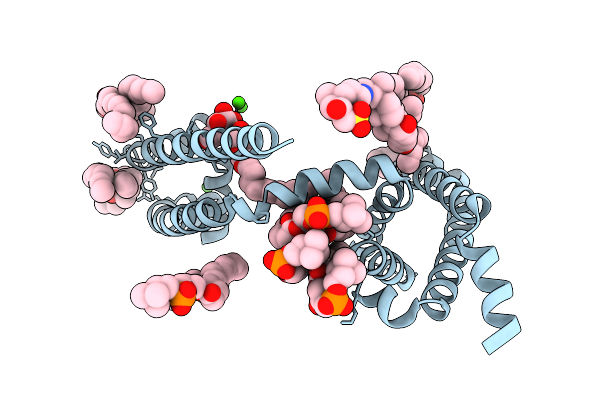
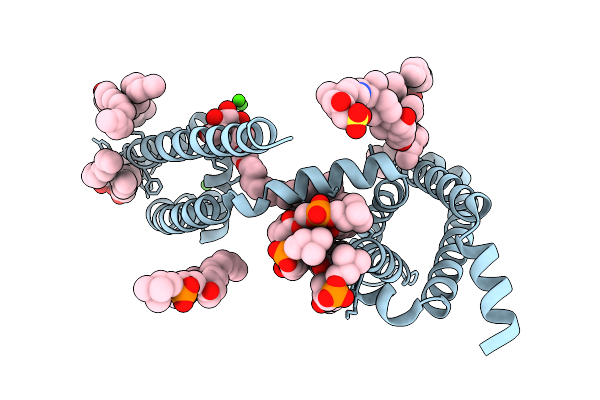 |
Crystal Structure Of Voltage-Gated Sodium Channel Navab N49K/S178T/T206A Mutant
Organism: Aliarcobacter butzleri
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: CA, 1N7, LMT, PX4 |
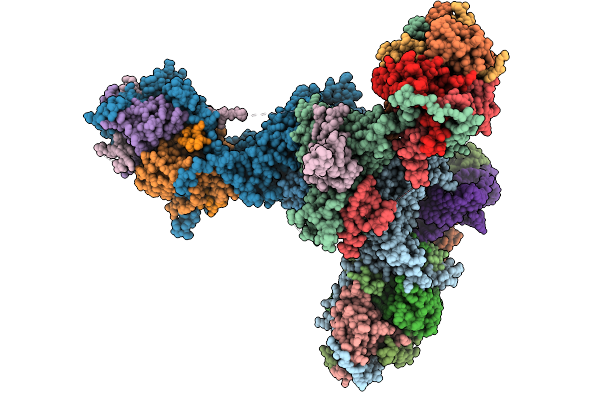 |
Trypanosoma Brucei Mitochondrial Rna-Editing Catalytic Complex 1, U-Deletion (Recc1)
Organism: Trypanosoma brucei
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: RNA BINDING PROTEIN Ligands: ZN, MG, 5GP |
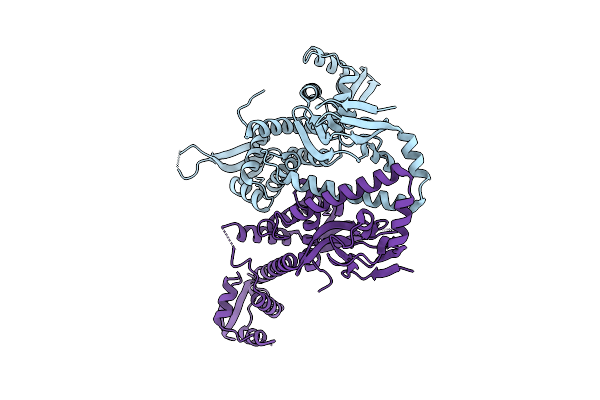 |
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2025-10-22 Classification: SIGNALING PROTEIN |
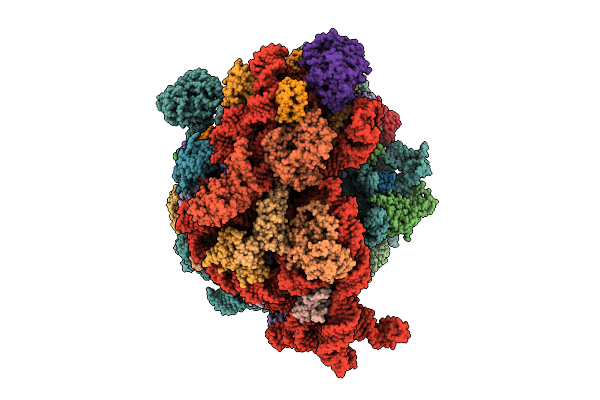 |
Cryo-Em Structure Of Escherichia Coli Hibernating Ribosome With Rnase I Mutant
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: RIBOSOME Ligands: MG, CA |
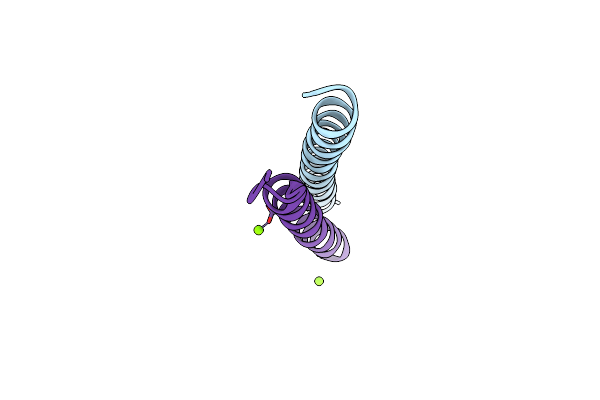 |
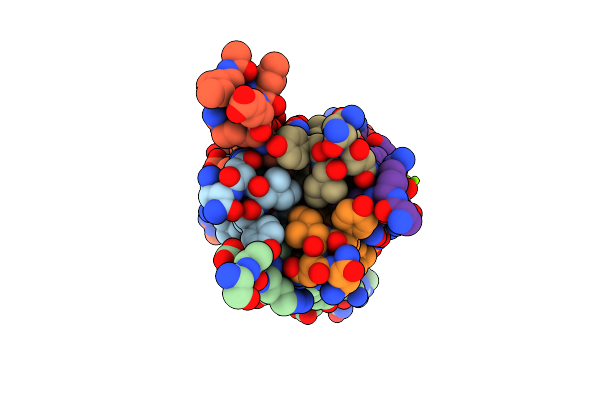
Crystal Structure Of Measles Virus Fusion Inhibitor M1 Complexed With F Protein Hr1 (Hr1-42) (H32 Space Group)
Organism: Measles virus strain edmonston
Method: X-RAY DIFFRACTION Resolution:1.16 Å Release Date: 2025-01-01 Classification: ANTIVIRAL PROTEIN Ligands: MG |
 |
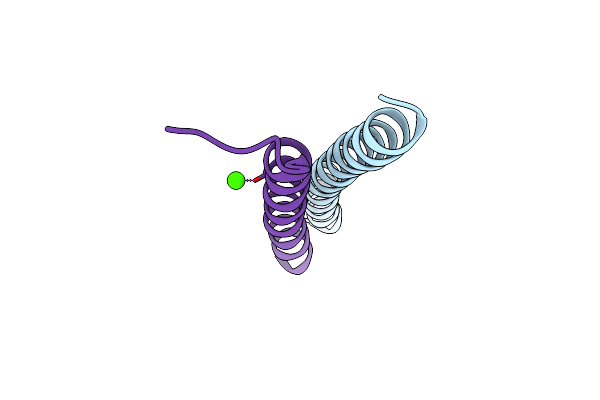
Crystal Structure Of Measles Virus Fusion Inhibitor M1 Complexed With F Protein Hr1 (Hr1-42) (P21212 Space Group)
Organism: Measles virus strain edmonston
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2025-01-01 Classification: ANTIVIRAL PROTEIN Ligands: MG |
 |
Crystal Structure Of Measles Virus Fusion Inhibitor M1 Complexed With F Protein Hr1 (Hr1-47) (P321 Space Group)
Organism: Measles virus strain edmonston
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2025-01-01 Classification: ANTIVIRAL PROTEIN Ligands: CA |
 |
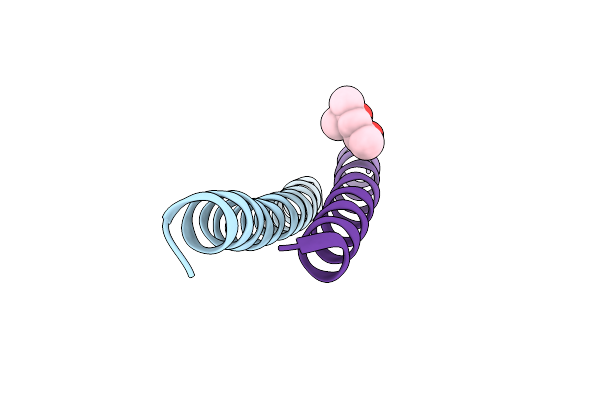
Crystal Structure Of Measles Virus Fusion Inhibitor M1Ek Complexed With F Protein Hr1 (Hr1-42) (P21 Space Group)
Organism: Measles virus strain edmonston
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2025-01-01 Classification: ANTIVIRAL PROTEIN |
 |
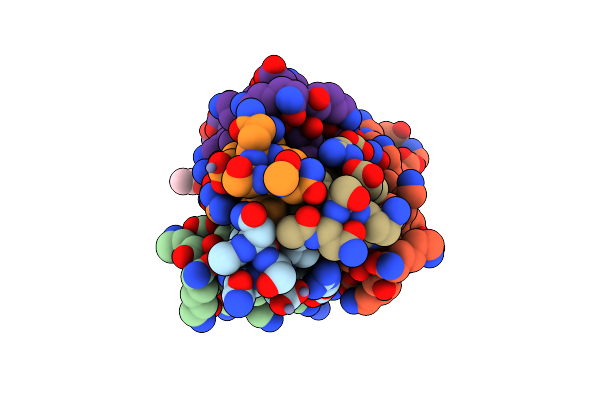
Crystal Structure Of Measles Virus Fusion Inhibitor Mek28 Complexed With F Protein Hr1 (Hr1-40) (H3 Space Group)
Organism: Measles virus strain edmonston
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2025-01-01 Classification: ANTIVIRAL PROTEIN Ligands: MPD |
 |
Crystal Structure Of Measles Virus Fusion Inhibitor Mek35Ge Complexed With F Protein Hr1 (Hr1-42) (P21212 Space Group)
Organism: Measles virus strain edmonston
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2025-01-01 Classification: ANTIVIRAL PROTEIN Ligands: ZN, ACT |
 |
Crystal Structure Of Measles Virus Fusion Inhibitor Mek35Ge Complexed With F Protein Hr1 (Hr1-42) (P2 Space Group)
Organism: Measles virus strain edmonston
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2025-01-01 Classification: ANTIVIRAL PROTEIN |

