Search Count: 3,753
All
Selected
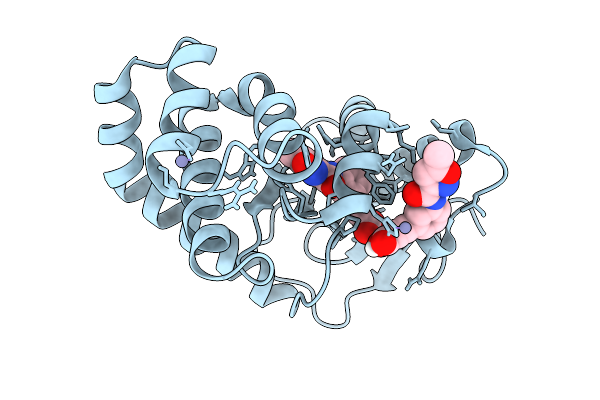 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: ZN, A1CJH |
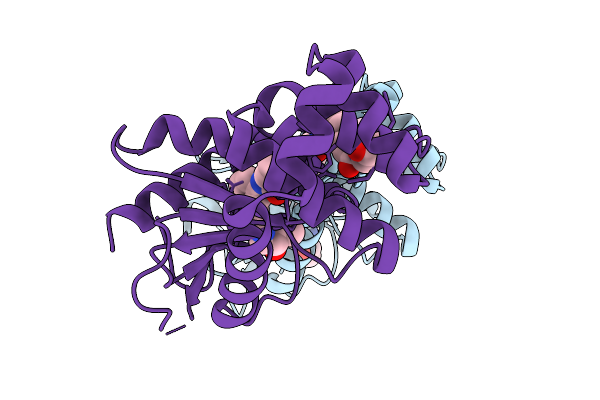 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CQW, A1CJJ |
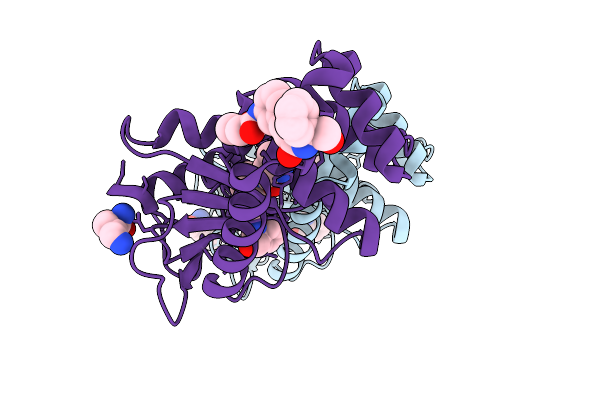 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: LYS, A1CJK |
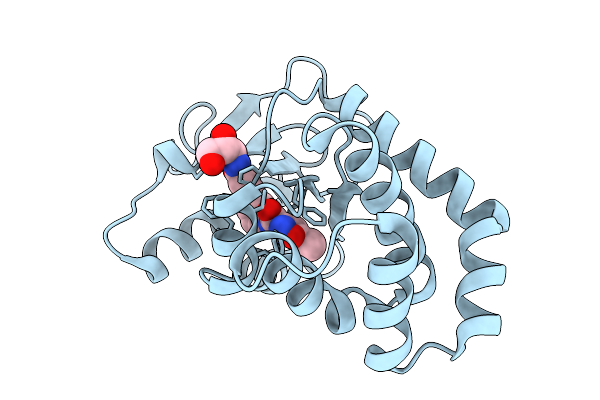 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJL |
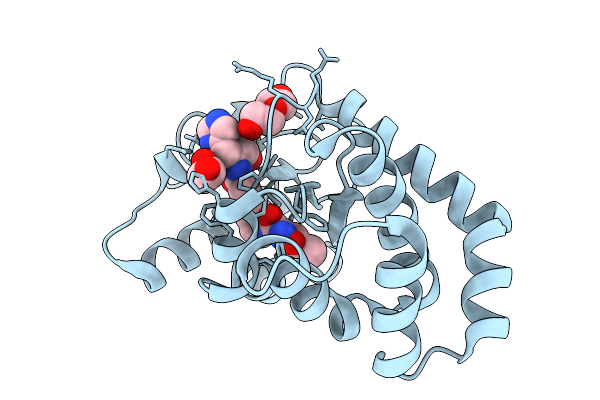 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJM, 1PE, EDO |
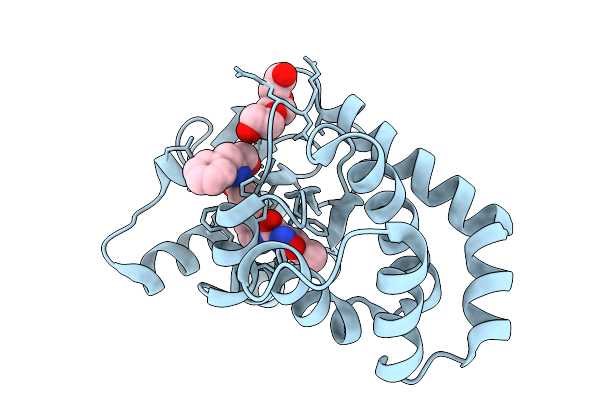 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJN, PGE |
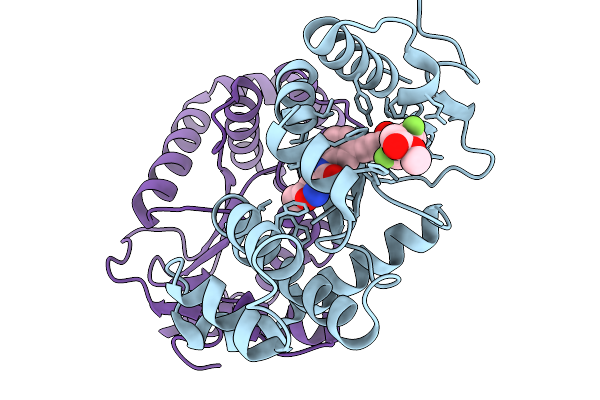 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJO |
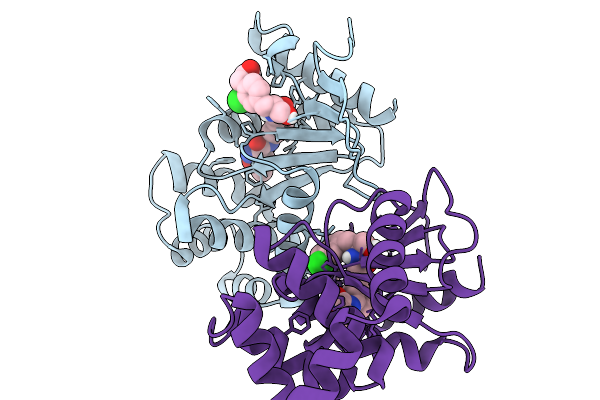 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJP |
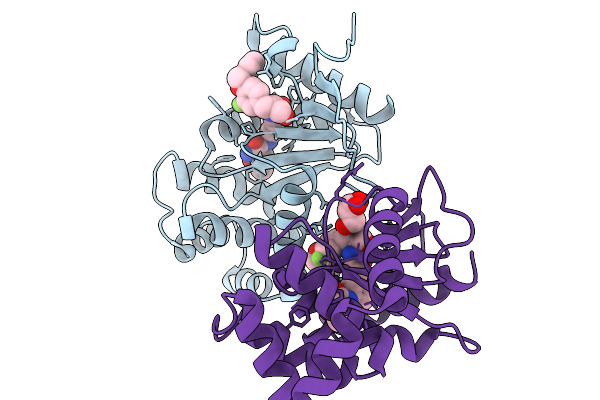 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJQ, GOL |
 |
Scia(H42Q) Endonuclease From Escherichia Phage T5 In Complex With 10 Bp Dna (Gcgc Sequence 2)
Organism: Escherichia phage t5
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN Ligands: IMD, ZN |
 |
Scia(H42A) Endonuclease From Escherichia Phage T5 In Complex With 10 Bp Dna (Gcgc Sequence 1)
Organism: Escherichia phage t5
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN Ligands: IMD, ACT, ZN |
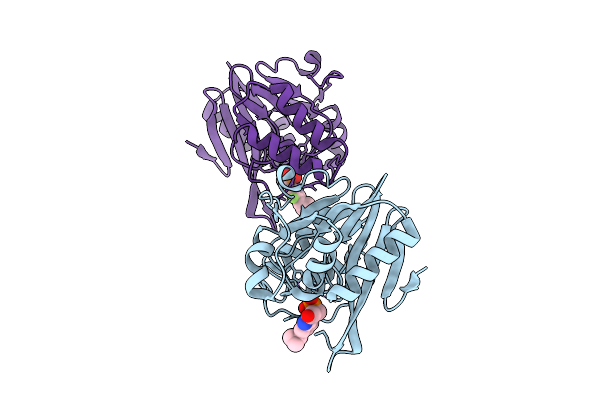 |
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: A1B7Y, ZN |
 |
Context-Dependent Variability Of Hif Heterodimers Influences Interactions With Macromolecular And Small Molecule Partners
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-03-11 Classification: PROTEIN BINDING |
 |
Crystal Structure Of Kirsten Rat Sarcoma G12C Complexed With Gmppnp And Covalently Bound To 1-[(2R,3R)-3-{[(7P)-7-(8-Ethynyl-7-Fluoronaphthalen-1-Yl)-8-Fluoro-2-{ [(2R,4R,7As)-2-Fluorotetrahydro-1H-Pyrrolizin-7A(5H)-Yl]Methoxy}Pyrido[4,3-D] Pyrimidin-4-Yl](Methyl)Amino}-2-Methylpyrrolidin-1-Yl]-3-(Pyrazin-2-Yl)Propan-1-One
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: GDP, A1C5K, CA |
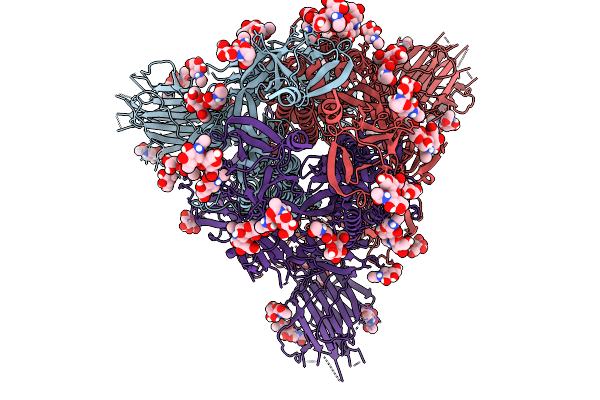 |
Cryo-Em Consensus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 1 Up & 2 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
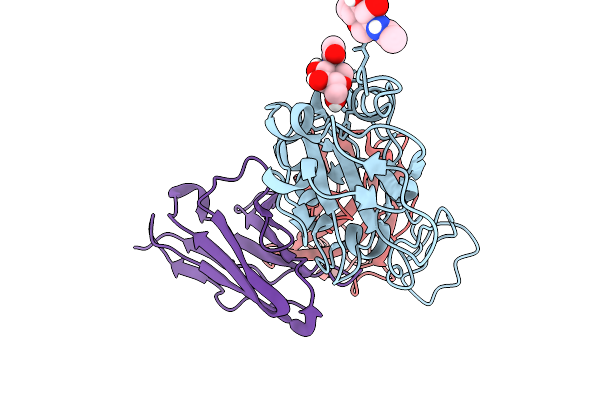 |
Cryo-Em Focus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 1 Up & 2 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
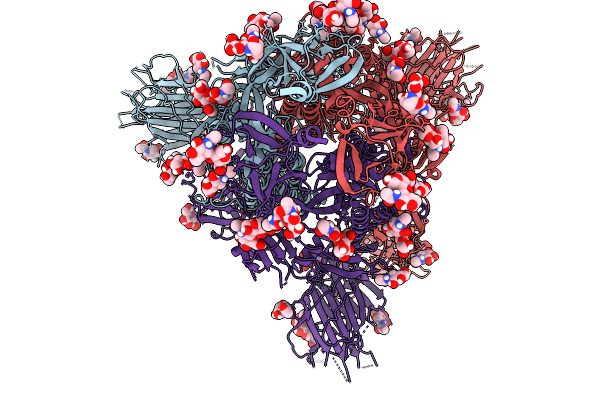 |
Cryo-Em Consensus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 2 Up & 1 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
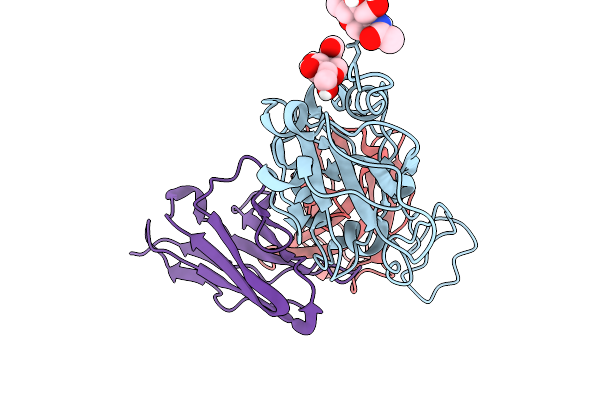 |
Cryo-Em Focus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 2 Up & 1 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
 |
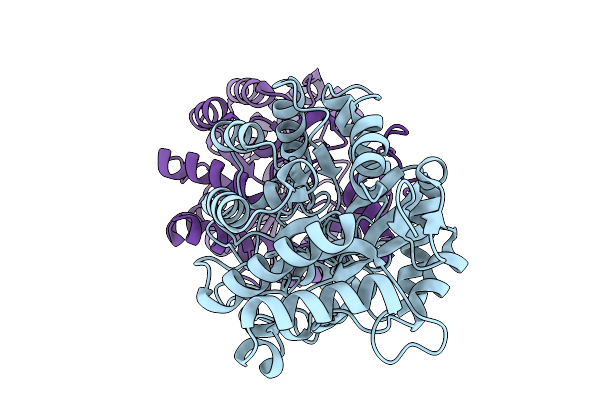
Cryo-Em Structure Of Human Atp Citrate Lyase In Complex With Inhibitor Evt0185-Coa
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: LYASE Ligands: ADP, W1K |
 |
Chemical Inhibition Of The N-Acetyltaurine Amidohydrolase Pter Reduces Food Intake And Obesity
Organism: Amphimedon queenslandica
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: ZN |

